Carina Valente
Dissertation presented to obtain the Ph.D degree in Biology | Molecular Biology
Instituto de Tecnologia Química e Biológica António Xavier | Universidade Nova de Lisboa
Oeiras,
December 2015
.
First Edition, October 2015 Second Edition, January 2016 © Carina Valente
Cover image by Carina Valente ISBN: 978-989-206061
Financial support from Fundação para a Ciência e a Tecnologia,
Portugal through grant SFRH/BD/70058/2010 awarded to Carina
Dissertation presented on December 4, 2015, to obtain a Ph.D. degree in Biology | Molecular Biology by Instituto de Tecnologia Química e Biológica António Xavier, Universidade Nova de Lisboa.
Supervisors:
Raquel Sá-Leão Hermínia de Lencastre
Examiners:
Angela B. Brueggemann Krzysztof Trzciński
Acknowledgments/Agradecimentos
The work presented in this thesis was only possible due to the support of several
people and institutions, that I would like to acknowledge here.
My PhD supervisor, Raquel Sá-Leão, for all the guidance and advice. Thank you for all
the opportunities you gave me and for helping me grow, both scientifically and as a
person. You once asked me if there was anyone that I admired scientifically: you are
on that list. You are my role model as a supervisor.
Professor Hermínia de Lencastre, co-supervisor of my thesis, for the opportunity of
doing this PhD and for all the advice along the years. For her demand of excellence,
which motivates us all to do our best.
ITQB, for the excellent working conditions and Fundação para a Ciência e a
Tecnologia for financial support.
The researchers and institutions with whom we have collaborated and that have
accepted me for internships during this PhD. Dr. Francisco Pinto at Faculdade de
Cièncias da Universidade de Lisboa; the late Professor Kathrin Mühlemann and Dr.
Sylvio Brugger at the Institute of Infectious Diseases, University of Bern; Dr. Jason
Hinds and Dr. Katherine Gould at the Bacterial Microarray Group, St. Georges
University of London; and Dr. Suzanne Dawid at the Department of Pediatrics and
Communicable Diseases, University of Michigan.
My Thesis Committee, Dr. Isabel Gordo and Professor Manuel Carmo Gomes, for their
critical appreciation of my work and for interesting discussions.
My former and current colleagues at the Microbiology of Human Pathogens Unit, for all
My friends, for the joyful moments, for the support during the not-so-good moments
and for reminding me, when needed, that there is life outside the lab.
My family, my anchor. My idols and advisors in everything. The ones I reach for help
and to share good news. The ones who put up with my bad temper and are okay with
it. The ones I know I will always be able to count on, no matter what. This thesis is
dedicated to you. A minha família, minha âncora. Os meus ídolos e conselheiros em
“[…] searching for the causes of things is the best way to spend a life.”
Aristotle
Abstract
Streptococcus pneumoniae (or pneumococcus) is a frequent colonizer of the nasopharynx and an important cause of infectious diseases, with a high rate of
morbidity and mortality worldwide, particularly among young children, the elderly and
the immunocompromised. Despite this high burden of morbidity and mortality, invasive
pneumococcal disease is incidental. Nasopharyngeal colonization is the preferred
lifestyle for the pneumococcus, its prevalence being particularly high among young
children
Colonization by more than one S. pneumoniae strain, or co-colonization, is frequent and is important for pneumococcal biology because it promotes intra-specific
competition and evolution. Moreover, understanding co-colonization is critical to
monitor the impact of pneumococcal conjugate vaccines (PCV) and the extent of
serotype replacement by minor serotypes when vaccination targets the most abundant
ones. Despite its relevance and the fact that it has been recognized for several
decades, co-colonization remains poorly studied, mainly because high throughput
methods of multi-type detection have been developed only recently.
In this thesis we conducted five studies that aimed at understanding the dynamics of
pneumococcal co-colonization and how it is affected by pneumococcal conjugate
vaccines and by specific properties and mechanisms intrinsic to S. pneumoniae.
In the first study we used a molecular-based strategy to detect co-colonization in
nasopharyngeal samples from the pre- and vaccine eras, in order to evaluate the
impact of the introduction of the seven-valent pneumococcal conjugate vaccine (PCV7)
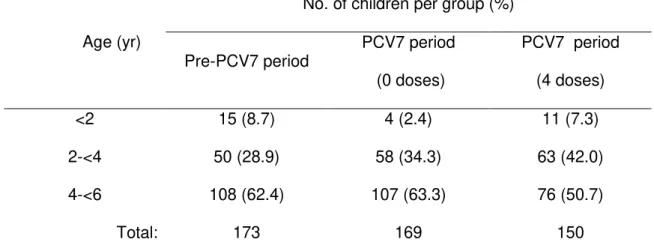
on pneumococcal colonization in Portuguese children. We showed that
co-colonization rates were significantly lower (p=0.004; Fisher’s exact test) among fully
vaccinated children (8.0%) than among children from the pre-PCV7 era (17.3%) or
vaccinated children there was an asymmetric distribution of non-vaccine types (NVT)
found in single and co-colonization events and we have attributed this asymmetry to
differences in the competitive abilities of serotypes. We proposed that some NVTs
prevalent in the PCV7 era are more competitive than others, impairing their
co-existence in the same niche. The results obtained in this study may have important
implications since a decrease in co-colonization events is expected to translate in
decreased opportunities for horizontal gene transfer, hindering pneumococcal evolution
events, and might represent a novel potential benefit of conjugate vaccines.
The replacement of PCV7 by PCV13 (13-valent PCV) has raised the question on
whether this vaccine would have a similar effect on co-colonization. To address this
question, in the second study we used a similar approach to the one used in the study
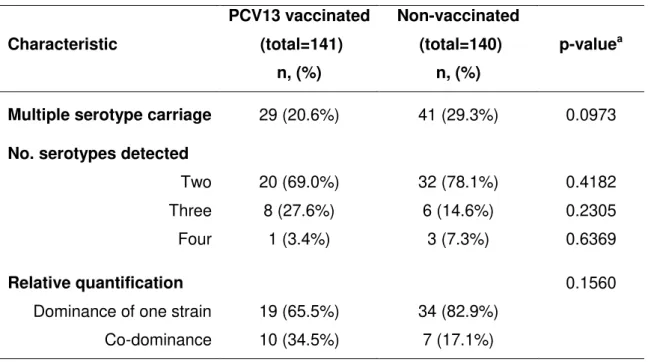
conducted in the PCV7 era. We showed lower co-colonization prevalence in PCV13
fully vaccinated children, compared to children who had not received any PCV (20.6%
vs. 29.3%), an observation that supports the results obtained in the first study. As
before, we reported an asymmetric redistribution of NVTs in single and co-colonization
events in the vaccinated group, which supported our hypothesis of different competitive
abilities among NVT serotypes. Moreover, we showed that PCV13 serotypes are still
highly prevalent in a population with no universal but very high vaccine coverage,
mainly among non-vaccinated children. In vaccinated children PCV13 serotypes were
found mainly as minor serotypes, while in the non-vaccinated group these serotypes
were highly prevalent in single carriage events and were present in high relative
abundance in most co-colonization events. Overall, the results obtained in this study
regarding the impact of PCV13 on co-colonization corroborated the potential impact of
PCVs on pneumococcal ecology by decreasing co-colonization events and, thus,
decreasing opportunities for horizontal gene transfer.
In the third study we used knowledge previously obtained through the use of highly
associated with the use of colony morphology for detection of co-colonization, a
convenient approach that has been the basis of most reports on co-colonization in
surveillance studies that do not systematically look for co-colonization. Comparison of
the serotype distribution of co-colonized samples detected through both methodologies
revealed that detection based on colony morphology resulted in a 20%
overrepresentation of serotypes that display very distinctive colony morphologies, such
as serotype 3 and non-encapsulated pneumococci, which did not reflect the real
epidemiology of these serotypes on the pneumococcal population.
In the fourth and fifth studies we explored how co-colonization might be affected by
specific properties and mechanisms intrinsic to S. pneumoniae biology.
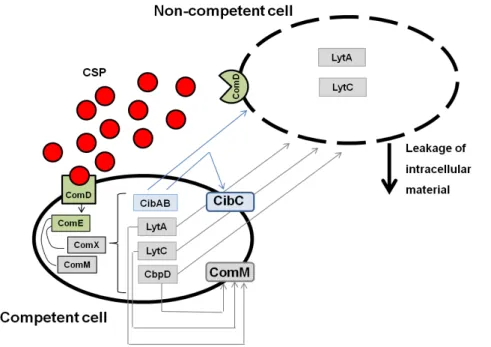
In the fourth study we determined the impact of pherotype-mediated competition (or
competence stimulating peptide, CSP) on pneumococcal co-colonization using a
collection of co-colonized samples containing two strains of different serotypes. CSP
assignment of all pneumococci revealed that in 52.5% of the samples both strains were
of the same pherotype, while in 47.5% co-colonizing strains were of a different
pherotype. Comparison of the observed proportions of concordant and discordant
pherotypes with the ones estimated (53.8% and 46.2%, respectively), revealed no
significant differences. The results obtained in this study supported the hypothesis that
there is a limited role of pherotype-mediated competition on co-colonization.
In the fifth study we explored the impact of the blp (bacteriocin-like peptide) locus and
bacteriocin secretion on co-colonization. We characterized the blp locus of a highly diverse collection of co-colonizing strains and determined their inhibitory activity
through competition overlay assays. For comparison, we performed the same
characterization in a collection of pneumococci isolated from single carriage events.
We showed that co-colonizing pneumococci present high genetic diversity at the level
showed also that pneumococcal strains co-colonize individuals independently of the
genetic content of their blp-locus and independently of their phenotypes of bacteriocin
secretion, and that phenotypes of bacteriocin secretion are the same in strains isolated
from single and co-colonization events. Overall, the results obtained in this study
support the hypothesis that there is a limited role of the blp-locus and bacteriocin secretion on co-colonization.
In conclusion, the results obtained in this thesis improved the scarce knowledge on
pneumococcal co-colonization through the use of innovative methodologies and by
addressing important questions such as the impact of PCVs and bacterial competition
Resumo
A bactéria Streptococcus pneumoniae (pneumococo) coloniza assintomatica e frequentemente a nasofaringe e é também uma importante causa de doença
infecciosa, com elevadas taxas de mortalidade e morbilidade, sobretudo em crianças,
idosos e indivíduos imuno-comprometidos. A colonização da nasofaringe é, no
entanto, preferencial no ciclo de vida do pneumococo, sendo particularmente elevada
em crianças até aos seis anos.
A colonização da nasofaringe por mais do que uma estirpe de S. pneumoniae, ou
co-colonização, é frequente. O estudo da co-colonização é importante dada a sua
relevância para a biologia do pneumococo, por promover interacções intra-específicas
e, consequentemente, a evolução da espécie. Além disso, compreender a
co-colonização é fundamental para a monitorização do impacto das vacinas
pneumocócicas conjugadas (PCV) e do grau de substituição de serótipos por serótipos
minoritários, num cenário em que a vacina é dirigida contra os serótipos mais
abundantes. Apesar de a co-colonização ser reconhecidamente importante e um
fenómeno identificado há já várias décadas, continua pouco estudada, sobretudo
devido ao facto de apenas recentemente terem sido desenvolvidos métodos
adequados à sua detecção.
No âmbito desta tese foram realizados cinco estudos com o objectivo de entender a
dinâmica da co-colonização por pneumococos e qual o impacto das PCVs e de certas
propriedades e mecanismos intrínsecos à biologia desta bactéria neste fenómeno.
Com o objectivo de avaliar o impacto da introdução da vacina pneumocócica
conjugada sete-valente (PCV7) na co-colonização por pneumococos em crianças
portuguesas, realizámos o primeiro estudo, recorrendo a métodos moleculares para
detectar co-colonização em amostras da nasofaringe de crianças obtidas antes e após
completa apresentavam uma prevalência de co-colonização significativamente inferior
(8.0%) à de crianças não vacinadas no mesmo período (18.3%) ou no período
pré-vacinal (17.3%). Nas crianças vacinadas observou-se uma distribuição assimétrica dos
serótipos não vacinais (NVT) nos eventos de co-colonização e colonização simples
(i.e., colonização por uma só estirpe), assimetria essa que atribuímos a diferenças nas
capacidades competitivas dos serótipos. Esta observação levou à proposta de que
alguns NVT prevalentes na era vacinal são muito competitivos, impedindo a sua
co-existência com outros serótipos no mesmo nicho. Os resultados deste estudo poderão
ter implicações relevantes dado que uma diminuição nos eventos de co-colonização
poderá traduzir-se numa diminuição das oportunidades para a ocorrência de
transferência horizontal de genes e, consequentemente, dos eventos de evolução,
representando potencialmente um novo benefício das vacinas conjugadas.
A substituição da PCV7 pela PCV13 (PCV treze-valente) levou-nos a questionar se
esta vacina teria um efeito semelhante ao da PCV7 na co-colonização. Deste modo,
no segundo estudo adoptámos uma abordagem semelhante à do estudo da era da
PCV7. Demonstrámos que a prevalência de co-colonização em crianças com
vacinação completa (20.6%) era inferior à de crianças não vacinadas do mesmo
período (29.3%). Em concordância com o estudo anterior, observámos uma
distribuição assimétrica de NVT em eventos de co-colonização e colonização simples,
facto que suporta a hipótese previamente proposta de diferenças nas capacidades
competitivas dos serótipos. Demonstrámos ainda que os serótipos incluídos na PCV13
são ainda prevalentes na população. Nas crianças vacinadas os serótipos incluídos na
PCV13 foram encontrados como um serótipo minoritário na maioria das ocorrências,
enquanto que no grupo de não-vacinados estes serótipos eram muito prevalentes em
eventos de colonização simples ou como serótipo maioritário em eventos de
co-colonização. Os resultados apresentados neste estudo relativos ao impacto da PCV13
pneumococo pela diminuição dos eventos de co-colonização e das oportunidades para
transferência horizontal de genes.
No terceiro estudo usámos informação obtida previamente através do uso de métodos
moleculares na detecção de co-colonização para quantificar o viés associado à
detecção de co-colonização com base na morfologia das colónias de pneumococos.
Dada a sua conveniência, esta abordagem baseada na morfologia constitui a base da
maioria dos dados reportados na literatura relativos à prevalência de co-colonização. A
comparação da distribuição de serótipos em amostras co-colonizadas com detecção
efectuada através de ambas as metodologias demonstrou que a detecção com base
na morfologia das colónias aumenta em 20% a prevalência de serótipos que
apresentam morfologias mais facilmente distinguíveis, como é o caso do serótipo 3 e
dos pneumococos não-encapsulados, podendo alterar artificialmente a prevalência
estimada destes serótipos na população.
No quarto e quinto estudos, explorámos o impacto de propriedades e mecanismos
intrínsecos à biologia do pneumococo na co-colonização.
No quarto estudo avaliámos o impacto da competição mediada pelo ferótipo (ou
péptido indutor da competência) na co-colonização, recorrendo a uma colecção de
amostras co-colonizadas por duas estirpes de pneumococos. A determinação do
ferótipo para todas as estirpes revelou que 52.5% das amostras continham estirpes
com o mesmo ferótipo, enquanto que nas restantes as estirpes eram de ferótipos
diferentes. A comparação das proporções de amostras com ferótipos concordantes
(52.5%) e discordantes (47.5%) com as proporções estimadas (53.8% e 46.2%,
respectivamente) não revelou diferenças significativas. Estes resultados sugerem que
a competição mediada pelo ferótipo tem um impacto limitado na co-colonização.
na co-colonização. Caracterizámos o locus de uma colecção muito diversa de
pneumococos identificados em eventos de co-colonização e determinámos a sua
capacidade inibitória através de ensaios de competição em meio sólido. Utilizámos
como controlo uma colecção de pneumococos isolados de eventos de colonização
simples. Demonstrámos que os pneumococos a co-colonizar apresentam uma elevada
diversidade genética ao nível do locus blp e exibem diferentes fenótipos de secreção
de bacteriocinas e de imunidade. Adicionalmente, reportámos que os pneumococos
co-colonizam um hospedeiro independentemente dos fenótipos de secreção de
bacteriocinas e que os mesmos não são diferentes em estirpes de co-colonização e de
colonização simples. Os resultados obtidos neste estudo sugerem que o impacto do
locus blp e da secreção de bacteriocinas na co-colonização é limitado.
Em suma, os resultados apresentados nesta tese aumentaram o conhecimento sobre
a co-colonização por pneumococos através do uso de metodologias inovadoras e da
resposta a questões importantes, tais como o impacto das PCV e dos mecanismos de
competição neste fenómeno, e a forma como os mesmos podem influenciar a ecologia
Thesis outline
The purpose of this thesis was to gain insights on the dynamics of pneumococcal
co-colonization and how it is affected by pneumococcal conjugate vaccines and by
specific properties and mechanisms intrinsic to S. pneumoniae biology.
Chapter I is a general introduction where important aspects of S. pneumoniae epidemiology and biology, relevant for the scope of the thesis, are presented. Among
these, are pneumococcal epidemiology, effects of anti-pneumococcal vaccination,
importance of co-colonization, methods for detection of co-colonization, and bacterial
properties and competition mechanisms that might impact on co-colonization.
Chapter II describes the assessment of co-colonization in a collection of
nasopharyngeal samples obtained from vaccinated and non-vaccinated children,
encompassing the availability of PCV7 in Portugal, aiming at determining the impact of
this vaccine on co-colonization. The analysis includes the combination of two sensitive
molecular methods for detection of co-colonization and comparison of co-colonization
patterns among children sampled in the pre-PCV7 era, and vaccinated and
non-vaccinated children sampled in the PCV7 era.
Chapter III describes a study conducted to evaluate the impact of PCV13 on
co-colonization. Samples obtained from PCV13 vaccinated and non-vaccinated children
were analyzed with a capsular microarray to detect co-colonization and the two groups
were compared regarding co-colonization patterns.
Chapter IV describes a study conducted to assess the bias in the serotype distribution
of pneumococci associated with co-colonization detection based on colony
distribution of co-colonized samples detected by molecular methods and by colony
morphology.
Chapter V describes the impact of pherotype-mediated fratricide on co-colonization.
The analysis included pherotype assignment of a co-colonized collection of
nasopharyngeal samples and estimation of the proportion of samples co-colonized with
strains of the same or of a different pherotype for comparison with the observed
proportions.
Chapter VI describes the impact of the blp (bacteriocin-like peptide) locus and bacteriocin secretion on pneumococcal co-colonization. The analyses included genetic
characterization of the blp locus of co-colonizing strains and assessment of the phenotype of bacteriocin secretion through overlay assays. Comparison with a control
collection of strains isolated from single carriage events was also performed.
Chapter VII presents general conclusions of the studies conducted in this thesis and
enumerates several questions that remain unanswered and could be the focus of future
research.
Chapters II, IV and V are reproductions of the following publications and can be read
independently:
Chapter II - Valente, C., J. Hinds, F. Pinto, S. D. Brugger, K. A. Gould, K.
Mühlemann, H. de Lencastre, R. Sá-Leão. 2012. Decrease in pneumococcal
co-colonization following vaccination with the seven-valent pneumococcal conjugate
Chapter IV - Valente, C., H. de Lencastre, R. Sá-Leão. 2013. Selection of distinctive
colony morphologies for detection of multiple carriage of Streptococcus pneumoniae. Pediatr. Infect. Dis. J.; 32(6):703-4. doi: 10.1097/INF.0b013e31828692be.
Chapter V - Valente, C., H. de Lencastre, R. Sá-Leão. 2012. Pherotypes of
co-colonizing pneumococci among Portuguese children. Microbe Drug Resist;
18(6):550-4. doi: 10.1089/mdr.2011.0228.
Chapter III has been submitted for publication and Chapter VI is nearly ready for
Table of contents
Acknowledgments/Agradecimentos... v
Abtract ... ix
Resumo ... xii
Thesis outline ... xvi
Table of contents ... xvii
Chapter I ... 1
Introduction
Streptococcus pneumoniae, on the vanguard of scientific discoveries ... 3 Streptococcus pneumoniae ... 4
Epidemiology of S. pneumoniae ... 5
Pneumococcal colonization ... 5
Pneumococcal disease ... 7
Non-susceptibility to antimicrobial agents ... 9
Anti-pneumococcal vaccination ... 11
Effect of pneumococcal vaccination ... 12
Pneumococcal co-colonization... 14
Biological significance ... 14
Epidemiology ... 16
The quest for the perfect detection method ... 17
Co-colonization determinants... 21
The contributions of the capsule and genetic background to (co)colonization .... 22
Chemical war I – competence-mediated fratricide ... 25
Chemical war II – bacteriocin production ... 29
The blp-bacteriocins ... 30
Aim of the thesis ... 35
Chapter II ... 51
Decrease in pneumococcal co-colonization following vaccination with the seven-valent
pneumococcal conjugate vaccine
Summary ... 53
Introduction ... 54
Materials and methods... 55
Results ... 59
Discussion ... 65
Acknowledgments ... 70
Funding ... 70
References ... 70
Supplementary material ... 73
Chapter III ... 75
Impact of the 13-valent pneumococcal conjugate vaccine on Streptococcus pneumoniae multiple serotype carriage
Summary ... 77
Introduction ... 78
Materials and methods... 79
Results ... 81
Discussion ... 87
Funding ... 91
References ... 91
Supplementary material ... 94
Chapter IV ... 95
Selection of distinctive colony morphologies for detection of multiple carriage of
Streptococcus pneumoniae
References ... 99
Chapter V ... 101
Pherotypes of co-colonizing pneumococci among Portuguese children
Summary ... 103
Introduction ... 103
Materials and methods... 105
Results ... 106
Discussion ... 109
Acknowledgments ... 112
Author disclosure statement ... 112
References ... 112
Chapter VI ... 115
Characterization of the blp locus of colonizing pneumococci and its impact on
co-colonization
Summary ... 117
Introduction ... 118
Materials and methods... 121
Results ... 128
Discussion ... 138
Funding ... 142
References ... 142
Supplementary material ... 146
Chapter VII ... 149
Concluding remarks
Chapter I
Streptococcus pneumoniae, on the vanguard of scientific
discoveries
Streptococcus pneumoniae, or pneumococcus, is an important human pathogen, continuing to kill thousands of people and being still “the captain of the men of death”,
as William Osler called it in 1918 (Osler, 1901).
The first reports about “elongated diplococci” in infected lungs are from Edwin Klebs
and date back to 1875. It was only in 1881, however, that this bacterium was first
isolated and its pathogenic potential was established, by both Louis Pasteur and
George Sternberg (reviewed in (Watson et al., 1993)).
Several designations were attributed to the pneumococcus along the years (reviewed
in (Watson et al., 1993)) taking into consideration the shape and type of disease
caused by the bacteria but, in 1974, the name Streptococcus pneumoniae was adopted
based on its morphology in chains when grown in liquid medium (Bergey, 1974).
Due to its important role as a human pathogen, S. pneumoniae is one of the most well
studied bacteria. For this reason, it is not surprising that it has been center stage to
important medical and scientific discoveries. In the field of medicine, the
pneumococcus was used in the development of Gram’s stain that allowed the
distinction between Gram positive and Gram negative pathogenic bacteria (reviewed in
(Watson et al., 1993)), the discovery of the ability of polysaccharides to induce antibody
production and their use as antigens in vaccines (Avery, 1917; Heidelberger & Avery,
1923), and the confirmation of the therapeutic efficacy of penicillin (Tillett et al., 1944).
In science, S. pneumoniae was the leading character in the discovery of the “Transforming Principle” and the mechanism of bacterial gene transfer (Griffith, 1928),
in the identification of DNA as the genetic material (Avery et al., 1944), in the discovery
of the protective role of the capsule against host phagocytic cells (Felton et al., 1955),
all of which have contributed to even greater discoveries and, as of today, are still
being used in the most various fields and applications.
Over 130 years have passed since the discovery of the pneumococcus and science
and medicine are still far from winning the battle against this highly successful
pathogen. Recent estimates on the number of serious cases of pneumococcal disease
reach over 14 million, from which over 0.8 million cases result in the death of children
below five years of age (O'Brien et al., 2009) .
Streptococcus pneumoniae
S. pneumoniae is a Gram positive bacterium with the form of an elongated coccus (lancet-shaped). It varies between 0.5 and 1.5 µm in diameter and can be found
isolated in single cells, in the form of diplococci, and in longer chains. Pneumococci are
facultative anaerobic, display α-hemolytic activity when grown on blood-supplemented
plates and are typically optochin susceptible and soluble in bile salts. It is a fastidious
organism, its growth being favored in media containing blood, at 37ºC, and in a
CO2-supplemented atmosphere (CDC, 2012; Sneath, 1986).
The genome of S. pneumoniae is a covalently closed circular DNA molecule of ~2M base pairs, varying, in size and content, from strain to strain. It contains over 1500
genes that are essential for cell viability and a variable number of genes essential for
virulence or to maintain a non-invasive phenotype (Bijlsma et al., 2007; Hava & Camilli,
2002; Lanie et al., 2007; Obert et al., 2006; Orihuela et al., 2004). S. pneumoniae possesses a highly plastic genome, with several recombination hotspots, which
constitutes an evolutive advantage and makes it one of the most well adapted
2007; Obert et al., 2006). In fact, analysis of 17 sequenced genomes showed that only
46% of the homologous gene clusters were common between strains (Hiller et al.,
2007).
Due to all the diversity in pneumococcal population and to the need to classify and
cluster pneumococcal strains in the context of epidemiological studies, a gold-standard
fingerprinting method, MLST (multi-locus sequence typing) is generally used as a
typing method, in which a sequence type (ST) is assigned to a strain based on the
combination of the allelic variants of seven housekeeping genes – aroE, gdh, gki, recP,
spi, xpt, and ddl (Enright & Spratt, 1998) (http://www.mlst.net/).
One of most distinctive characteristics of the pneumococcus is its polysaccharide
capsule, which led Avery to name it “the sugar-coated microbe” (Bardossi, 1988). This
capsule has antigenic properties which can be used as an identification method,
through the Quellung reaction, by the use of specific antisera (Sorensen, 1993). The
capsule is the main pneumococcal virulence factor and has several distinct functions
that will be discussed later in this chapter. Based on different antigenic properties
conferred by biochemical and structural differences of the polysaccharide capsule, over
95 capsular types, or serotypes, have been described up to now (Calix & Nahm, 2010;
Jin et al., 2009; Oliver et al., 2013; Park et al., 2007; Park et al., 2015).
Epidemiology of S. pneumoniae
Pneumococcal colonization
S. pneumoniae is a frequent inhabitant of the human nasopharynx, co-existing commensally with the human host. Colonization can occur soon after birth and remains
high in the first three years of life, decreasing until the age of ten and remaining low
pneumococcus at least once in life and each serotype can colonize for several weeks
or months, being then replaced by another serotype or reacquired (Gray et al., 1980;
Sá-Leão et al., 2008).
Carriage is usually asymptomatic and its duration varies according to the serotype of
the strain, the age of the child and the immunological status, as it allows the individual
to acquire B cell mediated immunity against the carried serotype. A low mucosal
immunity will result in persistent and repeated colonization events, while a stronger
local response will accelerate clearance and prevent re-colonization (Bogaert et al.,
2004a; Weinberger et al., 2008).
Asymptomatic carriers constitute a reservoir of pneumococci in the community and are
important vehicles of transmission, which can occur via direct contact between
individuals, by aerosols, or by contact with contaminated abiotic surfaces (Musher,
2003; Walsh & Camilli, 2011).
Risk factors for pneumococcal carriage include young age (up to 2 years of age),
regular contact with young children, crowding (as occurs in day care centers, prisons,
hospitals, and military camps), previous respiratory disease, both infectious and
chronic, and cigarette smoking (Abdullahi et al., 2012; Almeida et al., 2014; Bogaert et
al., 2004a; Gray et al., 1980; Labout et al., 2008).
Several studies have been conducted worldwide on the subject of pneumococcal
carriage (Adegbola et al., 2014; Cohen et al., 2012; Daana et al., 2015; Vestrheim et
al., 2010) (Nunes et al., submitted), most of them focused on young children attending day care centers, as this is the age group where colonization is more frequent and
crowding is high in day care centers, favoring transmission (Bogaert et al., 2004a;
Sá-Leão et al., 2008). These studies have shown that the carriage rates and the
distribution of the carried serotypes vary across geographical locations, reaching
(Vestrheim et al., 2010). In Portugal, studies conducted in the Lisbon area report a
pneumococcal carriage rate of ~64% in 2010 (Nunes et al., submitted).
Pneumococcal disease
From the nasopharynx pneumococci can spread to other body sites to cause a wide
range of diseases that vary in prevalence and severity. The most common forms of
disease are mild infections such as acute otitis media (AOM) and sinusitis. The
nasopharynx is connected to the middle ear via the Eustachian tube which, if clogged,
can trap bacteria and cause a middle ear infection. Despite being a mild infection, AOM
has a high socio-economic impact as it is the leading bacterial infection in children in
high income countries and the primary reason for antibiotic prescription (Hau et al.,
2013). In fact, by the age of 3 years, over 80% of the children are estimated to have an
AOM episode and over 40% to have recurrent episodes (Vergison et al., 2010).
Overall, the serotypes more frequently associated with AOM are serotype 3, 6A, 6B,
9V, 11A, 14, 19A, 19F and 23F, although this may vary with geographical location and
vaccine coverage (discussed below in this chapter) (Rodgers et al., 2009).
The pneumococcus can also be responsible for more severe infections, such as
pneumonia (non-invasive disease), bacteremia and meningitis (invasive disease).
Pneumonia is the non-invasive pneumococcal disease with highest burden in adults. It
has high incidence rates and high mortality risk, especially in the elderly, and is the
most common infectious source for invasive pneumococcal disease (Drijkoningen &
Rohde, 2013).
The incidence of invasive pneumococcal disease (IPD) in any population depends on
several factors, such as geographic location, season of the year, serotype prevalence,
age, co-morbidities, and vaccine coverage (Drijkoningen & Rohde, 2013; Hausdorff et
age, children younger than 2 years of age, and those with certain underlying conditions
that compromise the immune system, such as HIV infection and chronic diseases.
The World Health Organization estimated the occurrence of 14.5 million global cases of
serious illness in children less than 5 years old (O'Brien et al., 2009). According to the
USA Active Bacterial Core surveillance (ABCs) database of the Emerging Infections
Program Network, in 2013, the incidence of invasive pneumococcal disease in
individuals >65 years of age was 30.5 cases per 100,000 population and, in infants <1
year, the incidence was 29.8 cases per 100,000 population (CDC).
The serotype prevalence in disease also varies with age, vaccine coverage and type of
disease. Serotypes 4, 6B, 9V, 14, 18C, 19F and 23F were the most prevalent in
pediatric invasive disease in the USA before the introduction of the seven-valent
pneumococcal conjugate vaccine (PCV7) (Hausdorff et al., 2005), while serotype 19A
emerged after PCV7 introduction (Richter et al., 2009). Serotypes 1, 4 and 14 have
been associated with bacteremia (Hausdorff et al., 2000). Serotype 1 has also been
associated with pneumonia, together with serotype 3 (Hausdorff et al., 2005).
Figure 1. Incidence rate of invasive pneumococcal disease in children under 5 years
In Portugal, a study has shown that serotypes 1, 7F and 19A accounted for 61% of
pediatric invasive pneumococcal disease cases between 2006 and 2008 (Aguiar et al.,
2010).
From the information provided above it is clear that, although there are over 95
serotypes described, only a few are responsible for causing disease. Several studies
have aimed at establishing differences in invasive potential among serotypes
(Brueggemann et al., 2003; Brueggemann et al., 2004; Greenberg et al.; Sá-Leão et
al., 2011; Sleeman et al., 2006). These studies had different designs and were conducted in different geographical locations but the overall results were similar, as
most reported a high invasive disease potential for serotypes 1, 3, 4, 5, 7F, 9V, 14, and
18C. The study conducted in Portugal identified the same serotypes as having a high
invasive potential, among others, while serotypes 6A, 6B, 11A, 15B/C, 16F, 19F, and
23F, among others, to be mostly associated with colonization (Sá-Leão et al., 2011).
More importantly, this study has shown that different genetic lineages of the same
serotype have different invasive potential, as is the case of serotypes 3, 6A, 6B, 19A,
19F, and 23F, among others.
Non-susceptibility to antimicrobial agents
Non-susceptibility of S. pneumoniae to commonly used antibiotics has been reported worldwide (CDC; ECDC, 2013) and has increased dramatically since the description of
the first clinical isolate resistant to an antimicrobial agent (penicillin) in 1967 (Hansman
et al., 1971).
The distribution of non-susceptible isolates seems to be associated with the capsular
pneumococcal conjugate vaccine coverage (Dagan & Klugman, 2008; Goossens,
2009; Hausdorff et al., 2005; Riedel et al., 2007).
In general, high rates of non-susceptibility are observed in countries with high antibiotic
consumption, such as Southern and Eastern European countries, whilst low
non-susceptibility rates are observed in countries with low antibiotic consumption, such as
Northern European countries (Goossens, 2009; Riedel et al., 2007). Accordingly,
strains isolated from carriage tend to be more frequently non-susceptible to
antimicrobial agents than strains isolated from disease episodes (Hausdorff et al.,
2005).
The latest report from the European Center for Disease Control and Prevention
(ECDC) showed that, in 2013, the percentage of isolates non-susceptible to penicillin
causing invasive disease ranged between < 1.1% (the Netherlands) and 40.0%
(Cyprus), while for macrolides non-susceptibility rates ranged between 1.5% (Latvia)
and 38.1% (Romania) (ECDC, 2013). In the USA, in the same year, the percentages of
penicillin and macrolide non-susceptible isolates were 4.7% and 28.5%, respectively
(CDC).
In Portugal, data from carriage isolates from 2010 showed percentages of
non-susceptibility to penicillin and macrolides of 19.6% and 25.2%, respectively. The same
study showed that in that period, non-susceptibility to those antimicrobial agents was
associated mainly with serotypes 6C, 15A, 19A, 19F, and non-encapsulated isolates
(Nunes et al., submitted).
A study focused on isolates causing invasive disease in Portugal, from 2006-2008,
showed that the rates of non-susceptibility to penicillin and macrolides were 18.7% and
antimicrobial agents was associated mainly with serotypes 19A, 14, 19F, 6C and 23F
(Aguiar et al., 2010).
Anti-pneumococcal vaccination
Being S. pneumoniae “the captain of the men of death” (Osler, 1901), it is not surprising that several attempts have been made throughout history to prevent
pneumococcal disease through vaccination.
Although not effective, the first attempt was the use of killed whole bacterial cells to
immunize mineworkers, in 1911, to prevent death by pneumonia (Wright et al., 1914).
The demonstration of the efficacy of pneumococcal vaccines was a long process,
mainly due to flawed study designs and serious adverse effects of the vaccines.
Although some clinical trials were undertaken, the discovery of antibiotics and their
efficacy against pneumococcal disease put this process on hold, until Robert Austrian
proved the efficacy of pneumococcal polysaccharide vaccines and, as a consequence,
the 14-valent polysaccharide vaccine was licensed (Austrian et al., 1976). In 1983 this
vaccine was extended to 23 serotypes and licensed, being still in the market with the
commercial name of Pneumovax© 23 (PPV23). PPV23 is immunogenic for serotypes
1, 2, 3, 4, 5, 6B, 7F, 8, 9N, 9V, 10A, 11A, 12F, 14, 15B, 17F, 18C, 19F, 19A, 20, 22F,
23F, and 33F when administered in adults, the elderly and children older than 2 years
of age, as in younger children, who have an underdeveloped immune system, it fails to
induce an adequate immune response (Bogaert et al., 2004b; Douglas et al., 1983).
In 2000, the first pneumococcal conjugate vaccine (PCV7) was licensed and introduced
in the National Immunization Plan in the USA, with the commercial name of Prevnar©,
and protecting against the seven serotypes more prevalent in pneumococcal disease in
children under 6 years of age: 4, 6B, 9V, 14, 18C, 19F, and 23F (CDC, 2000). The
polysaccharide of these serotypes was conjugated to a protein carrier, CRM197,
Two additional conjugate vaccines have been developed in more recent years: a 10-
and 13-valent vaccine, in 2009 and 2010, respectively (Synflorix© and Prevnar©13,
respectively). Synflorix© (PCV10) includes the 7 serotypes present in PCV7 and 3
additional serotypes - 1, 5 and 7F – conjugated to Haemophilus influenza protein D as
the carrier protein. Prevenar©13 (PCV13) replaced PCV7 in 2010 and includes six
additional serotypes, conjugated to the same carrier protein, CRM197: 1, 3, 5, 6A, 7F,
and 19A.
In Portugal, PCV13 was introduced in the National Immunization Plan for children born
after January 2015 on a scheme of two doses followed by a booster dose.
Nevertheless, PCVs have been available in the market since June 2001 (PCV7), April
2009 (PCV10) and January 2010 (PCV13) and their use in young children has been
recommended by the Pediatric Portuguese Society. Estimates on PCV7 coverage in
2009 reached 62%, while PCV10 coverage was estimated to be 13% in the same year
(data from the National Statistics Institute (INE) and IMS).
Effects of pneumococcal vaccination
The introduction of PCV7 has led to several changes in pneumococcal epidemiology
and population structure. These changes are more or less significant according to the
vaccine coverage and geographic location and included: (i) decrease in IPD in
vaccinated children, (ii) decrease in IPD in non-vaccinated children and in adults, as a
reflex of the herd-immunity effect, (iii) increase in the incidence of IPD caused by
non-vaccine serotypes, and (iv) replacement of non-vaccine types by non-non-vaccine types in
carriage. The USA, a country with immediate massive PCV7 use, is a good example of
all these changes. A 77% decrease in IPD cases in vaccinated children was reported
four years after vaccine introduction (CDC, 2008) and later a 45% decrease in IPD in
all age groups was observed, accompanied by a 94% reduction of vaccine types
to non-vaccine types also increased in the first years of vaccination, particularly due to
serotype 19A (Hicks et al., 2007).
In Europe, the same effects were observed but with some variability among countries,
depending on vaccine coverage. Decrease in IPD incidences varied between 28% and
68% and serotype replacement was also observed, with non-vaccine serotypes 1, 19A,
3, 6A, and 7F being the ones with higher incidence in IPD after widespread use of
PCV7 (reviewed in (Isaacman et al., 2010)).
In Portugal, the same scenario was observed. The incidence of IPD cases associated
with PCV7 serotypes was reduced from 56% (before PCV7 introduction) to 17% by
2006-2008, in children aged up to 17 years of age (Aguiar et al., 2010). Carriage of
PCV7 serotypes also decreased, while the prevalence of non-PCV7 serotypes
increased, maintaining the overall prevalence of carriage in the population (Sá-Leão et
al., 2009) (Nunes et al., submitted).
Since serotypes included in PCV7 are frequently resistant to antimicrobial agents,
introduction of PCV7 vaccination was predicted to have an impact on the levels of
antimicrobial resistance worldwide (Dagan & Klugman, 2008; Kyaw et al., 2006). This
decrease was, in fact, observed following the first years of vaccination but was rapidly
compensated by the emergence of drug-resistant non-PCV7 serotypes (Mera et al.,
2009). In Portugal, antimicrobial resistance rates were maintained, despite the
widespread use of PCV7 (Simões et al., 2011) (Nunes et al., submitted).
PCV10 and PCV13 have similar effects on the population as the ones described for
PCV7, with reports on the impact these vaccines becoming more frequent (Hammitt et
al., 2014; Jokinen et al., 2015; Moore et al., 2015; Waight et al., 2015). In the USA a study reported a decrease of 93% in IPD cases attributable to PCV13-only serotypes,
in children under 5 years old, and a decrease of 58-72% in adults, depending on age,
serotypes IPD cases was of 69% for all ages (Waight et al., 2015). However, both
studies reported an increase in IPD cases associated with non-PCV13 serotypes,
particularly in children under 5 years old, evidence of the serotype replacement
expected to occur.
Despite the serotype replacement that has been observed and that is attributable to the
fact that current vaccines only target a limited number of serotypes, the strategy of
targeting the most prevalent serotypes in IPD has been highly successful. However,
the adaptability of the pneumococcal population to these human interventions has been
a driving force for the development of the ideal vaccine. This vaccine would be a
conserved, universally present surface protein, capable of eliciting a good immune
response in all age groups. No such protein has been described so far but several
initiatives are ongoing, using surface proteins, combinations of proteins or an whole cell
approach to find a vaccine that could ultimately overcome the limitations of currently
used conjugate vaccines (reviewed in (Feldman & Anderson, 2014)).
Pneumococcal co-colonization
Biological significance
The preferential niche of the pneumococcus is the nasopharynx, a polymicrobial
environment where the pneumococcus is forced to interact with other microorganisms
of the same or different species. Simultaneous colonization by more than one
pneumococcal strain, or co-colonization, is frequent and is of high clinical,
epidemiological and ecological relevance (O'Brien & Nohynek, 2003; Shak et al.,
2013).
It has been shown that colonizing pneumococcal strains can interfere both negatively
2000; Marks et al., 2012b). Marks et al. (Marks et al., 2012b) have shown in a mouse model that strains with poor colonization efficiency have their efficiency improved when
in co-colonization with a highly efficient strain that is already established. This
improvement was independent of genetic exchange between strains. On the other
hand, Lipsitch et al. (Lipsitch et al., 2000) have shown, also in a mouse model, that carriage of an established strain can inhibit acquisition of a second strain in a strain-
and density-dependent manner. The inhibition was observed through a reduced
colonization probability and/or through reduced cell counts in the colonized animals.
Additionally, some molecular mechanisms have been shown to contribute to inter-strain
interactions, namely bacteriocin production and competence-mediated fratricide, which
will be discussed later in this chapter (Dawid et al., 2007; Guiral et al., 2005).
Co-colonization is a requirement for horizontal gene transfer between pneumococci
through homologous recombination, a mechanism important for evolution in this
species (Coffey et al., 1991; Feil et al., 2000; Smith et al., 1993; Spratt et al., 2001).
Several studies reported the occurrence of genetic exchange in biofilm models (Carrolo
et al., 2014; Marks et al., 2012b; Wei & Havarstein, 2012) and also in in vivo models (Marks et al., 2012b). Additionally, Hiller et al. (Hiller et al., 2010) have shown the occurrence of genetic rearrangements in vivo that generated genetic diversity during a
pediatric chronic infection. Several population genetic studies have also shown the
occurrence of genetic recombination, including capsule switching events (Croucher et
al., 2011; Donkor et al., 2011). Donkor et al. (Donkor et al., 2011) have used an MLST approach to characterize pediatric colonization strains and they found evidence of
extensive recombination among those strains, which they have related to the high
prevalence of co-colonization in the population.
Besides the evolutionary implications, DNA exchange between co-colonizing strains
can have important clinical and epidemiological consequences, not only because it has
significant changes in the virulence of a strain (Hiller et al., 2011), but also because this
genetic exchange has played a role in the development and evolution of antibiotic
resistance (Dowson et al., 1989). The occurrence of capsule switching events is also of
importance as it can have an impact on the virulence of strains and, in the context
vaccine effectiveness, can result in the emergence of vaccine escape recombinants
(Brueggemann et al., 2007). In the same context, understanding co-colonization is
critical to monitor the extent of serotype replacement by minor serotypes when
vaccination targets the most abundant ones (Lipsitch, 1999; Lipsitch, 2001; Spratt &
Greenwood, 2000).
Another important clinical implication is the increasing evidence that co-colonization
might be associated with higher colonization densities. Brugger et al (Brugger et al.,
2010) showed that colonization density was significantly higher in nasopharyngeal
swabs containing multiple pneumococcal strains, compared to swabs with a single
strain. Accordingly, Margolis et al (Margolis et al., 2010) have shown, in a neonatal rat
model, that sequential colonization with two pneumococcal strains results in increased
colonization densities to allow the co-existence of the pulsed and the established
strains. This is of great importance due to the increasing interest in using
nasopharyngeal colonization density as a diagnostic tool for pneumonia (Albrich et al.,
2012). Moreover, a study that assessed co-colonization events in samples from healthy
children and children with acute respiratory infection has shown that the latter group
was significantly more co-colonized (Dhoubhadel et al., 2014).
Epidemiology
The epidemiology of co-colonization is generally unknown. There are several
surveillance studies that report colonization by more than one strain in the same host,
methods of detection and most have not systematically looked at co-colonization (for a
review see (O'Brien & Nohynek, 2003)).
Studies that used the same detection method (serotyping of multiple colonies) suggest
that co-colonization prevalence is highly variable across geographical settings, ranging
between 3% and 36% (Gratten et al., 1989; Hansman & Morris, 1988; Ussery et al.,
1996). Also, there is some evidence that co-colonization prevalence in a given setting
might be associated with the carriage levels in that setting, i.e., settings with higher carriage prevalence will have higher co-colonization levels (Brugger et al., 2010;
Kandasamy et al., 2015; Turner et al., 2011).
There is little data on the effect of vaccination on co-colonization. Brugger et al. (Brugger et al., 2010) have used a highly sensitive method to detect co-colonization in
nasopharyngeal samples collected between 2004 and 2009, spanning PCV7
introduction in Switzerland. They concluded that the rate of co-colonization was similar
before and after the introduction of PCV7 and that it was associated with younger age,
as children younger than 2 years of age were significantly more co-colonized. This
study has also shown that co-colonization rates are independent of the vaccination
status of the child, which is in disagreement with the results obtained in Portugal,
where it was shown that, in the vaccine era, children with four PCV7 doses were
significantly less co-colonized than non-vaccinated children (see chapter II).
The quest for the perfect detection method
The first reports on co-colonization were obtained by successive intraperitoneal saliva
injections in mice to detect multiple serotypes (Gundel et al. (Gundel M, 1933), reviewed in (Shak et al., 2013)). Using this strategy, Gundel et al. reported a 73% rate of co-colonization. Although impractical, this methodology was the basis for the
by serotyping of several colonies, to detect co-colonization. The latter strategy was
used for several years and constitutes the basis of most reports on co-colonization
(Barker et al., 1989; Gratten et al., 1989; Hansman & Morris, 1988; Huebner et al.,
2000; Ussery et al., 1996). Despite being the most straightforward approach for
detection of co-colonization, serotyping of multiple colonies has proved to be of little
value due to its low sensitivity to detect less abundant serotypes in a nasopharyngeal
sample. Huebner at al. (Huebner et al., 2000) have shown that, to have a 95% chance
of detecting a serotype present in a sample in a relative abundance of 1%, one would
need to serotype 299 colonies. This is highly demanding in feasibility and cost.
For the reasons mentioned above, several alternative serotyping methods have been
developed, taking into consideration the ability to detect co-colonization. The criteria to
define a good detection method rely on the ability of that method to (i) detect minor
serotypes with high specificity, (ii) detect serotypes directly from the nasopharyngeal
specimen, (iii) detect all known serotypes, (iv) distinguish pneumococci from closely
related species, (v) be quantitative, and (vi) be affordable (Satzke et al., 2012; Satzke
et al., 2014).
These methods can be divided into phenotypic and genotypic and some are variants of
the same technology (Table I).
Most phenotypic methods are either bead-based (Park et al., 2000; Sheppard et al.,
2011; Whaley et al., 2010) or blot-based immunoassays (Bogaert et al., 2004c;
Bronsdon et al., 2004). The blot-based methods have the advantage of being
cost-effective because they use highly diluted typing sera but they are technically
demanding and can be of difficult interpretation due to cross-reactions. The
bead-based methods are technically demanding and of difficult implementation, as they
Turner et al. (Turner et al., 2011) have also developed a phenotypic method based on
latex agglutination. This method uses an enrichment step of the nasopharyngeal
specimen according to the WHO recommendations and resuspension of the primary
selective growth on a saline solution to be tested against all available typing sera,
allowing the detection of virtually all capsular types or groups. The authors were able to
detect minor serotypes that corresponded to down to 25% of the total pneumococcal
population.
The genotypic methods include PCR-based methods, including real-time PCR, and
microarray-based methods. Their main advantage is their higher sensitivity, when
compared to phenotypic methods, which enables the detection minor serotypes
present in a very low abundance that could be below the culturable limit.
Several multiplex-PCR systems were developed that are able to detect co-colonization,
although all of them require a previous enrichment of the nasopharyngeal specimen
(Morais et al., 2007; Rivera-Olivero et al., 2009; Yu et al., 2011). These methods have
the limitation of having low discriminative power in closely-related capsular types, of not
being quantitative and the fact that most of them target a limited number of capsular
types. They present high sensitivity and are of straightforward implementation and
execution.
Real-time PCR based methods have also been developed (Azzari et al., 2010; Pimenta
et al., 2012). These methods are extremely sensitive but are limited by the low discriminative power in closely-related capsular types and by the optimization
difficulties for multiplexing. Azzari et al. (Azzari et al., 2010) have shown, however, the
main advantage of this approach, by demonstrating its ability to detect co-colonization
directly from the nasopharyngeal sample with a four times higher sensitivity than
sequential multiplex PCR. Another key advantage of this method is the fact that it is
The main drawback of both conventional and real-time PCR based methods is the
possible occurrence of false positive results associated with the fact that these
methods use as target fragments of capsular genes that can potentially be present in
other inhabitants of the nasopharynx, such as closely related Streptococcus spp. (Carvalho Mda et al., 2013).
Brugger et al. (Brugger et al., 2009) have developed a PCR-based method which takes
advantage of a highly conserved and specific genetic region within the pneumococcal
species, adjacent to the pneumolysin gene (ply), which encodes for a major virulence
factor and is highly specific for pneumococcus. This variable noncoding region adjacent
to the ply gene (plyNCR) is amplified by PCR and the PCR product is then digested for
restriction fragment length polymorphisms (RFLP). The restriction is performed with
four enzymes used independently and the samples are analyzed by capillary
electrophoresis for exact size assessment of the digestion fragments. When the sum of
the restriction fragments exceeds the size of the undigested product (c.a. 1400 bp),
there is evidence that more than one pneumococcal strain exist in the sample. This
method has high throughput but requires expensive equipment and reagents and a
fairly high level of technical expertise, as interpretation of results can be misguided by
incomplete digestion or presence of closely-related Streptococcus spp. The greatest disadvantage of this method is related to the fact that it does not discriminate the
serotypes of the co-colonizing strains. For this reason, it has to be coupled with another
genotypic method that has this ability.
Genotypic methods based on comparative genomic hybridization (CGH) have also
been developed (Hinds et al., 2010; Tomita et al., 2011; Turner et al., 2011; Wang et
al., 2007). However, to our best knowledge, only the microarray developed by Hinds et al. (Hinds et al., 2010; Turner et al., 2011) has the ability to detect all capsular types described up to now. In this method each capsular gene is targeted by ten 60-mer
capsular types. As detection is based on fluorescently labeled oligomers, this
microarray is able to quantify the relative abundance of each capsular type in the total
of pneumococcal DNA present in the samples. Despite being very sensitive, this
method is not suitable for detection directly from the nasopharyngeal specimen,
although optimization is ongoing (Satzke et al., 2012). The main drawback of this
method is, however, the requirement of very expensive reagents and equipment and
high level of technical expertise.
Due to the existence of all these different approaches and to the increasing relevance
attributed to co-colonization in vaccine surveillance studies, an initiative was
undertaken under the scope of the PneuCarriage project in which 20 methods were
compared for their ability to detect co-colonization with a good cost/benefit ratio. This
assessment is being done by using a reference collection of spiked samples mixed with
real samples, in which methods with the best performance in detecting minor serotypes
with good positive predictive value will be selected for a second round that consists in
the analysis of 200 field samples (Satzke et al., 2012). First reports announced that
four methods were selected for the second stage of evaluation: the microarray
developed by Hinds et al. (Hinds et al., 2010; Turner et al., 2011), the latex agglutination method developed by Turner et al. (Turner et al., 2011), the plyNCR-RFLP coupled with multiplex PCR, developed by Brugger et al. (Brugger et al., 2009), and the multiplex PCR and real-time PCR developed by Azzari et al. (Azzari et al., 2010).
Co-colonization determinants
Co-colonization has been known to occur for many years, although the factors
determining its occurrence and the interactions that occur between strains inside the
Several theoretical models have been used to predict which are the determinants of
co-colonization and strain interactions, all of them considering the capsular type as the
decisive factor to distinguish between strains (Auranen et al., 2009; Cobey & Lipsitch,
2012; Zhang et al., 2004). These studies have resulted in the formulation of important
premises about between-strain competition and carriage of multiple serotypes:
co-colonization does not seem to accelerate clearance (Auranen et al., 2009), established
colonization with one serotype reduces risk of acquisition of a second serotype
(Auranen et al., 2009), serotype-specific immunity stabilizes competition, and acquired
immunity to non-capsular antigens reduces fitness differences between strains, being
relevant for competition (Cobey & Lipsitch, 2012; Zhang et al., 2004). Notwithstanding,
these theoretical principles have not been sufficiently addressed experimentally.
Experimental and epidemiological data on this subject is scarce and the bacterial
factors that might affect this phenomenon are generally unknown. In the following
sections information regarding some bacterial factors that might have an impact on
co-colonization and that are relevant in the context of this thesis will be presented.
The contributions of the capsule and genetic background to
(co)colonization
The ability of a strain to colonize an individual is intrinsically related to its ability to resist
clearance. With the objective of determining the invasive potential of different
serotypes, Sleeman et al. (Sleeman et al., 2006) have shown that less invasive serotypes are carried for longer periods. Most studies that have addressed which
factors contribute to the a success of a strain in colonization have focused on the role
of the capsule, as it is the main target of the immune system in the clearance process
capsule prevents clearance and aggregation, affects colonization and adherence, helps
the pneumococcus to survive in the lungs and spread to the bloodstream, and
contributes to antibiotic tolerance. Pneumococci regulate the amount of capsular
material produced during colonization and invasion. Transparent capsules (thin) are
favored during initial colonization stages to promote adherence to host epithelial cells,
while opaque capsules (thick) are favored during invasion to resist
complement-mediated opsonophagocytosis (Fernebro et al., 2004; Lysenko et al., 2005; Mac &
Kraus, 1950; Morona et al., 2004; Nelson et al., 2007; Weiser et al., 1994).
Weinberger et al. (Weinberger et al., 2009) have shown that the structure of the capsular polysaccharide can predict serotype prevalence. These authors and others
have shown that serotypes with capsule structures with more carbon in their
polysaccharide repeat unit required more energy for capsule production, displaying
thinner capsules. Therefore, these serotypes were less resistant to phagocytosis and
clearance, which correlated with their lower prevalence (Hathaway et al., 2012;
Weinberger et al., 2009). Other capsule-related properties have been studied to explain
the influence of the capsular type on pneumococcal colonization, such as the surface
charge. Li et al. (Li et al., 2013) have shown that the surface charge correlates with the
prevalence of serotypes in colonization, as more negatively charged capsular types are
more prevalent than less charged capsular type and capsules more negatively charged
were shown to repel host immune cells that contribute to clearance, such as
neutrophils and macrophages.
Several epidemiological studies have shown that there is high variability in strains of
the same serotype (Brueggemann et al., 2003; Hanage et al., 2005; Sá-Leão et al.,
2011; Sandgren et al., 2004) and this observation is supported by the increasing
available genomic data (Chewapreecha et al., 2014; Hakenbeck et al., 2001; Hiller et