267
online | memorias.ioc.fiocruz.br Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 106(3): 267-273, May 2011
Tuberculosis (TB) infects a significant proportion of the world population and constitutes a major public health problem, particularly in developing regions. This disease has remained endemic for many decades due the human immunodeficiency virus (HIV) epidemic, the large num-ber of people trapped in poverty, and the disruption of the TB control program (WHO 2006). In this context, Brazil continues to be one of the 22 countries with the highest TB burden; altogether, these 22 countries account for 90% of all TB cases worldwide. However, as of 2007, a 26% decrease in TB incidence and a 32% decrease in TB mor-tality were verified for Brazil (SBPT 2009). In 2004, ap-proximately 80,000 TB cases were reported in Brazil with an incidence rate of 37.1/100,000 (WHO 2006), while, in the state of Minas Gerais (MG), 6,000 new cases were re-ported with an incidence of 45/100,000 (WHO 2006). Of these TB patients, 16.2% were coinfected with HIV and had a TB-related death rate of 7.3%. MG is one of the most populous states in Brazil and Belo Horizonte is its capital and the fifth largest city by number of residents in Brazil. In Belo Horizonte, the TB incidence was 37.1/100,000 in 2004 and it decreased to 31/100,000 in 2008. Consequent-ly, TB is far from being under control this region and more data are required to implement specific control measures.
In the above context, the application of molecular methods permitting both rapid Mycobacterium tubercu-losis diagnosis and genotyping (Kamerbeek et al. 1997, Kremer et al. 1999) is useful for understanding the pre-vailing TB genotypes and their circulation within the population. Spoligotyping is the second most widely used meth od for M. tuberculosis (Mtb)complex genotype af-ter IS6110-based fingerprinting (Prodinger 2007) that in combination with mycobacterial interspersed repetitive units-variable-number of tandem DNA repeats (MIRU-VNTR) has been used to replace typing via restric tion fragment length polymorphisms (RFLP) based on the insertion sequence IS6110. The IS6110 method has been considered the “gold standard” for genotyping Mtbsince 1993. However, the IS6110 method is an expensive, labori ous and lengthy methodology that requires weeks of Mtb culturing and specific software to analyse the RFLP band patterns, all of which make the interpretation and exchange of the data difficult. In addition, the IS6110
method has limited use for genotyping the Mtbisolates that contain only a few copies of IS6110 (Cave et al. 1991). Spoligotyping is based on a polymorphism in the chro-mosomal direct repeat locus (Kamerbeek et al. 1997)and has been applied to the characterization of the Mtb com-plex. Further, spoligotyping has gained increased inter-national acceptance as a rapid, first line and discrimina-tory test (Sola et al. 2001, Filliol et al. 2002, Brudey et al. 2006). The spoligotyping method was also the basis for the creation of the largest genotype database for M. tu-berculosis, containing a global distribution and phyloge-netic analysis for worldwide spoligotypes (Brudey et al. 2006). This database provides information that is useful for comparing the patterns found within different world regions, enhancing our understanding of disease distri-bution dynamics. Indeed, molecular epidemiological
ap-Financial support: FAPEMIG, CAPES
The SITVIT2 database project was partially financed by the Regional Council of Guadeloupe (CR/08-1612: Biodiversité et Risque Infectieux dans les modèles insulaires).
+ Corresponding author: silvanaspindola@gmail.com Received 17 August 2010
Accepted 6 April 2011
Spoligotyping of clinical
Mycobacterium tuberculosis
isolates
from the state of Minas Gerais, Brazil
Silvana Spíndola de Miranda/+, Wânia da Silva Carvalho, Philip Noel Suffys, Afrânio Lineu Kritski,
Martha Oliveira, Noemi Zarate, Thierry Zozio, Nalin Rastogi,Brigitte Gicquel
Departamento de Clínica Médica/Pneumologia, Faculdade de Medicina, Universidade Federal de Minas Gerais, Av. Alfredo Balena 190, 301030-100 Belo Horizonte, MG, Brasil
We performed spoligotyping on 114 strains of the Mycobacterium tuberculosis (Mtb) complex that had been isolated from patients in Minas Gerais Health Units during 2004. A total of 82/114 (72%) clinical isolates were clus-tered and 32/114 (28%) were unique. Seven shared types containing nine strains were newly created. A total of nine patterns corresponded to unreported orphan strains, as evaluated against all of the strains recorded in the SITVIT2 proprietary database in the Institut Pasteur de la Guadeloupe. The major clades were composed of isolates that belong to the following genotypes: Latin-America and Mediterranean (63/114, 55.3%) (the ill-defined T superfamily) (12/114, 10.5%), Haarlem (8/114, 7%), X clade (6/114, 5.3%), S clade (3/114, 2.6%) and the East-African Indian and Manu types, each with 1/114 (0.9%) isolates. A considerable number of strains (n = 20, 17.5%) showed patterns that did not fall within any of the previously described major clades. We conclude the bulk of tuberculosis (TB) (92/114, 80.7%) in our location is recent evolutionary strains that belong to the principal genetic groups 2/3. Further studies on epidemiology of TB are required to understand Mtbbiodiversity and TB transmission in this region.
TB genotyping in Minas Gerais, Brazil • Silvana Spíndola de Miranda et al. 268
proaches have shown that certain emerging Mtbstrains, which induce more severe forms of TB (such as Beijing), manifest higher failure/relapse than others (Glynn et al. 2002, Lan et al. 2003). These more severe forms may, in turn, accentuate the TB burden even in countries with good TB control programs (Vree et al. 2007). In addition, strains from the Beijing/W and Haarlem families of Mtb are emerging in certain global regions and are associated with drug resistance (EU 2006, Marais et al. 2006). Al-most no genotyping information is available for the Mtb strains circulating in the MG area in Brazil (Dalla Costa et al. 2009). Thus, we decided to characterise the Mtb complex clinical isolates from this state by spoligotyp-ing and compare the data with patterns found around the world to pinpoint the predominant Mtbclones.
PATIENTS, MATERIALS AND METHODS
Patients and bacterial isolates -We studied a total of 114 TB patients that presented with positive Mtbcultures at the Health Units in MG over a one year period (January-December 2004). We included basic demographic data via a standard questionnaire from the files provided at the hospital. The majority of the TB patients included in the study were residents in the cities of Belo Horizonte, Uber-lândia and Juiz de Fora (2nd and 3rd largest cities in MG) (MS 2003). Cultures were grown on Lowenstein-Jensen medium, followed by drug-susceptibility tests for rifam-picin (40 µg/mL), isoniazid (INH) (0.2 µg/mL), pyrazin-amide (100 µg/mL), streptomycin (4 µg/mL), ethambutol (2 µg/mL) and ethionamide (20 µg/mL). These experi-ments were performed using the proportion method at
Tree of spoligotype patterns Mycobacterium tuberculosis isolates (n = 57 profiles) by using the Phylogenetic Analysis Using Parsimony 4.0b software and the unweighted pair group method with arithmetic mean method. EAI: East-African-Indian; LAM: Latin-American-Mediterra-nean; SIT: Spoligotype International Type as defined in the international database of the Institute Pasteur de la Guadeloupe.
3 4 1491 2883 1983 17 20 2753 179 42 60 64 95 93 176 177 866 1906 2860 2884 33 211 orphan6 2548 orphan1
orphan7 1106 34 784 1166 1877
50 53 52 119 834
orphan2 451 2885 2025 157 1655
51 137 439 orphan8 602
2859 orphan4 orphan5
orphan9
orphan3 92 46 1410
1904 2882
Manu T S LAM EAI
Clade Apoligotype
SIT
269 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 106(3), May 2011
the TB Reference Laboratory (Ezequiel Dias Foundation) (Canetti et al. 1963). The study was approved by the local Institutional Review Board/Independent Ethics Commit-tee. Only the isolates that were positive in the cultures and susceptibility tests were entered in the study.
Spoligotyping and database comparison - A loop-ful of culture was suspended in tris-ethylenediamine tetraacetic acid (EDTA) (TE) buffer [10 mM Tris-HCl, 1 mM EDTA (pH 8.0)] and inactivated at 80ºC for 20 min. Nucleic acids were extracted using three con-secutive cycles of boiling for 3 min and cooling for 3 min, followed by centrifugation at 12,000 g for 5 min. The pellet was resuspended in the same TE buffer, stored at -20ºC and subjected to spoligotyping using a previously published protocol (Kamerbeek et al. 1997) and a com-mercially available kit (Isogen Life Science BV, Utrecht, The Netherlands). The hybridised spoligotypes in binary format were converted to an octal code for comparison with the SITVIT2 proprietary database from the Pasteur Institute of Guadeloupe, which is an updated version of the previously released SpolDB4 database (available from: www.pasteur-guadeloupe.fr:8081/SITVITDemo) (Brudey et al. 2006). At the time of the study herein, SIT-VIT2 contained genotyping information on 70,000 Mtb clinical isolates from 160 different countries of origin. In this database, a Spoligotype International Type (SIT) designation is given to isolates with spoligotypes shared by two or more patients, as opposed to “orphan” which designates patterns reported only in a single isolate. The major phylogenetic clades were assigned according to the signatures provided by SpolDB4, which defines 62 ge-netic lineages/sub-lineages (Brudey et al. 2006). These include specific signatures for various Mtbcomplex spe-cies and the rules defining the major lineages/sublineag-es for Mtb sensu stricto. These clades include Central-Asian (CAS), East-African-Indian (EAI), Haarlem (H), Latin-American and Mediterranean (LAM), Manu fam-ily,the S clade, the IS6110-low banding X clade and an
ill-defined T clade (defined “by default”). In parallel, a phylogenetic tree for all of the spoligotype patterns was constructed using the Phylogenetic Analysis Using Par-simony (PAUP) 4.0b software and the unweighted pair group method with arithmetic mean (UPGMA) method.
RESULTS AND DISCUSSION
TB remains a great public health concern in MG and in a country where TB is endemic, identifying the genotypes of the predominant strains is critical to study the disease transmission patterns and epidemiological features. The present study was performed on strains collected over one year and comprised a total of 114 patients, all fully characterised via spoligotyping. The results obtained are summarized in Figure and Tables I-III. Table I summarises the orphan patterns (n = 9). Table II shows seven newly-defined SITs for nine strains in this investigation. Table III describes 41 SITs contain-ing 96 isolates that matched a preexistcontain-ing shared type in the SITVIT2 database. Figure shows an UPGMA tree and illustrates the similarity between the 57 different patterns observed in this study.
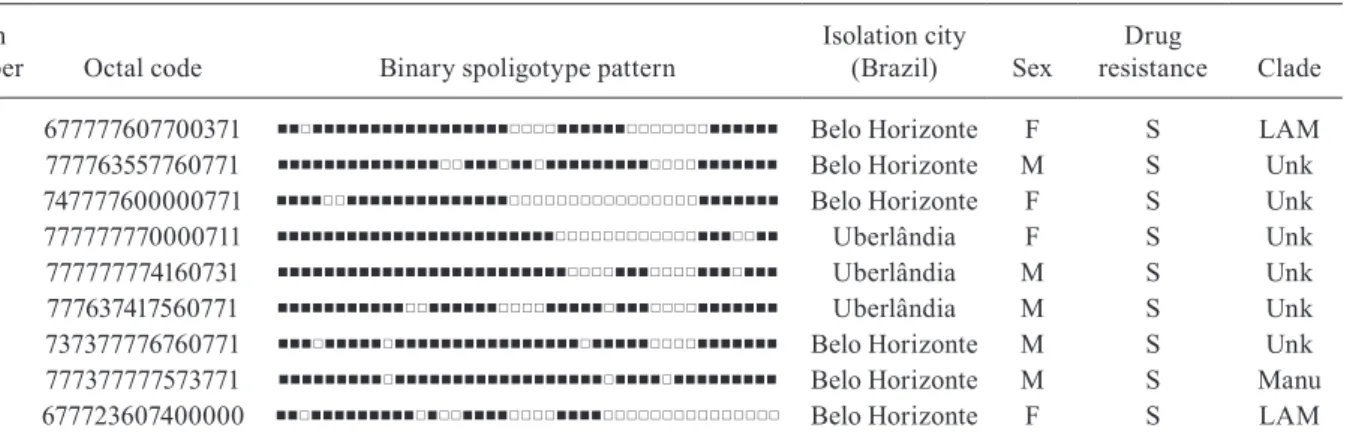
Regarding the orphan patterns (Table I), the major-ity of the patients were residents in the cities of Belo Horizonte and Uberlândia (the latter is a city with high incidence of TB in MG) (MS 2003). Fifty-six per cent of patients were male and the mean age varied between 45-59 years old, as observed in other studies in Brazil (Malaspina et al. 2008, Scholante Silva et al. 2009). All of the orphan patterns corresponded to pansusceptible strains. Note that the proportion of multidrug resistant (MDR) Mtbstrains in MG is low; between 2002-2008, less than 3.5% of all Brazilian MDR-TB cases con-cerned the residents of MG (MS 2007). Regarding the seven newly created shared types (Table II), two strains matched between them, one matched a Brazilian strain from a previous study, one matched a strain from Cuba, one matched a strain from Italy, three matched a strain from the United States of America and, finally, one
TABLE I
Description of the orphan strains (n = 9)
Strain
number Octal code Binary spoligotype pattern
Isolation city
(Brazil) Sex
Drug
resistance Clade
25 677777607700371 Belo Horizonte F S LAM
37 777763557760771 Belo Horizonte M S Unk
49 747777600000771 Belo Horizonte F S Unk
54 777777770000711 Uberlândia F S Unk
81 777777774160731 Uberlândia M S Unk
108 777637417560771 Uberlândia M S Unk
113 737377776760771 Belo Horizonte M S Unk
114 777377777573771 Belo Horizonte M S Manu
39a 677723607400000 Belo Horizonte F S LAM
TB g
en
ot
yp
ing i
n M
in
as G
era
is, B
ra
zil • S
ilv
an
a S
pí
nd
ola d
e M
ira
nd
a e
t a
l.
27
0
TABLE II
Description of seven newly-defined Spoligotype International Type (SIT) for nine strains
New SIT
Strain
number Octal code Binary spoligotype pattern
Match with a strain from
(patient origin)a Sex
Drug resistance
Isolation city
(Brazil) Clade
2859 110 777177600000000 BRA (NA) M S Belo Horizonte Unk
2885 93 777777477720771 CUB (NA) M S Uberlândia H
2882 40 677677606000171 ITA (NA) M INH Belo Horizonte LAM
2884 77 761777607760771 MTQ (NA) M S Belo Horizonte LAM
2753 18 677737607760571 This study (BRA) F S Juiz de Fora LAM
2753 105 677737607760571 This study (BRA) F S Juiz de Fora LAM
2860 33 757777607560771 USA (BRA) M S Belo Horizonte LAM
2860 95 757777607560771 USA (BRA) M S Belo Horizonte LAM
2883 6 774000017760771 USA (NA) F S Belo Horizonte Unk
a: the three letter country codes available from en.wikipedia.org/wiki/ISO_3166-1_alpha-3; H: Haarlem; INH: drug-resistance to isoniazid; LAM: Latin-American and Mediterranean; NA: information not available in database; S: pan-susceptible to first-line drugs; Unk: unknown patterns within any of the major clades described in database.
TABLE III
Description of 41 shared types containing 96 isolates that matched a preexisting shared type in the database
SIT Octal code Binary spoligotype pattern
Total in this study
(n)
Total in this study
(%)
Total in this study as compared to
database
(%) Clade
Worldwide distributiona (> 5% of total SIT)
3 000000007720771 1 0.9 1.19 H3-var USA, MEX, BRA, AUT
4 000000007760771 1 0.9 0.43 LAM USA, ZAF, ITA, TUR, BRA
17 677737607760771 3 2.6 0.56 LAM2 VEN, USA, BRA, ESP
20 677777607760771 5 4.4 0.80 LAM1 USA, BRA, NAM, PRT, VEN
33 776177607760771 5 4.4 0.61 LAM3 ZAF, USA, BRA, ESP, ARG, ITA
34 776377777760771 2 1.8 0.31 S USA, ZAF, ITA, CAN
42 777777607760771 19 16.7 0.93 LAM9 USA, BRA, ITA, VEN
46 777777770000000 2 1.8 1.56 Unk USA, CZE, IDN, FIN, IND, POL, AUT
50 777777777720771 6 5.3 0.26 H3 USA, AUT, ITA, ESP, CZE, BRA
51 777777777760700 3 2.6 1.49 Unk USA, ITA, AUT, MYS, HTI, BRA
52 777777777760731 1 0.9 0.18 T2 USA, BEL, FRA, ITA
27
1
Mem Inst Oswaldo Cruz
, Rio de Janeiro, V
ol.
106
(3), May 2011
SIT Octal code Binary spoligotype pattern
Total in this study
(n)
Total in this study
(%)
Total in this study as compared to
database
(%) Clade
Worldwide distributiona (> 5% of total SIT)
60 777777607760731 2 1.8 1.02 LAM4 ZAF, BRA, USA, VEN, GMB, ITA
64 777777607560771 3 2.6 1.17 LAM6 USA, BRA, GUF, PRT
92 700076777760771 1 0.9 0.30 X3 ZAF, USA, BRA
93 777737607760771 1 0.9 0.35 LAM5 VEN, USA, BRA, ITA, HTI
95 777777607560731 2 1.8 6.45 LAM BRA, USA, GUF, PRY
119 777776777760771 3 2.6 0.31 X1 USA, ZAF, MEX
137 777776777760601 1 0.9 0.10 X2 USA, ZAF, GBR
157 777737777760471 1 0.9 4.55 T BRA, PRT, ITA
176 777737607560771 1 0.9 3.33 LAM5- var USA, BRA, GUF
177 377777607760771 1 0.9 1.92 LAM BRA, ITA, PRY, USA, SAU
179 677737607760711 2 1.8 15.38 LAM2- var BRA, GUF, USA, PRT
211 776137607760771 2 1.8 2.99 LAM3- var USA, MEX, BRA, ESP
439 777376777760601 1 0.9 25.00 X2- var USA, PRY, BRA
451 777777477760771 3 2.6 2.38 T (H37Rv) USA, SAU, GBR
602 777777770000771 1 0.9 1.37 Unk GEO, TUR, BRA, USA, AUT, ZAF
784 776377777760731 1 0.9 2.00 S CAN, SAU, USA, BRA, ITA, ZAF, TUR
834 777767777740771 1 0.9 16.67 Unk FRA, BRA, IDN, PRT
866 577777607760771 1 0.9 2.86 LAM BRA, ITA, USA, FRA, ESP
1106 777777607760400 1 0.9 6.67 LAM PRT, BRA, PER, ITA
1166 777377777760771 1 0.9 3.13 T USA, BRA, ZAF, GRC, SAU, ESP, VEN, ITA
1410 777777700000000 1 0.9 6.67 Unk BRA, GMB, SAU, ESP, AUT, PRT, ITA, BGD, POL
1491 740000007760731 3 2.6 25.00 LAM BRA, MYS, TWN, USA, GBR
1655 777723777760771 1 0.9 3.33 Unk IDN, SAU, ITA, ZAF
1877 737377777760771 1 0.9 10.00 Unk SAU, IND, USA
1904 37676000000371 1 0.9 25.00 Unk ITA, VNM, AUT
1906 777777605560771 2 1.8 50.00 LAM BRA
1983 474000377413031 1 0.9 33.33 EAI3-IND BRA, ITA, IND
2025 777777777700371 1 0.9 20.00 Unk BRA, USA, JPN, ITA
2548 777777607600331 1 0.9 50.00 LAM BRA
TB genotyping in Minas Gerais, Brazil • Silvana Spíndola de Miranda et al. 272
matched a strain from Martinique (French West Indies). Lastly, for each of the preexisting shared types shown in Table III, the results summarise their binary and octal groupings, the total number of strains, their geographi-cal distribution, their genotypic lineage, as well as their percentage in this study compared with the same vari-able in the SITVIT2 database.
Irrespective of the database comparison, 77 (67.5%) of the clinical samples were clustered (20 clusters con-taining 2-19 isolates). Another 37 isolates (32.4%) had unique patterns, including nine orphan strains. In addi-tion, 105 out of 114 isolates from this study belonged to 48 shared types. Seven of these shared types contained nine strains and were newly created (Table II), whereas 41 contained 96 isolates. To discern the worldwide dis-tribution of the major shared types encountered in this study (Table III), the strains observed herein were com-pared with those classified as SIT in the database. SIT42 (LAM9 sub-lineage) was predominant in our setting, in 19/114 (16.7%) of the isolates. The next most prominent types were SIT50 (H lineage) and SIT53 (ill-defined T group of strains), with 6/114, or 5.3%, strains in each case. Next, we observed SIT33 and SIT20 (also of H lineage) in 5/114, or 4.4%, of the isolates in each case. These patients were also inhabitants of areas with a high TB prevalence. The majority of them were male and young adults; an important and productive age range in this country. Only a single case of drug-resistance from an INH-monoresis-tant strain was isolated in the city of Belo Horizonte.
The overall repartition of strains, according to ma-jor Mtb families (but not SITs stricto sensu), showed that major lineages in our setting ranked in the follow-ing order (Table III): LAM 63/114 or 55.3%, ill-defined T superfamily (defined by default) 12/114 or 10.5%, H 8/114 or 7%, X clade 6/114 or 5.3%, S clade 3/114 or 2.6% and EAI and Manu clades, 1/114 or 0.9% each. Previous studies analysing Mtb isolates from the state of Rio Grande do Sul in Southern Brazil reported the prevalence of LAM, unknown,He T, as described in our study; however, the authors analysed a lower number of isolates within a region of low TB prevalence (Noguti et al. 2010, Von Groll et al. 2010). Interestingly, no Beijing and CAS strains were found in the investigation herein, an observation that is corroborated by a recent report from Ritacco et al. 2008. A global perspective on the spoligotype binary patterns, related SITs, and genotypic lineages is illustrated in the Phylogenetic tree shown in Figure. The majority of LAM and T strains in our study originated in patients from Belo Horizonte. Approxi-mately 30% of these patients were either in hospitals or health care units, which could suggest an eventual link (such as nosocomial transmission). However, to support such a link, clonality of these isolates should be con-firmed using more discriminatory second-line typing methods, such as either IS6110-RFLP (IS6110-RFLP) or MIRU-VNTRs, which are most appropriate for defining recent transmission (Filliol et al. 2006). In a prospective study developed using Mtbisolates from a reference TB hospital in Brazil, T was the most frequent family; T was also the most frequent in our study at 10.5% (Malaspina et al. 2008, Cafrune et al. 2009).
In conclusion, we observed a high rate of cluster-ing (77/114, 67.5%; 20 clusters containcluster-ing 2-19 isolates). Further, the bulk of TB strains in our location (92/114, 80.7%) are recent evolutionary strains that belong to the principal genetic groups 2/3. Together, these data show that spoligotyping per se is not sufficient in our location for assessing the burden of recently transmitted TB. This methodology is known to overestimate the proportion of clustered cases (Filliol et al. 2006), hence, secondary typ-ing is necessary for such extended studies. Nonetheless, we have been able to provide an initial insight into the ge-netic relatedness of Mtbisolates within MG. In our study, given the small pool of analysed strains and that the study period was only one year, the rate of clustering (67.5%) was significantly high. Furthermore, the genotype for these strains suggests that the prevailing TB in Brazil might have been imported via European settlers during the conquest of the new world (Brudey et al. 2006). Last, but not least, a considerable number of strains (20/114 or 17.5%) showed patterns that did not belong to any of the previously described major clades, underlining that Bra-zil is a fertile land for studying Mtbgenotypic diversity. This type of study is important for trac ing relation-ships among the strains and it may also aid in interrupt-ing the chain of transmission between differ ent commu-nities and identifying the principal clades responsible for spreading the disease. Of course, additional studies us-ing more discriminatory secondary line typus-ing methods (IS6110-RFLP or MIRU-VNTRs), more isolates and a longer time period are critical if we are to fully under-stand the epidemiology of TB in this region.
REFERENCES
Brudey K, Driscoll JR, Rigouts L, Prodinger WM, Gori A, Al-Hajoj SA, Allix C, Aristimuño L, Arora J, Baumanis V, Binder L, Ca-frune P, Cataldi A, Cheong S, Diel R, Ellermeier C, Evans JT, Fauville-Dufaux M, Ferdinand S, Garcia de Viedma D, Garzel-li C, Gazzola L, Gomes HM, Guttierez MC, Hawkey PM, van Helden PD, Kadival GV, Kreiswirth BN, Kremer K, Kubin M, Kulkarni SP, Liens B, Lillebaek T, Ho ML, Martin C, Martin C, Mokrousov I, Narvskaïa O, Ngeow YF, Naumann L, Niemann S, Parwati I, Rahim Z, Rasolofo-Razanamparany V, Rasolonavalo-na T, Rossetti ML, Rüsch-Gerdes S, Sajduda A, Samper S, Shemyakin IG, Singh UB, Somoskovi A, Skuce RA, van Soolin-gen D, Streicher EM, Suffys PN, Tortoli E, Tracevska T, Vincent V, Victor TC, Warren RM, Yap SF, Zaman K, Portaels F, Rastogi N, Sola C 2006. Mycobacterium tuberculosis complex genetic di-versity: mining the fourth international spoligotyping database (SpolDB4) for classification, population genetics and epidemiol-ogy. BMC Microbiol 6: 23.
Cafrune PI, Possuelo LG, Ribeiro AW, Ribeiro MO, Unis G, Jarcze-wski CA, Rossetti ML, Zaha A 2009. Prospective study applying spoligotyping directly to DNA from sputum sample of patients of having tuberculosis. Can J Microbiol55: 895-900.
Canetti G, Rist N, Grosset J 1963. Measurement of sensitivity of the tuberculous bacillus to antibacillary drugs by the method of pro-portions. Methodology, resistance criteria, results and interpreta-tion. Rev Tuberc Pneumol (Paris) 27: 217-272.
273 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 106(3), May 2011
Dalla Costa ER, Ribeiro MO, Silva MS, Arnold LS, Rostirolla DC, Cafrune PI, Espinoza RC, Palaci M, Telles MA, Ritacco V, Suffys PN, Lopes ML, Campelo CL, Miranda SS, Kremer K, da Silva PE, Fonseca Lde S, Ho JL, Kritski AL, Rossetti ML 2009. Cor-relations of mutations in katG, oxyR-ahpC and inhA genes and in vitro susceptibility in Mycobacterium tuberculosis clinical strains segregated by spoligotype families from tuberculosis prevalent countries in South America. BMC Microbiology 19;9:39.
EU - European Union. European Concerted Action on New Generation Genetic Markers and Techniques for the Epidemiology and Con-trol of Tuberculosis 2006. Beijing⁄W genotype Mycobacterium tuberculosis and drug resistance. Emerg Infect Dis 12: 736-743.
Filliol I, Ferdinand S, Sola C, Thonnon J, Rastogi N 2002. Spoligo-typing and IS6110-RFLP typing of Mycobacterium tuberculosis
from French Guiana: a comparison of results with international databases underlines interregional transmission from neighbor-ing countries. Res Microbiol 153: 81-88.
Filliol I, Motiwala AS, Cavatore M, Qi W, Hazbon MH, Bobadilla del Valle M, Fyfe J, Garcia-Garcia L, Rastogi N, Sola C, Zozio T, Guerrero MI, Leon CI, Crabtree J, Angiuoli S, Eisenach KD, Durmaz R, Joloba ML, Rendon A, Sifuentes-Osornio J, Ponce de Leon A, Cave MD, Fleischmann R, Whittam TS, Alland D 2006. Global phylogeny of Mycobacterium tuberculosis based on single nucleotide polymorphism (SNP) analysis: insights into tubercu-losis evolution, phylogenetic accuracy of other DNA fingerprint-ing systems, and recommendations for a minimal standard SNP set. J Bacteriol 18: 759-772.
Glynn JR, Whiteley J, Bifani PJ, Kremer K, van Soolingen D 2002. Worldwide occurrence of Beijing/W strains of Mycobacterium tuberculosis: a systematic review. Emerg Infect Dis 8: 843-849.
Kamerbeek J, Schouls L, Kolk A, van Agterveld M, van Soolingen D, Kuijper S, Bunschoten A, Molhuizen H, Shaw R, Goyal M, van Embden J 1997. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology.
J Clin Microbiol 35: 907-914.
Kremer K, van Soolingen D, Frothingham R, Haas WH, Hermans PW, Martín C, Palittapongarnpim P, Plikaytis BB, Riley LW, Yakrus MA, Musser JM, van Embden JD 1999. Comparison of methods based on different molecular epidemiological markers for typing of Mycobacterium tuberculosis complex strains: in-terlaboratory study of discriminatory power and reproducibility.
J Clin Microbiol 37: 2607-2618.
Lan NT, Lien HT, Tung le B, Borgdorff MW, Kremer K, van Soolin-gen D 2003. Mycobacterium tuberculosis Beijing genotype and risk for treatment failure and relapse, Vietnam. Emerg Infect Dis 9: 1633-1635.
Malaspina AC, Cavalcanti HR, Leite CQ, Machado SM, Viana BH, Silva RM, Hage EF, Figueiredo WM, Marques E, Ferrazoli L, Arbex M, Lessi M, Fonseca LS, Rigouts L, Saad MH 2008. Use-fulness of Mycobacterium tuberculosis molecular typing in a tuberculosis low-endemic agro-industrial setting of Brazil. Jpn J Infect Dis 61: 231-233.
Marais BJ, Victor TC, Hesseling AC, Barnard M, Jordaan A, Brittle W, Reuter H, Beyers N, van Helden PD, Warren RM, Schaaf HS 2006. Beijing and Haarlem genotypes are overrepresented among children with drug-resistant tuberculosis in the Western Cape province of South Africa. J Clin Microbiol 44: 3539-3543.
MS - Ministério da Saúde 2003. Secretaria de Vigilância em Saúde. Departamento de Vigilância Epidemiológica. Coordenação Geral de Doenças Endêmicas. Área Técnica de Pneumologia Sanitária. Programa Nacional de Controle da Tuberculose [cited 2003 Nov 3]. Available from: www.opas.org.br/prevencao/site/UploadArq/ ProgramaTB.pdf.
MS - Ministério da Saúde 2007. Sistema de Vigilância em Saúde. Cen-tro de Referência Professor Hélio Fraga. Projeto MSH. Informe Técnico. Sistema de vigilância epidemiológica multirresistente Boletim 1, 1994-2006. Rev Bras Pneumol Sanit 15: 39-46.
Noguti EN, Leite CQF, Malaspina AC, Santos ACB, Hirata RDC, Hirata MH, Mamizuka EM, Cardoso RF 2010. Genotyping of
Mycobacterium tuberculosis isolates from a low-endemic set-ting in northwestern state of Paraná in Southern Brazil. Mem Inst Oswaldo Cruz 105:779-785.
Prodinger WM 2007. Molecular epidemiology of tuberculosis: toy or tool? A review of the literature and examples from Central Eu-rope. Wien Klin Wochenschr 119: 80-89.
Ritacco V, López B, Cafrune PI, Ferrazoli L, Suffys PN, Candia N, Vásquez L, Realpe T, Fernández J, Lima KV, Zurita J, Robledo J, Rossetti ML, Kritski AL, Telles MA, Palomino JC, Heersma H, van Soolingen D, Kremer K, Barrera L 2008. Mycobacterium tuberculosis strains of the Beijing genotype are rarely observed in tuberculosis patients in South America. Mem Inst Oswaldo Cruz 103:489-492.
SBPT - Sociedade Brasileira de Pneumologia e Tisiologia 2009. III Diretrizes para Tuberculose da Sociedade Brasileira de Pneumo-logia e TisioPneumo-logia. J Bras Pneumol 35: 1018-1048.
Scholante Silva AB, Von Groll A, Félix C, Conceição FR, Spies FS, Scaini CJ, Rossetti ML, Borsuk S, Dellagostin OA, Almeida da Silva PE 2009. Clonal diversity of M. tuberculosis isolated in a sea port city in Brazil. Tuberculosis (Edinb) 89: 443-447.
Sola C, Filliol I, Gutierrez MC, Mokrousov I, Vincent V, Rastogi N 2001. Spoligotype database of Mycobacterium tuberculosis: bio-geographic distribution of shared types and epidemiologic and phylogenetic perspectives. Emerg Infect Dis 7: 390-396.
Von Groll A, Martin A, Felix C, Prata PF, Honscha G, Portaels F, Vandame P, da Silva PE, Palomino JC 2010. Fitness study of the RDRio lineage and Latin American-Mediterranean family of
Mycobacterium tuberculosis in the city of Rio Grande, Brazil.
FEMS Immunol Med Microbiol58: 119-127.
Vree M, Bui DD, Dinh NS, Nguyen VC, Borgdorff MW, Cobelens FG 2007. Tuberculosis trends, Vietnam. Emerg Infect Dis 13: 796-797.