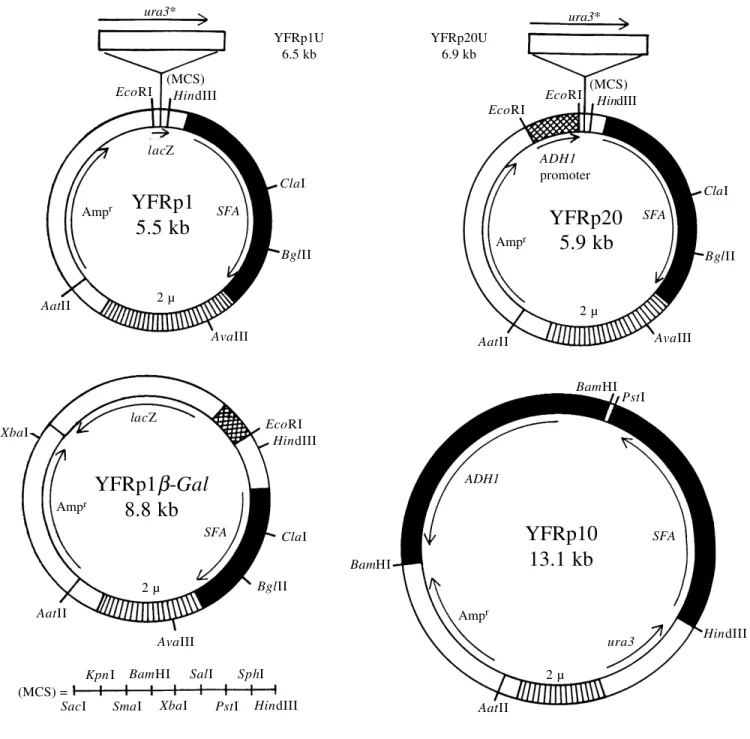
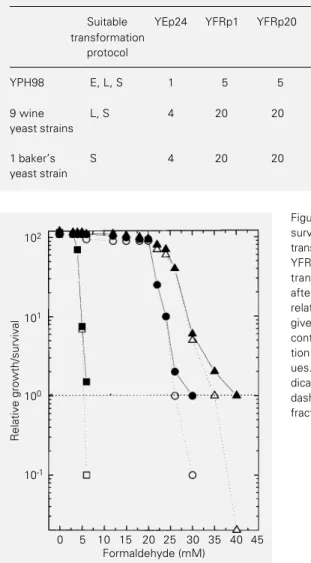
Genetic engineering of baker’s and wine yeasts using formaldehyde hyperresistance-mediating plasmids
Texto
Imagem
Documentos relacionados
Uma das explicações para a não utilização dos recursos do Fundo foi devido ao processo de reconstrução dos países europeus, e devido ao grande fluxo de capitais no
Neste trabalho o objetivo central foi a ampliação e adequação do procedimento e programa computacional baseado no programa comercial MSC.PATRAN, para a geração automática de modelos
Ousasse apontar algumas hipóteses para a solução desse problema público a partir do exposto dos autores usados como base para fundamentação teórica, da análise dos dados
Tendo como base teórica a Análise do Discurso de Linha Francesa, esta pesquisa pretendeu demonstrar que o discurso, no ciberespaço, assim como em toda e qualquer enunciação, não
[r]
Os modelos desenvolvidos por Kable & Jeffcry (19RO), Skilakakis (1981) c Milgroom & Fry (19RR), ('onfirmam o resultado obtido, visto que, quanto maior a cfiráda do
“A diferença de juízo entre Sócrates e Górgias sobre o poder da retórica mostra como, do ponto de vista da construção das personagens, elas possuem valores absolutamente
Para determinar o teor em água, a fonte emite neutrões, quer a partir da superfície do terreno (“transmissão indireta”), quer a partir do interior do mesmo