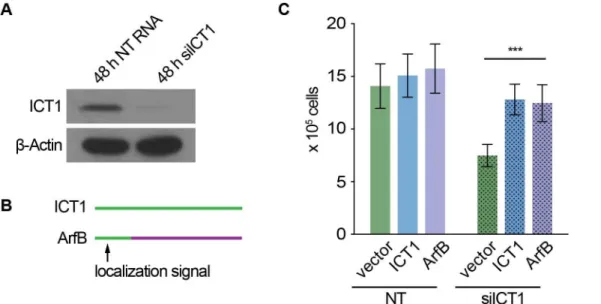
Human Cells Require Non-stop Ribosome Rescue Activity in Mitochondria.
Texto
Imagem

Documentos relacionados
Results indicate that for a population of adolescent athletes with an amateur sports level, the dyskinesis prevalence is high; however, it is not associated with pain and does
The results indicate that the increase of biodiesel in mineral diesel reduces torque and power, increases the specific fuel consumption and practically does not change the
As destinações possíveis para grãos quimicamente tratados são o envio para aterro, a incineração, o coprocessamento em fornos de cimento, a compostagem e
O senhor tem que trazer soluções para certos problemas específicos - por exemplo, como melhorar as anemotecnicas atualmente utilizadas, como obter mais rapidamente
Results indicate that, on average, the impact of the concession announcements on stock returns is negative and suggest that the participation in these projects do not add value to
Conclusions: These results indicate that a single application of ultrasound and ultrasound associated with stretching were not able to modify the activity pattern of the
geniculatus to the enclosures of domestic pigs near human dwellings on the river flood plain and the attack of these triatomines on people within these houses indicate that
Todavia, é possível entender que a baixa densidade do grafo não interferiu, de forma alguma, no objetivo desenhado, afinal como o intuito deste trabalho era avaliar as categorias