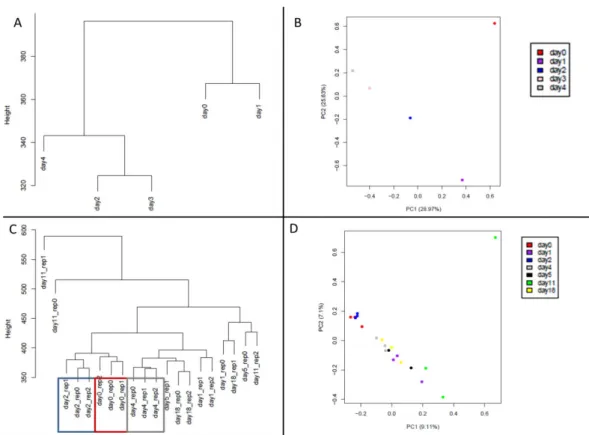
Assessing pseudogene expression during neural differentiation by RNA-Seq
Texto
Imagem
Documentos relacionados
To facilitate an analysis of the role of BRaf kinase in brain development, postnatal neural stem cell proliferation and neuronal differentiation, we created a conditional allele
From the two different developmental stages (myce- lium and fruiting body), we identified differentially expressed genes by transcriptome analysis and found that numerous
In the present study, we identified ubiquitous and tissue-specific [Na + ] i /[K + ] i -sensitive transcriptomes by comparative analysis of differentially expressed genes in
4 ) 48 out of 90 selected differentially expressed genes were confirmed by qRT-PCR, including genes implicated in energy metabolism, neuronal differentiation, signaling,
Through DGE analysis, we obtained 1,566 differentially expressed genes in SOF/NC, and 1,099 genes in LOF/NC, where the differentially expressed genes related to lipid metabolism
We analyzed the expression of hundreds of miRNAs predicted to be expressed during myogenic differentiation using a customized microarray and identified several known and
Using data from a microarray analysis of gene expression along a time course of differentiation (Figure S1A) [22], we analyzed the linear chromosomal distribution of co-regulated
We sought to identify patterns of gene expression in the MI during pod differentiation, determine biological processes associated with pod differentiation and determination,