Evaluation of gene selection metrics for tumor cell classification
Texto
Imagem


Documentos relacionados
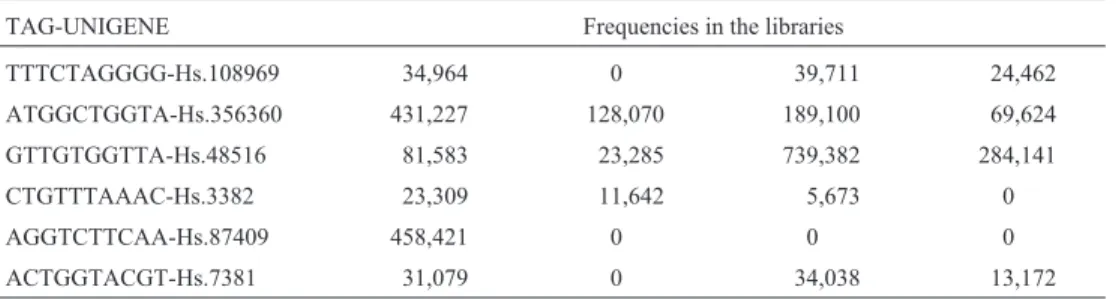
A set of genes related to secondary metabolism was extracted from the sugarcane expressed sequence tag (SUCEST) database and was used to investigate both the gene expression pattern
Primarily, the tumor suppressor gene expression was assigned to genes involved in the control of strategic points of the chain of events related to cell growth and
Therefore, both the additive variance and the deviations of dominance contribute to the estimated gains via selection indexes and for the gains expressed by the progenies
Comparison with gene expression-based metrics To determine whether a similar accuracy could have been obtained using expression data alone (i.e., without adding constraints based on
Results: Microarray based gene expression data and protein-protein interaction (PPI) databases were combined to construct the PPI networks of differentially expressed (DE) genes in
To characterize the expression profile of androgen receptor target genes in prostate cancer cells, we used expression microarray analysis on the PC346 cell line panel incubated with
In the present study, we used microarray-based methodology to assess the gene expression pro- file in the livers of mice infected with F. hepatica metacercariae, which was combined
Because of the increas- ing the speed of the decomposition of the anhydrite, together with the growth of the temperature of casting the bronze to the plaster mould, the gases