Major Article
INTRODUCTION
Address to:Dr.Carlos José Pereira da Cunha de Araújo-Coutinho. Laboratório de Entomologia Médica/SUCEN. Rua Cardeal Arcoverde no 2878, 05408-003 São Paulo, SP, Brasil.
Phone: 55 11 3032-2228 e-mail: [email protected] Received 18 July 2014
Accepted 9 October 2014
Description of microsporidia in simulids: molecular and
morphological characterization of microsporidia in the
larvae of
Simulium pertinax
Kollar (Diptera: Simuliidae)
Isabel Maria Vicente Guedes de Carvalho
[1], Artur Trancoso Lopo de Queiroz
[2],
Rosiane Brito de Moraes
[4], Helio Benites Gil
[3], Rafael Alves
[5],
Andréa de Barros Pinto Viviani
[6], James John Becnel
[7]and Carlos José Pereira da Cunha de Araujo-Coutinho
[4][1]. Laboratório de Parasitologia, Instituto Butantan, São Paulo, SP. [2]. Laboratório de Imunoparasitologia, Centro de Pesquisa Gonçalo Moniz, Fundação Oswaldo Cruz, Salvador, BA. [3]. Disciplina de Infectologia, Universidade Federal de São Paulo, São Paulo, SP. [4]. Laboratório de Entomologia Médica, Superintendência de Controle de Endemias, São Paulo, SP. [5]. Departamento de Medicina, Disciplina de Gastroenterologia, Universidade Federal de São Paulo, São Paulo, SP. [6]. Laboratório de Simulídeos, Superintendência de Controle de Endemias, Caraguatatuba, SP. [7]. United States Department of Agriculture, Mosquito and Fly Research Unit, Gainesville, Florida, USA.
ABSTRACT
Introduction: Microsporidia constitute the most common black fl y pathogens, although the species’ diversity, seasonal occurrence
and transmission mechanisms remain poorly understood. Infections by this agent are often chronic and non-lethal, but they can cause reduced fecundity and decreased longevity. The objective of this study was to identify microsporidia infecting Simulium
(Chirostilbia) pertinax (Kollar, 1832) larvae from Caraguatatuba, State of São Paulo, Brazil, by molecular and morphological characterization. Methods: Larvae were collected at a single point in a stream in a rural area of the city and were kept under artifi cial aeration until analysis. Polydispyrenia spp. infection was characterized by the presence of at least 32 mononuclear spores measuring 6.9 ± 1.0 x 5.0 ± 0.7µm in persistent sporophorous vesicles. Similarly, Amblyospora spp. were characterized by the presence of eight uninucleate spores measuring 4.5 x 3.5µm in sporophorous vesicles. Results: The molecular analysis confi rmed
the presence of microsporidian DNA in the 8 samples (prevalence of 0.51%). Six samples (Brazilian larvae) were related to
Polydispyrenia simulii and Caudospora palustris reference sequences but in separate clusters. One sample was clustered with
Amblyospora spp. Edhazardia aedis was the positive control taxon. Conclusions: Samples identifi ed as Polydispyrenia spp. and
Amblyospora spp. were grouped with P. simulii and Amblyospora spp., respectively, corroborating previous results. However, the 16S gene tree showed a considerable distance between the black fl y-infecting Amblyospora spp. and the mosquito-infecting spp. This distance suggests that these two groups are not congeneric. Additional genomic region evaluation is necessary to obtain a coherent phylogeny for this group.
Keywords: Microsporidae. Amblyospora spp. Polydispyrenia spp.Phylogenetic analysis.
Black fl ies (Diptera: Simuliidae) cause severe medical and veterinary problems worldwide. Simuliidae species are able to transmit parasites that can result in severe disease in humans and animals. In addition, their bites can cause allergic reactions and dermatitis in sensitized individuals, resulting in severe economic losses to tourism centers and negatively impacting animal production1-3. Black fl y control remains a
major public health challenge. Microsporidia are unicellular, eukaryotic organisms that are obligate, intracellular parasites with public health relevance4. Several studies have suggested a new classifi cation for microsporidia as fungi, but Ebersberger5 stated that phylogenetic analysis did not support fungal characterization for this group.
Microsporidia are the most common black fl y pathogens, although the species’ diversity, seasonal occurrence and transmission mechanisms remain poorly understood6,7. Infections caused by this agent are often chronic and non-lethal, but they can cause sub-lethal host effects, such as reduced fecundity, decreased life span and general loss of vigor8.
The objective of this study was to identify microsporidian species infecting Simulium (Chirostilbia) pertinax (Kollar, 1832) larvae from Caraguatatuba City, on the north coast of State of São Paulo, by molecular and morphological characterization.
METHODS
fl y bites annoy visitors and have deleterious effects on the local economy. Monitoring and controlling black fl ies are essential to avoiding seasonal population outbreaks.
Sampling and biological material processing
The sampling period was from May to August 2013, and the samples were collected from a stream in Caraguatatuba City, located on the north coast of the State of São Paulo, Brazil,which has a total area of 458,097km2 and had a population at that time of 100,8409. All of the larvae were held in aerated containers with water from the breeding site until examination. Tissues showing evidence of infection (whitish abdomens or whitish digestive tracts) were dissected in NaCl 0.9% solution, and fat bodies and adjacent tissues were removed10. Processed samples were frozen in 1.5ml tubes with 30µl of diethylpyrocarbonate (DEPC)(Invitrogen® Life Technologies, Carlsbad, CA, USA). Fresh smears of fat bodies were made, fi xed with methanol for 5min and stained with 10% Giemsa in 7.4 pH buffer for 20min. The slides were washed in water and dried at 25°C overnight11 for further morphological analysis of spores.
Morphological analysis
The Nis Elements F 3.0 NIKON H550S software, with phase III objective scale 100X settings, was used for spore measurement. Morphological characterization was performed according to Sprague12.
Molecular assay
Molecular assays were performed with frozen tissues from infected larvae, and Aedes aegypti larvae infected with
Edhazardia aedis were used as positive controls.
DNA extraction
Larvae exhibiting symptoms of infection had deoxyribonucleic acid (DNA) extracted using a viral DNA kit (QIAamp® viral RNA, Qiagen, Inc, Hilden, Germany). Healthy larvae (Figure 1A)
were discarded. Tissue samples were processed with a proteinase K kit, incubated at 56°C for 2h and mixed every 20min. The supernatants were used to amplify the r16S ribosomal gene13.
Small subunit ribosomal gene (SSUrDNA) PCR (r16S)
Polymerase chain reaction (PCR) amplifi cation was performed with 18f (CAC CAG GTT GAT TCT GCC) and 1492r (GGT TAC CTT GTT ACG ACT T), according to Vossbrinck et al.14.
The amplifi cation products were visualized on 2% agarose gels, with positive and negative controls and a 100 bps ladder (Invitrogen® Life Technologies, Carlsbad, CA, USA), following electrophoresis.
Nucleotide sequencing
PCR products were purifi ed with the Illustra GFX PCR DNA and Gel Band Purifi cation Kit (GE Healthcare Limited, Little Chalfont, Buckinghamshire, UK) and were quantifi ed with 2% agarose gel ethidium bromide staining, according to the Low
DNA Mass Ladder (Invitrogen®) protocol. The products were sequenced using an ABI PRISM Big Dye Terminator Cycle Sequencing Ready Reaction kit (PE Applied Biosystems), following the standard manufacturer protocols. The data were analyzed with the phred/phrap software, and the contigs were assembled with the cap3 software15.
Phylogenetic analysis
The analyses were performed using the Seaview software16. A phylogenetic tree was constructed, with reference sequences32-46 from Table 1 (supplementary fi le), using the maximum likelihood method with the general time reversible (GTR) model of nucleotide substitution and gamma distribution (G) (GTR + G)17. The model was selected by the Modeltest software, version 3.0.618, and was optimized by the Seaview software. We calculated the bootstrap values with 1,000 replications to support the verification of branches in the topologies of the trees obtained, and bootstrap values greater than 70 were considered signifi cant.
Nucleotide sequences and accession numbers
The nucleotide sequences obtained in this work were submitted to the GenBank nucleotide sequences databank under the following accession numbers GenBank: KC855552-KC855557 (L1_L6); and GenBank: KC855558 (L2).
A
B
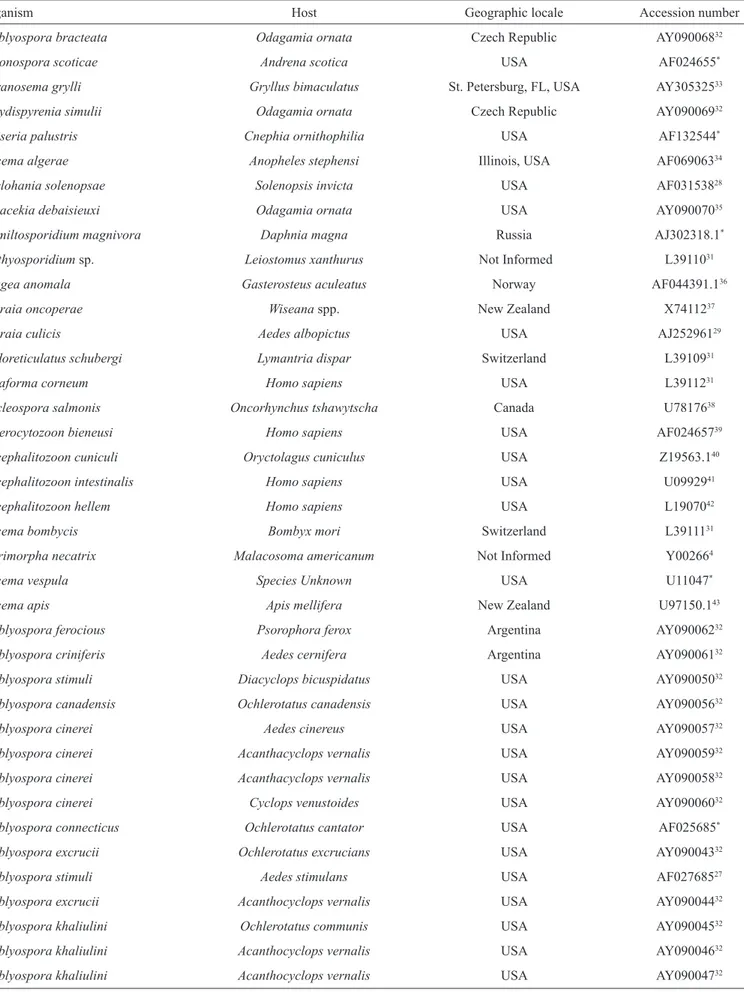
TABLE 1 - Sequences and accession numbers used for phylogenetic analysis.
Organism Host Geographic locale Accession number
Amblyospora bracteata Odagamia ornata Czech Republic AY09006832
Antonospora scoticae Andrena scotica USA AF024655*
Paranosema grylli Gryllus bimaculatus St. Petersburg, FL, USA AY30532533
Polydispyrenia simulii Odagamia ornata Czech Republic AY09006932
Weiseria palustris Cnephia ornithophilia USA AF132544*
Nosema algerae Anopheles stephensi Illinois, USA AF06906334
Thelohania solenopsae Solenopsis invicta USA AF03153828
Janacekia debaisieuxi Odagamia ornata USA AY09007035
Hamiltosporidium magnivora Daphnia magna Russia AJ302318.1*
Ichthyosporidium sp. Leiostomus xanthurus Not Informed L3911031
Glugea anomala Gasterosteus aculeatus Norway AF044391.136
Vavraia oncoperae Wiseana spp. New Zealand X7411237
Vavraia culicis Aedes albopictus USA AJ25296129
Endoreticulatus schubergi Lymantria dispar Switzerland L3910931
Vittaforma corneum Homo sapiens USA L3911231
Nucleospora salmonis Oncorhynchus tshawytscha Canada U7817638
Enterocytozoon bieneusi Homo sapiens USA AF02465739
Encephalitozoon cuniculi Oryctolagus cuniculus USA Z19563.140
Encephalitozoon intestinalis Homo sapiens USA U0992941
Encephalitozoon hellem Homo sapiens USA L1907042
Nosema bombycis Bombyx mori Switzerland L3911131
Vairimorpha necatrix Malacosoma americanum Not Informed Y002664
Nosema vespula Species Unknown USA U11047*
Nosema apis Apis mellifera New Zealand U97150.143
Amblyospora ferocious Psorophora ferox Argentina AY09006232
Amblyospora criniferis Aedes cernifera Argentina AY09006132
Amblyospora stimuli Diacyclops bicuspidatus USA AY09005032
Amblyospora canadensis Ochlerotatus canadensis USA AY09005632
Amblyospora cinerei Aedes cinereus USA AY09005732
Amblyospora cinerei Acanthacyclops vernalis USA AY09005932
Amblyospora cinerei Acanthacyclops vernalis USA AY09005832
Amblyospora cinerei Cyclops venustoides USA AY09006032
Amblyospora connecticus Ochlerotatus cantator USA AF025685*
Amblyospora excrucii Ochlerotatus excrucians USA AY09004332
Amblyospora stimuli Aedes stimulans USA AF02768527
Amblyospora excrucii Acanthocyclops vernalis USA AY09004432
Amblyospora khaliulini Ochlerotatus communis USA AY09004532
Amblyospora khaliulini Acanthocyclops vernalis USA AY09004632
Amblyospora khaliulini Acanthocyclops vernalis USA AY09004732
TABLE 1 - Continuation.
Organism Host Geographic locale Accession number
Amblyospora weiseri Ochlerotatus cantans USA AY09004832
Amblyospora stictici Ochlerotatus sticticus USA AY09004932
Edhazardia aedis Aedes aegypti Thailand AF02768427
Amblyospora sp. Cyclops strenuus Czech Republic AY09005532
Amblyospora californica Culex tarsulis USA U6847344
Amblyospora sp. Culex nigripalpus USA AY09005332
Amblyospora sp. Culex salinarius USA U6847444
Amblyospora salinaria Culex salinarius USA AY32627032
Culicospora magna Culex restuans USA AY09005432
Culicospora magna Culex restuans USA AY32626932
Intrapredatorus barri Culex fuscanus Norway AY01335945
Amblyospora indicola Culex sitiens India AY09005132
Amblyospora opacita Culex territans USA AY09005232
Hyalinocysta chapmani Culiseta melanura USA AF48383746
Hyalinocysta chapmani Orthocyclops modestus USA AF48383846
Culicosporella lunata Culex pilosus USA AF02768327
Parathelohania anophelis Anopheles quadrimaculatus USA AF02768227
Parathelohania obesa Anopheles crucians USA AY09006532
Trichotuzetia guttata Cyclops vicinus Czech Republic AY32626832
Hazardia milleri Culex quinquefasciatus Argentina AY09006732
Hazardia sp. Anopheles crucians USA AY09006632
Marsoniella elegans Cyclops vicinus Czech Republic AY09004132
Gurleya vavrai Daphnia longispina Finland AF39452630
Gurleya daphniae Daphnia pulex Austria AF43932030
Larssonia obtusa Daphnia pulex Sweden AF39452730
Berwaldia schaefernai Daphnia galeata Czech Republic AY09004232
Varimorpha sp. Solenopsis richteri USA AF03153928
Amblyospora sp. Simulium sp. UK AJ25294929
USA: United States of America; FL: Florida; UK: United Kingdom.*Unpublished.
RESULTS
A total of 1,574 S. pertinax larvae were examined. Eight larvae exhibited symptoms of microsporidian infection localized to the fat body (Figure 1B).
Morphological characterization indicated Polydispyrenia
spp. infections in 7 larvae (Figure 2A), representing 87.5% of the infected larvae. Amblyospora sp. infection was observed in one larva (12.5% of the infected larvae) (Figure 2B). The prevalence of microsporidia parasitizing larvae of S. pertinax was 0.51%.
Polydispyrenia spp. infections were characterized by the presence of at least 32 mononuclear spores contained within a persistent sporophorous vesicle, with the spores measuring 6.9 ± 1.0 x 5.0 ± 0.7µm (n = 23). Similarly, Amblyospora spp. were characterized by the presence of eight uninucleate spores contained within a sporophorous vesicle, with the spores measuring 4.5 x 3.5µm (n = 12).
The PCR products targeting the 16S region and electrophoresis agarose gel analysis confi rmed the presence of microsporidian DNA in 8 samples.
FIGURE 2 - Phase-contrast microscopy of smear slides of Simulium pertinax infected by microsporidia. Sporophorous vesicle of Polydispyrenia sp. containing 32 mononuclear spores (A). Octospores of Amblyospora spp. containing 8 uninucleate spores each (B).
A
B
[GenBank: AY090069] and Caudospora palustris [GenBank:
AF132544] reference sequences (with 100% bootstrapping). One sample (L2) was clustered with Amblyospora spp. [GenBank:
AJ252949] with 100% bootstrapping. The Edhazardia aedis
positive control (CONT+) taxon was clustered with Edhazardia aedis [GenBank: AF027684] with 100% bootstrapping.
DISCUSSION
Herein, we reported microsporidia parasitizing S. pertinax
larvae in theState of São Paulo, with a prevalence of 0.51%. Araújo-Coutinho6 previously reported a 0.5-2.0% prevalence of microsporidia in S. pertinax in State of Rio de Janeiro. Our study showed a similar prevalence to that previously reported by Crosskey19 in other populations of black fl ies, with rates
of up to 1%. Polydispyrenia spp. were the most prevalent parasitic species in S. pertinax from Caraguatatuba/SP in this study, while Amblyospora spp. showed a higher prevalence in Rio de Janeiro6. This difference could be explained by the small sample size, which prevented further analysis of the species population dynamics between S. pertinax from Rio de Janeiro and Caraguatatuba.
In this study, spores of the Polydispyrenia spp. measured 6.9 ± 1.0μm in length x 5.0 ± 0.7μm in width. Araújo-Coutinho6 reported spores of a similar size for a Polydispyrenia sp. from
S. pertinax that was ovocylindrical and measured 7.0 ± 0.6 x 4.9 ± 0.8μm. However, Castello-Branco and Andrade20 reported larger-sized spores measuring 8.3μm in length x 6.3μm in width for P. simulii from S. pertinax collectedin State ofSão Paulo,
Brazil. Sprague12 stated that the spore dimensions were 4.5 to 5.5μm x 2.5 to 3.5μm for P. simulii with the hosts listed as
S. pertinax and S. perfl avum from Brazil.
In this study, for Amblyospora spp. from Caraguatatuba, the spore measurement was 4.5μm in length x 3.5μm in width, similar to that found by Araújo-Coutinho6 for Amblyospora spp. infecting S. pertinax in the State ofRio de Janeiro. Both of these results were similar to those from Amblyosporabracteata and
Amblyospora varians, described in black fl ies in North America
and Europe21. According to Sprague12
, the morphological similarity between species of microsporidia, particularly the spore measurements, makes identifi cation diffi cult, and other methods are needed for identifi cation. Such evidence indicates that spore dimension diversity is too variable; thus, molecular analysis could help in species identifi cation.
Our sample, identifi ed morphologically as Polydispyrenia
spp., was grouped with the P. simulii and C. palustris clusters. This identifi cation corroborated previous results22-26 regarding the phylogeny of these parasites.
The genera Parathelohania, Hazardia, Marsoniella, Gurleya, Larssonia, Berwaldia, Varimorpha, Amblyospora
and the Amblyospora sp. from S. pertinax in this study form a separate group from the main Amblyospora cluster (Figure 3). Excluding the Varimorpha sp., which wascharacterized in an ant species, Solenopsis richteri (Forel, 1909), all genera in this group are parasites of aquatics hosts27-30.
Because the Amblyospora group is divided into two clades, corresponding to the hosts (Culex or Aedes/Ochlerotatus)28, the aquatic group also demonstrated distinct phylogenetic characteristics according to the host. The genera that infect both
Culex quinquefasciatus (SAY, 1823)and crustaceans (Hazardia,
Marsoniella, Gurleya, Larssonia and Berwaldia) are the main members of this clade. The genera that infect anopheline mosquitoes (Parathelohania), simulids (Amblyospora spp 3 in this study) and a species of ant (Varimorpha sp.), are more closely related to the aquatic group than to the main
Amblyospora group. The Amblyospora spp. in this study were clustered with Amblyospora sp. (AJ252949) from Simulium
spp. from the Paleartic29
,; confi rming the morphological and molecular similarities between these 2 species.
FIGURE 3 - Phylogenetic tree generated for microsporidia. Unrooted tree constructed with the maximum likelihood method using the general time reversible model of nucleotide substitution and gamma distribution (GTR + G), using Seaview software. The robustness of
the phylogenetic groups was evaluated using 1,000 bootstrap replicates, and bootstrap values greater than 70 were considered signifi cant.
and the main group of Amblyospora spp., which infects mosquitoes, indicating that these groups are not congeneric.
The differences between taxonomic relationships, based on phylogenetic placement and classical morphological characteristics, could probably be explained by the possibility that some of these characteristics (diplokaryon, sporophorous vesicles, and meiosis) appear to have multiple origins31. Thus, molecular analysis of other genomic regions could improve the phylogenetic understanding of microsporidia. This work
contributes to the phylogenetic analysis of microsporidia because it provides two genus sequences from these parasites.
The authors declare that there is no confl ict of interest.
FINANCIAL SUPPORT
Fundação de Amparo à Pesquisa do Estado de São Paulo
(FAPESP) 2012/23947-0
REFERENCES
1. Shelley A, Hernandez L, Maia-Herzog M, Luna Dias A, Garritano D. The blackfl ies (Diptera: Simuliidae) of Brazil. In: Arias JR, Golovatch S, Wantzen KM, Dominguez E, editors. Aquatic biodiversity in Latin America (ABLA). Vol. 6. Pensoft:Sofi a-Moscow; 2010. p. 821. 2. Coscarón S, Coscarón-Arias CL. Neotropical Simuliidae (Diptera:
Insecta). In: Adis J, Arias JR, Rueda-Delgado G, Wantzen KM, editors. Aquatic biodiversity in Latim America (ABLA). Vol. 3. Pensoft: Sofi a-Moscow; 2007. p. 685.
3. Ribeiro do Amaral-Calvão AM, Maia-Herzog M. Adolpho Lutz's collection of black fl ies (Diptera: Simuliidae), its history and importance. Hist Cienc Saude Manguinhos 2003; 10:259-271.
4. Vossbrinck CR, Maddox JB, Friedman S, Debrunner-Vossbrink BA, Woese CR. Ribosomal RNA sequence suggests microsporidia are extremely ancient eukaryotes. Nature 1987; 326: 411-414.
5. Ebersberger I, Simões RM, Kupczok A, Gube M, Kothe E, Voigt K, et al. A consistent phylogenetic backbone for the fungi. Mol Biol Evol 2012; 29:1319-1334.
6. Araújo-Coutinho CJ, Nascimento ES, Figueiro R, Becnel JJ. Seasonality and prevalence rates of microsporidia in Simulium pertinax (Diptera: Simuliidae) larvae in the region of Serra dos Órgãos, Rio de Janeiro, Brasil. J Invertebr Pathol 2004; 85:188-191.
7. Ginarte CA, Andrade CFS, Gaona JC. Larvas de simulídeos (Diptera, Simuliidae) do Centro Oeste, Sudeste e Sul do Brasil, parasitadas por microsporídeos (Protozoa) e mermitídeos (Nematoda). Ser Zool 2003; 93:325-334.
8. Castello-Branco A. Effects of Polydispyrenia simulii (Microspora; Duboscqiidae) on Development of the Gonads of Simulium pertinax (Diptera; Simuliidae). Mem Inst Oswaldo Cruz 1997; 94:421-424. 9. Online Archive of Instituto Brasileiro de Geografi a e Estatística
[Internet][cited 2013 May]. Available at: http://cod.ibge.gov.br/2335A. 10. Undeen A H, Vavra J. Research Methods for Entomopathogenic
Protozoa In: Lacey LA. Manual of Techniques in Insect Pathology. Chapter 4. 1st ed. Great Britain: Academic Press; 1997. p. 115-151. 11. Becnel J J. Preparations of Entomopathogens. In: Lacey LA. Manual of
Techniques in Insect Pathology.Chapter VIII-1. 1st ed. Great Britain: Academic Press; 1997; p. 337-353.
12. Sprague V, Becnel JJ, Hazard EI. Taxonomy of Phylum Microspora. Crit Rev Microbiol 1992; 18:285-395.
13. Adler PH, Becnel JJ, Moser B. Molecular charaterization and taxonomy of a new species of Caudosporidae (Microsporidia) from black fl ies (Diptera: Simuliidae), with host-derived relationships of the North American caudosporids. J Invertebr Pathol 2000; 75:133-143.
14. Vossbrinck CR, Andreadis TG, Vavra J, Debrunner-Vossbrink BA. Verifi cation of intermediate hosts in the life cycles of Microsporidia by small subunit rDNA sequencing. J Eukaryot Microbiol 1998; 45:290-292. 15. Huang X, Madan A. CAP3: A DNA sequence assembly program.
Genome Res 1999; 9:868-877.
16. Gouy M, Guindon S, Gascuel O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 2010; 27:221-224.
17. Waddell PJ, Steel MA. General time-reversible distances with unequal rates across sites: mixing gamma and inverse Gaussian distributions with invariant sites. Mol Phylogenet Evol 1997; 8:398-414.
18. Posada D, Crandall KA. MODELTEST: testing the model of DNA substitution. Bioinformatics 1998; 14:817-818.
19. Crosskey RW. The Natural History of Black fl ies. 1st ed. West Sussex, England: Wiley; 1990.
20. Castello-Branco A, Andrade CFS. Studies on Polydispyrenia simulii (Microspora; Pleistophoridae) in Simulium pertinax (Diptera; Simulidae) in Brazil. Mem Inst Oswaldo Cruz 1993; 88:167.
21. Vávra J, Undeen AH. Microsporidia (Microspora: Microsporida) from Newfoundland blakfl ies (Diptera: Simuliidae). Can J Zool 1981; 59:1431-1446.
22. Lom J, Nilsen F, Dykova I. Thelohania contejeani Henneguy, 1892: dimorphic life cycle and taxonomic affi nities, as indicated by ultrastrutural and molecular study. Parasitol Res 2001; 87:860-872. 23. Terry RS, Smith JE, Sharpe RG, Rigaud T, Littlewood TDJ, Ironside
JE, et al. Widespread vertical transmission and associated host sex-ratio distortion within eukaryotic Phylum Microspora. Proc R Soc Lond 2004; 271:1783-1789.
24. Vossbrinck CR, Debrunner-Vossbrinck BA. Molecular Phylogeny of the Microsporidia: ecological, ultrastructural and taxonomic considerations. Folia Parasitol 2005; 52:131-142.
25. Smith JE. The ecology and evolution of microsporidian parasites. Parasitol 2009; 136:1901-1914.
26. Dong S, Shen Z, Xu L, Zhu F. Sequence and phylogenetic analisys of SSU rRNA gene of fi ve Microsporidia. Curr Microbiol 2010; 60:30-37. 27. Baker MD, Vossbrinck CR, Becnel JJ, Andreadis TG. Phylogeny of
Amblyospora (Microsporida: Amblyosporidae) and related genera based on small subunit ribosomal DNA data: A possible example of host parasite cospeciation. J Invertebr Pathol 1998; 71:199-206. 28. Moser BA, Becnel JJ, Maruniak J, Patterson RS. Analysis of the
ribosomal DNA sequences of the microsporidia Thelohania and Vairimorpha of fi re ants. J Invertebr Pathol 1998; 72:154-159.
29. Cheney SA, Lafranchi-Tristem NJ, Canning EU. Phylogenetic relationships of Pleistophora-like microsporidia based on small subunit ribosomal DNA sequences and implications for the source of Trachipleistophora hominis infections. J Eukaryot Microbiol 2000; 47:280-287.
30. Refardt D, Canning EU, Mathis A, Cheney SA, Lafranchi-Tristem NJ, Ebert D. Small subunit ribosomal DNA phylogeny of microsporidia that infect Daphnia (Crustacea: Cladocera). Parasitol 2002; 124:381-389.
31. Baker MD, Vossbrinck CR, Didier ES, Maddox JV, Shadduck JA. Small subunit ribosomal DNA phylogeny of various microsporidia with emphasis on AIDS related forms. J Eukaryot Microbiol 1995; 42:564-570.
32. Vossbrinck CR, Andreadis TG, Vavra J, Becnel JJ. Molecular phylogeny and evolution of mosquito parasitic Microsporidia (Microsporidia: Amblyosporidae). J Eukaryot Microbiol 2004; 51:88-95.
33. Sokolova YY, Dolgikh VV, Morzhina EV, Nassonova ES, Issi IV, Terry RS, et al. Establishment of the new genus Paranosema based on the ultrastructure and molecular phylogeny of the type species Paranosema grylli Gen. Nov., Comb. Nov. (Sokolova, Selezniov, Dolgikh, Issi 1994), from the cricket Gryllus bimaculatus. Deg J Invertebr Pathol 2003; 84:159-172.
34. Muller A, Trammer T, Chioralia G, Seitz HM, Diehl V, Franzen C. Ribosomal RNA of Nosema algerae and phylogenetic relationship to other microsporidia. Parasitol Res 2000; 86:18-23.
35. Vossbrinck CR, Andreadis TG, Vavra J, Becnel JJ. Molecular phylogeny and evolution of mosquito parasitic Microsporidia (Microsporidia: Amblyosporidae). J Eukaryot Microbiol 2004; 51:88-95.
36. Nilsen F, Endresen C, Hordvik I. Molecular phylogeny of microsporidians with particular reference to species that infect the muscles of fi sh. J Eukaryot Microbiol 1998; 45:535-543.
37. Malone LA, Broadwell AH, Lindridge ET, McIvor CA, Ninham J. Ribosomal RNA genes of two microsporidia, Nosema apis and Vavraia oncoperae are very variable. J Invertebr Pathol 1994; 64:151-152. 38. Docker MF, Kent ML, Hervio DL, Khattra JS, Weiss LM, Cali A,
39. Silva AJ, Schwartz DA, Visvesvara GS, Moura H, Slemenda SB, Pieniazek NJ. Sensitive PCR diagnosis of Infections by Enterocytozoon bieneusi (microsporidia) using primers based on the region coding for small-subunit rRNA. J Clin Microbiol 1996; 34:986-987.
40. Zhu X, Wittner M, Tanowitz HB, Cali A, Weiss LM. Nucleotide sequence of the small ribosomal RNA of Encephalitozoon Cuniculi. Nucleic Acids Res 1993; 21:1315.
41. Visvesvara GS, Silva AJ, Croppo GP, Pieniazek NJ, Leitch GJ, Ferguson D, et al. In vitro culture and serologic and molecular identifi cation of Septata intestinalis isolated from urine of a patient with AIDS. J Clin Microbiol 1995; 33:930-936.
42. Visvesvara GS, Leitch GJ, Silva AJ, Croppo GP, Moura H, Wallace S, et al. Polyclonal and monoclonal antibody and PCR-amplifi ed small-subunit rRNA identifi cation of a microsporidian, Encephalitozoon hellem, isolated from an AIDS patient with disseminated infection. J Clin Microbiol 1994; 32:2760-2768.
43. Gatehouse HS, Malone LA. The ribosomal RNA gene region of Nosema apis (Microspora): DNA sequence for small and large subunit rRNA genes and evidence of a large tandem repeat unit size. J Invertebr Pathol 1998; 71:97-105.
44. Baker MD, Vossbrinck CR, Becnel JJ, Maddox JV. Phylogenetic position of Amblyospora Hazard & Oldacre (Microspora: Amblyosporidae) based on small subunit rRNA data and its implication for the evolution of the microsporidia. J Eukaryot Microbiol 1997; 44:220-225.
45. Nilse F, Che WJ. rDNA phylogeny of Intrapredatorus barri (Microsporida: Amblyosporidae) parasitic to Culex fuscanus Wiedemann (Diptera: Culicidae) Parasitology 2001; 122:617-623. 46. Andreadis TG, Vossbrinck CR. Life cycle, ultrastructure and molecular