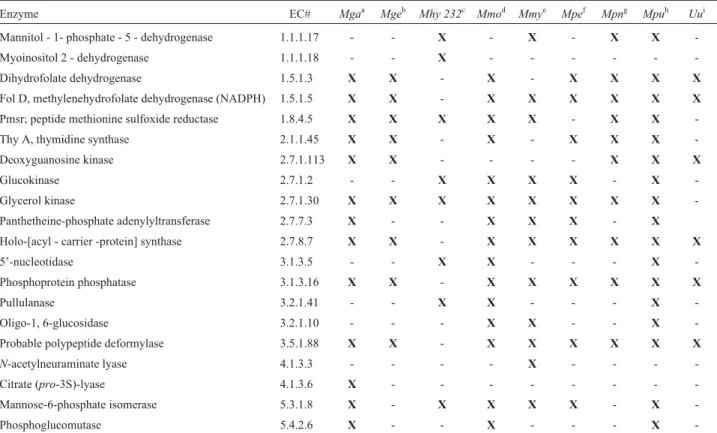
Differential metabolism of Mycoplasma species as revealed by their genomes
Texto
Imagem

Documentos relacionados
The complete genomes of a non-pathogenic (J) and pathogenic (7448) strain of Mycoplasma hyopneumoniae, as well as of a pathogenic (53) strain of Mycoplasma synoviae have been
This is a report on the analysis of genes involved in translation of the complete genomes of Mycoplasma hyopneumoniae strain J and 7448 and Mycoplasma synoviae.. In both genomes 31
Gene ontology (GO) term enrichment analysis of the genes with differential H3K4me3 peaks, revealed statistically significantly enriched GO terms only in the genes with
The amino acid consensus sequences corresponding to the PBP genes in the genomes and the clinical isolates were highly conserved.. The changes found in amino acid sequences
The differential expression analysis of RNA-seq data in control and Al stress revealed that more genes up-regulated in roots than leaves ( S7 Table and S8 Table), which suggested
Analysis of the predicted proteins of the three sequenced strains using a 95% amino acid similarity cutoff revealed that 69% of the predicted proteins of the three sequenced
Eighteen top Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were significantly involved in physiological responses, particularly in lipid metabolism, including
Analysis of RPR genes in various animal genomes revealed a striking divide in the animal kingdom that separates insects and crustaceans into a single group in which RPR genes