Universidade de Lisboa
Faculdade de Ciências
Departamento de Biologia Animal
UNDERSTANDING THE ROLES OF PIWIL3 AND
TACC3 PROTEINS DURING BOVINE OOCYTE
DEVELOPMENT
ANA RITA CANHOTO LEITOGUINHO
Dissertação
Mestrado em Biologia Evolutiva e do Desenvolvimento
2012
Universidade de Lisboa
Faculdade de Ciências
Departamento de Biologia Animal
UNDERSTANDING THE ROLES OF PIWIL3 AND
TACC3 PROTEINS DURING BOVINE OOCYTE
DEVELOPMENT
ANA RITA CANHOTO LEITOGUINHO
Dissertação
Mestrado em Biologia Evolutiva e do Desenvolvimento
Dissertação orientada por:
Professora Dra. Maria Gabriela Rodrigues (DBA/FCUL)
Professor Dr. Bernard Roelen (BRC/UU)
2012
Acknowledgements
I would like to express my deep and sincere gratitude to Bernard for this opportunity and for being always there when I needed. Also a special thanks to the entire lab, in particular to Mahdi for his patience, help and great talks, to Eric for the tips on music and to Leni for her constant guidance and assistance in the Lab. From the warm and dry part of Europe, my sincere thanks to Gabriela for her support and great advices. Para a minha mãe, pai e irmã um muito obrigado por me providenciarem tudo o que sempre precisei, mesmo nestes tempos difíceis. Nunca teria conseguido sem a vossa ajuda.
To all my friends, can’t thank you enough. Thank you to Nuno, Luís, Silvia, Yuri and Filipa for your help during rough times and for your friendship. Also, to all Utrecht friends, a big cheers for your welcoming feeling. Last but not least, a loving gratitude to Stephin for his boundless wisdom, smile and company. This thesis is for all of you. Ana Rita Leitoguinho / Toga In loving memory of my grandfather
Abstract
In order to achieve a healthy embryo, the germline needs to properly develop while protecting its genome against endogenous and exogenous adversities. The PIWI proteins, a subgroup of Argonaute proteins, are involved in the search and silencing mechanism of transposons ‐ genetic transposable elements‐ through a mechanism of complementarity with piRNAs (Piwi‐interacting RNAs). These proteins are germline‐specific and are thought to be also male‐specific in mammals. PIWIL3 (Piwi‐like 3) is part of this family and it is present in the human and bovine genome, while rodents have PIWIL1, PIWIl2 and PIWIL4, but lack a gene for PIWIl3. Recent findings on bovine cells suggested a role for PIWIL3 in the female germline, the oocyte. Here, PIWIL3 protein was detected both in the bovine oocyte and in somatic cells surrounding it, using an antibody directed against the Human PIWIL3 suggesting a role for this protein in protecting the oocyte genome.
Besides protecting the genome, the oocyte also needs to enroll meiosis. Transforming acidic coiled‐coil 3 (TACC3) acts in a complex with ch‐TOG and clathrin to ensure mitotic spindle stability and organization. TACC3 is phosphorylated and activated by Aurora‐A kinase and it is responsible for TACC3’s proper placement in the centrosomes. The order of recruitment of these proteins to the spindle is still open for debate, as it is the role of TACC3 during meiosis. Our antibody designed to identify bovine PIWIL3 was in fact recognizing TACC3 and we could therefore study the spatiotemporal localization of TACC3 in the oocyte and during embryo development. When bovine oocytes are matured with MLN8054, an Aurora‐A inhibitor, they exhibit (i) dispersion of TACC3 protein in the cytoplasm (ii) formation of an abnormal meiotic spindle (ii) meiotic arrest in the metaphase I stage and (iii) decreased percentage of blastocyst formation suggesting a role of TACC3 in meiotic spindle assembly, vital for oocyte and early embryo development.
Resumo em Português
Todos os indivíduos que se reproduzem sexualmente combinam a sua informação genética para criar um novo ser e apesar de todas as células possuírem esta informação, apenas as células sexuais são capazes de a transmitir à descendência. É por isso vital proteger estas células de fatores externos ou internos que possam afetar o seu genoma bem como certificar um correto desenvolvimento do oócito com o intuito de formar um embrião saudável.
O processo de maturação do oócito envolve o desenvolvimento da célula bem como a criação de estruturas de suporte que o protegem de agressões externas. A Foliculogénese é processo responsável pelo suporte físico do oócito, altamente regulado hormonalmente pelo ovário, onde várias camadas celulares são formadas consecutivamente em volta do oócito, contribuindo para que o resultado final seja uma célula sexual protegida, formando o folículo antral. Concomitantemente, também a própria célula sofre alterações críticas. Durante a maturação nuclear do oócito, ou oogénese, é essencial que este reduza o seu número de cromossomas a metade, formando um gameta haplóide (com metade da informação genética). Após fertilização, este resultará num zigoto diplóide (com o total da informação genética) com informação materna e paterna que diferem não apenas na informação genética mas também em fatores associados que são transmitidos não por via genética, mas por via epigenética.
Para além dos rearranjos nucleares acima referidos, também o genoma da célula sexual está sujeito a agressões endógenas, como os transposões, elementos móveis do genoma que se podem inserir em diversas zonas, podendo interferir com a transcrição de genes vitais para a célula. Existem vários mecanismos de defesa celular que silenciam estes elementos, um destes é o RNA de interferência (RNAi) que, com ajuda de proteínas Argonautas, consegue localizar e silenciar RNA mensageiro (mRNA) através de um mecanismo de complementaridade, evitando a sua propagação pelo genoma. Esta família de Argonautas contém o grupo de proteínas PIWI cuja expressão é restrita às células da linha germinal. Recentemente descobriu‐se que estas proteínas PIWI são alvo de um tipo particular de RNAi, os piRNAs (Piwi‐interacting RNAs) cuja principal função é silenciar elementos transponíveis na linha germinal. Tendo como base mutantes PIWI em Drosophila que ativam transposões, o estudo de proteínas PIWI e piRNA tem avançado bastante nos últimos tempos revelando também em C. elegans, Zebrafish e ratinho o papel destas
proteínas na formação da linha germinal. No ratinho existem três genes da família PIWI: Miwi (Piwi‐like 1), Mili (Piwi‐like 2) e Miwi2 (Piwi‐like 4) e mutantes destes genes resultam em esterilidade masculina, em contraste com Drosophila e Zebrafish onde ambas as linhas germinais masculina e feminina são afetadas. Até à data não se sabe a função destas proteínas na linha germinal feminina de mamíferos, razão pela qual a descoberta de um quarto gene Piwi‐like (PIWIL3) em mamíferos que não o ratinho suscitou bastante interesse. Recentemente, este gene foi descrito em oócitos de bovinos e a facilidade em obter oócitos de vaca fizeram com que este organismo fosse o escolhido para o estudo desta proteína PIWIL3.
Os nossos resultados revelaram primeiro que tudo que o anticorpo e primers construídos anteriormente para a proteína PIWIL3 em vaca estavam mal construídos. Propomos aqui um padrão de expressão proteico diferente do anteriormente documentado. Com o uso de anticorpos anti‐humano dirigidos a PIWIL3, identificou‐se a proteína nas células de suporte que rodeiam o oócito, sugerindo que a expressão desta proteína PIWI em vaca não é específica para a linha germinal. Esta presença de PIWIL3 na linha somática que rodeia o oócito pode ser essencial para a proteção do genoma do oócito, essencial para a correta propagação da informação genética à descendência.
Aquando do estudo de PIWIL3, descobriu‐se que o anticorpo estava a reconhecer uma outra proteína em vez de PIWIL3, a TACC3 (Transforming Acidid Coiled‐Coil protein 3), o que fez com que este projeto se começasse a focar no papel desta outra proteína durante o desenvolvimento do oócito.
Também uma correta divisão celular é essencial para uma célula sexual e as proteínas TACC (Transforming acidic coiled‐coil proteins) estão de mãos dadas com esta prática. Estas proteínas estão presentes em diferentes organismos e localizam‐se tanto nos microtúbulos que se formam durante o processo de separação dos cromossomas como nos centrossomas, estruturas que se localizam nos dois pólos opostos do fuso mitótico. Em ratinho são conhecidas três proteínas TACC ‐ TACC1, TACC2 e TACC3 ‐ e sabe‐se que TACC3 é essencial para a correta formação do fuso mitótico durante uma mitose. Mutantes TACC3 revelaram um fuso mitótico anormal que, juntamente com cromossomas desalinhados na placa metafásica, originam incorreta segregação cromossómica, resultando em células‐filha com diferente conteúdo cromossómico. O recrutamento de TACC3 para os microtúbulos que formam o fuso é regulado por uma kinase, a Aurora‐A, que é responsável pela fosforilação e ativação desta proteína em diversas espécies como Drosophila, C.elegans, Xenopus e
Humanos. Quando esta kinase é inibida, por exemplo pelo composto MLN8054, a mitose falha de forma semelhante aos mutantes de TACC3, contudo alguma células conseguem dividir‐se, ainda que originando células aneuplóides ou tetraplóides. Porém, TACC3 não atua sozinha e recentemente foi descoberto que o faz em conjunto com outras duas proteínas num complexo proteíco denominado TACC3/ch‐TOG/clatrina. TACC3, clatrina e ch‐TOG (Colonic‐hepatic Tumour Overexpressed Gene) atuam como pontes entre microtúbulos e têm como função estabilizar estas fibras. Apesar de se saber que todos os intervenientes do complexo são importantes, a ordem de recrutamento dasdiferentes proteínas ainda é alvo de uma acesa discussão entre defensores que afirmam que clatrina recruta TACC3 e outros que defendem ser TACC3 o primeiro componente do complexo, recrutando clatrina.
Apesar de ter sido estudada em oócitos de ratinho, informação sobre a função da proteína TACC3 durante a meiose de mamíferos é quase nula, o que realça a importância deste estudo. Os nossos resultados demonstram que a proteína se encontra rodeando a cromatina do oócito e no fuso meiótico do mesmo durante a meiose, sugerindo um papel semelhante ao documentado no ratinho quanto à manutenção dos microtúbulos do fuso. Quanto à sua função, foi utilizado um inibidor de Aurora‐A, MLN8054, que resultou numa paragem no desenvolvimento do oócito na metafase I, efeito semelhante aos previamente descritos em células humanas. Também o fuso meiótico revelou sérias perturbações quando os oócitos maturam num meio com o inibidor o que, juntamente com a dispersão da proteína TACC3 no citoplasma e a diminuição da percentagem de blastocistos formados, sugere que este inibidor estará a influenciar a expressão ou ativação de TACC3. Apesar de não conseguirmos saber ao certo com este estudo se TACC3 esta de fato inibida, é essencial averiguar o padrão de fosforilação desta proteína para poder estabelecer uma ligação causa‐ efeito entre o inibidor e a fosforilação/correta localização de TACC3. Também nesta tese é proposto um modelo de recrutamento do complexo que se baseia na possibilidade de ser TACC3 o primeiro e talvez principal interveniente.
Em suma, os resultados desta tese referentes a TACC3 e a PIWIL3 revelam que ambas aparentam ter uma função durante o desenvolvimento da linha sexual feminina e é considerado de extrema importância estudar ambas. Quanto aos entraves que o projeto PIWIL3 sofreu, apesar de ter afetado parte da investigação da PIWIL3, serviu também para conhecer esta nova proteína, a TACC3, em oócitos. Esta tese propõe um padrão de expressão diferente de PIWIl3 com uma possível função também na linha somática em redor do oócito. Quanto a TACC3 os nossos resultados revelaram que quando a sua ação é inibida,
os oócitos não progridem na oogénese e revelam anomalias no fuso. Assim é sugerido que esta proteína tenha um papel crucial no processo de formação e correto alinhamento dos cromossomas na placa metafásica durante o desenvolvimento do oócito de vaca.
Table of Contents
Acknowledgements ... I Abstract ... II Resumo em Português ... III Table of Contents ... VII Index of Figures ... IX Index of Tables ... X Index of Supplementary Tables ... X Introduction ... 1 Follicle development protects the cell from outside adversities ... 1 Oocyte Development and Maturation ... 2 Protecting germ cell’s integrity via Argonaute proteins ... 5 Unraveling PIWI proteins in germ cells throughout species ... 6 Disclosures in the PIWIL3 former research ... 7 Transforming Acidic Coiled‐Coil (TACC) Proteins ... 9 Phosphorylation and activation of TACC3 by Aurora‐A kinase ... 11TACC3/ch‐TOG/clathrin complex stabilizes kinetochore fibers by inter‐microtubule bridging ... 12 TACC3/ch‐TOG/clathrin complex activation, two hypotheses ... 13 Material and methods ... 15 Oocyte collection and in vitro maturation ... 15 In vitro fertilization and embryo culture ... 15 Immunocytochemistry ... 16 Immunofluorescence ... 17 Western blotting ... 18 Aurora‐A inhibition experiment ... 19 Results ... 20 Sorting out the bovine Piwi‐like 3 antibody and unraveling Transforming Acidic Coiled‐ Coil protein 3 (TACC3) ... 20
Detection of PIWIL3 expression in bovine oocytes using antibodies directed against the human sequence ... 22 TACC3’s expression pattern during oocyte and early zygote development: its presence surrounding chromatin and the meiotic spindle ... 24 TACC3’s presence surrounding the metaphase plate in the metaphase II ... 26 TACC3 partially overlaps with α‐tubulin during oocyte and early zygote development ... 27
Discerning TACC3 function: using MLN8054 compound, an Aurora‐A inhibitor, decreases oocyte developmental progress and causes spindle anomalies ... 29 Effects of MLN8054 on TACC3 protein expression ... 31 Discussion ... 33 PIWIL3 during bovine oocyte development ... 33 TACC3 protein in bovine oocyte development ... 35 Future perspectives ... 39 Concluding remarks ... 39 References ... 40 Supplementary Information ... XI
Index of Figures
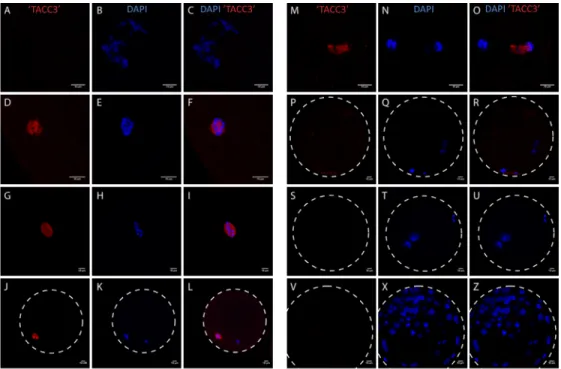
Figure 1‐ Schematic representation of bovine folliculogenesis. ... 2 Figure 2‐ Schematic representation of bovine oogenesis ... 3 Figure 3 ‐ Schematic representation of siRNA‐guided mRNA cleavage. ... 5 Figure 4 ‐ Homology between mammalian PIWI proteins. ... 7 Figure 5 ‐ mRNA quantification of PIWIl3 and PIWIL3 expression pattern ... 8Figure 6 ‐ Centrosome and spindle localization of TACC proteins in C. elegans, D. melanogaster, X. laevis and Human. ... 10 Figure 7 ‐ Depletion of Xenopus’ TACC3 (xTACC3): effects on the spindle. ... 11 Figure 8 ‐ Two models representing the recruitment of the complex TACC3/ch‐TOG/clathrin to the spindle. ... 13 Figure 9 ‐ Schematic representation of two Piwi‐like 3 aminoacid sequences ... 20 Figure 10 ‐ Double stainings TACC3 and ’TACC3’ during oocyte maturation and blastocyst. 21 Figure 11 ‐ Merge channels of (A) DAPI and PIWIL1 and (B) DAPI and PIWIL2 expression patterns during metaphase II. ... 22
Figure 12 ‐ Schematic representation of the human Piwi‐like 3 aminoacid sequence and bovine Piwi‐like 3 aminoacid sequences ... 22
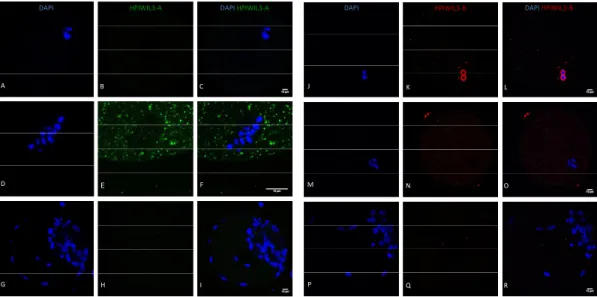
Figure 13 ‐ HPIWIL3‐A and HPIWIL3‐B stainings on different staged oocytes and blastocysts. ... 23
Figure 14 ‐ Immunocytochemistry results for HPIWIL3‐A and PIWIL3‐B in human testis, bovine testis and bovine ovary. ... 24 Figure 15 ‐ ‘TACC3’ expression pattern during oocyte development and early embryogenesis. ... 25 Figure 16 ‐ Microfilament and microtubule stainings on metaphase II oocytes ... 26 Figure 17 ‐ Double stainings ‘TACC3’ and actin on metaphase II oocytes. ... 27 Figure 18 – Double stainings of ‘TACC3’ and microtubules on oocytes and early zygotes. ... 29 Figure 19 ‐Effect of MLN8054 throughout oocyte developmental progress ... 30
Figure 20 – Effect of MLN8054 on oocyte polar body percentage and percentage of blastocyst formation from cleaved embryos ... 30
Figure 21 ‐ Effects of MLN8054 on oocyte meiotic spindle during oocyte maturation process ... 31
Figure 22 ‐ Effects of MLN8054 on TACC3 protein expression pattern during metaphase I and II oocytes. ... 32
Figure 23 ‐ Western Blot on TACC3 protein and β‐actin in four different groups ... 32
Figure 24 – Proposed TACC3/ch‐TOG/clathrin recruitment to the spindle and centrosomes during bovine oocyte development ... 38
Index of Tables
Table 1 ‐ Composition of multiple culture mediums used during oocyte in‐vitro maturation and fertilization and embryo culture. ... 15 Table 2 ‐ Detailed description of the antibodies used. ... 17 Table 3 ‐ Detailed description of the Buffers used on the Western Blot. ... 19 Table 4 ‐ Experimental design for the MLN8054 experiment. ... 19Index of Supplementary Tables
Supplementary Table 1 ‐ Effect of MLN8054 during oocyte meiosis progression ... XI Supplementary Table 2 ‐ Effect of MLN8054 during polar body extrusion. ... XI Supplementary Table 3 ‐ Effect of MLN8054 on blastocyst formation. ... XI
Introduction
For centuries, the question of how a new being comes into existence has provided a constant intellectual challenge and, since ancient Greece, several philosophers have attempted to connect fertilization events with the creation of a new being. Once the in vivo fertilization mechanism was understood, the molecular mechanisms underlying the process presented the subsequent challenge. In 1978, the first human birth originating from a successful in‐vitro‐fertilization (IVF) was announced 1, an accomplishment which earned Robert Edwards the Nobel Prize in Physiology or Medicine in 2010. The process of embryonic development goes through distinct stages with specific cell types, which differ from somatic ones in their capacity to pass our genetic code to the next generation. These are called germ cells and give rise to gametes in organisms with a sexual reproductive system. Germline development involves specification of primordial germ cells (PGCs) and their migration to special regions which will develop into the gonads, where the germ cells can be stored and preserved. The PGCs in mammals arise through a complex signaling process that, together with transcriptional regulators, inhibit somatic gene expression while activating germline genes, thereby preserving germ cell pluripotency 2. Later on, the germ cells undergo two divisions, a reductional and a non reductional one, after which they can be called either an oocyte or a sperm cell. Despite oocyte development, also the outside of the cell has to be protected and so, a follicle is matured in order to protect the oocyte integrity.
Follicle development protects the cell from outside adversities
Whilst oocyte maturation is taking place, also the exterior of the oocyte undergoes significant changes inside the ovarium through a process termed folliculogenesis (Fig. 1).
Coincident with the start of meiosis, oocytes become enclosed by a single layer of somatic cells, the so‐called pre‐granulosa cells 3, thus forming the primordial follicles (Fig. 1)
that are thought to comprise the pool of resting follicles which defines postnatal ovarian life span 4. When the follicles are recruited to undergo meiosis, the pre‐granulosa cells surrounding the oocyte change from a flattened to a cuboidal shape and are then called granulosa cells 5.
Moreover, both the oocyte and the granulosa cells grow further and the primary follicle (Fig. 1) becomes surrounded by a multiple layer of stroma cells 5, forming the theca. The oocyte becomes involved by the zona pellucida, characterizing the secondary follicle, and during this phase, it builds up a store of mRNAs proteins, crucial for its development competence 6. A subsequent tertiary follicle has a complete zona pellucida and starts to
develop an antral cavity (Fig. 1). The antral follicle is characterized by further proliferation and differentiation, where internal and external theca layers can be distinguished and the internal cavity is filled with follicular fluid (Fig.1). This cavity pushes the oocyte to one side of the follicle and it becomes surrounded by cumulus cells, forming the cumulus‐oocyte complex (COC) 7. In cattle, a tight communication between the cumulus cells, the follicular fluid and the oocyte is crucial for the oocyte’s proper development and is thought to directly influence its developmental competence both in vivo and in vitro 7,8.
Oocyte Development and Maturation
During oocyte development, two distinct maturation processes can be distinguished, nuclear maturation and cytoplasmic maturation. At the stage of cytoplasmic maturation, mRNA, proteins and nutrients accumulate in the cytoplasm, thus allowing the oocyte to sustain early embryonic development 9. Figure 1‐ Schematic representation of bovine folliculogenesis. Five phases are represented from primordial follicle to antral follicle. In bovine oocytes, primordial follicle reaches 30µm, primary follicle reach between 30‐60µm, secondary follicle 60µm, tertiary follicle 100µm and antral follicle may reach 120µm. O‐ Oocyte; ZP‐ Zona pellucida, GV‐Germinal vesicle, CC‐ Cumulus cells, GC‐ Granulosa cells, IT‐ Internal theca, ET‐ External theca. Bovine FolliculogenesisAll through mammalian nuclear maturation, the oocyte undergoes meiotic cell division, which implies a reduction in chromosome number, accomplished through two consecutive nuclear divisions termed meiosis I II. This process arrests in two distinct phases, the prophase I and the metaphase II stage, and it is unique to diploid germ cells. The process of oocyte meiosis, can be described through different stages 2,10 (Fig. 2). Meiosis starts with the prophase I, the longest and most complex phase of the meiotic cell division. This phase can be divided into five distinct stages: leptotene, zygotene, pachytene, diplotene and diakinesis. At leptotene (Fig. 2, stage 1), the chromosomes, which at this stage already consist of two sister chromatids, thicken and become visible while the centrosomes begin to move toward opposite poles. During zygotene (Fig. 2, stage 2), each chromosome seeks out its homologous partner and both are “zipped” together by a protein structure called synaptonemal complex, in a process known as synapsis. This will only be completed at the pachytene stage (Fig. 2, stage 3) where Crossing‐over occurs, i.e., the genetic exchange between non‐sister chromatids of a homologous pair. In the penultimate stage, termed diplotene (Fig. 2, stage 4), the synaptonemal complex dissolves and a tetrad of four chromatids becomes visible. Additionally, the former crossing‐over points appear as chiasmata, which hold nonsister chromatids together. Diplotene is the first regulatory check‐ point in oocyte meiosis, which can result in a meiotic arrest in a variety of species, including cow and human. Prophase I is then completed by the diakinesis stage (Fig. 2, stage 5), which
Figure 2‐ Schematic representation of bovine oogenesis. The different phases are divided in 12 steps in which the first 5
belong to the prophase I stage. The arrests represented in the figure concern the two known meiotic arrests known in bovine during prophase I and metaphase II. A‐ First polar body B‐ Sperm cell C‐ Second polar body D‐ Male and female pronuclei.
is characterized by further chromatid condensation (Germinal Vesicle (GV)), breakdown of the nuclear membrane ‐ in a process termed Germinal Vesicle Breakdown (GVBD) ‐ and spindle formation 2,10.
Subsequently, the cells enter metaphase I, when the tetrad lines up along the metaphase plate and each chromosome of a homologous pair attaches to microtubule fibers from opposite poles. This is accomplished through the fusion of both sister chromatids’ kinetochore, thus providing each chromosome with only one functional kinetochore which attaches to opposite spindles. Simultaneously, sister chromatids attach to microtubules from the same pole (Fig. 2, stage 6) 2,10. At the onset of the next stage, the anaphase I, the chiasmata dissolves, allowing the homologous to move towards opposite spindle poles. The centrosome, however, does not divide, meaning that homologous chromosomes with sister chromatids move towards opposite sides (Fig. 2, stage 7) 2,10. Succeeding anaphase I is the
telophase I, at the end of meiosis I. During this reductional division, each daughter cell receives half the number of chromosomes, all consisting of two sister chromatids, culminating with an asymmetric cell division and the extrusion of the first polar body, which usually does not undergo the second meiotic division (Fig. 2, stage 8) 2,10.
The oocyte now enters the second division process, meiosis II. Starting the metaphase II, the chromosomes align at the metaphase plate and the kinetochores of sister chromatids attach to opposite microtubule spindle fibers. Most vertebrate oocytes arrest again at this stage, where they can remain for a long time until fertilization occurs (Fig. 2, stage 9) 2,10. Upon fertilization, the kinetochore of sister chromatids is disrupted and microtubule fibers can attach to each centrosome of each sister chromatid, therefore pulling them on opposite directions (Fig. 2, stage 10) 2,10. This stage is termed anaphase II and after it is completed, the telophase II takes place, where another asymmetrical division is complete and the oocyte is provided with the second polar body. The larger cell, the oocyte, occupies 95% of the cytoplasm and is now a functional female pronuclei (Fig. 2, stage 11) 2,10.
By the end of oocyte meiosis, the zygote is formed (Fig. 2, stage 12) and the two pronuclei, arising from the female and male progenitor, approach each other. Their membranes break down and DNA replication takes place as they form a common mitotic spindle and align in a common metaphase plate. A true diploid nucleus is first seen not in the zygote but at the 2‐cell stage, after the first of many mitotic divisions that will ultimately originate a full individual. Besides ascending from different genders, the pronuclei are not equivalent and if the zygote’s genetic material is derived solely from one parent, normal development will not take place. Different methylation patterns and epigenetic events
contribute to the difference in both genomes 2,11. Although interchange of the genome such as the one that occurs during meiosis is important for evolutionary success, it is simultaneously important to protect the germ cell genome from potentially precarious mutations, particularly since these would be carried out throughout generations.
Protecting germ cell’s integrity via Argonaute proteins
In addition to follicular and cytosolic, also the nuclear environment is vital for a proper oocyte formation. Germ cells exhibit an unforeseen diversity of RNA interference (RNAi) mechanisms that are caught up in many gene‐regulatory mechanisms, such as genome defense against viruses and transposable elements, developmental competence and silencing activity 12,13.
Small interference RNA (siRNA) is a 20‐30 base‐pair double stranded fragment 14 and one strand, the guide RNA, can be incorporated into the protein complex RISC (RNA‐Induced Silencing Complex). The nuclease component of RISC known as Argonaute (AGO) uses these small RNAs to select mRNA for degradation 12,14–16 and these proteins are central in RNA‐ mediated gene silencing processes.
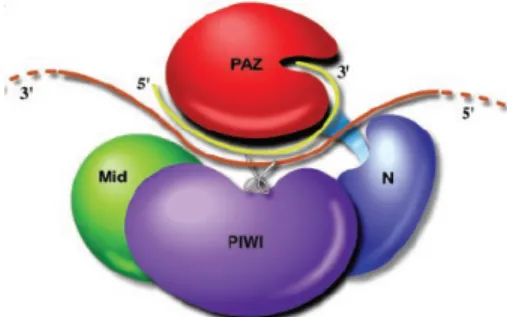
The AGO proteins consist of four domains (Fig. 3): the N‐terminal domain; the PAZ domain; the MID domain; and the PIWI domain, which has the important endonucleolytic activity in some AGOs 17,18. Hence, Argonaute, the signature component of RISC, seems to have slicer activity 16.
The PIWI subclass of Argonaute proteins has recently emerged in model organisms as a target for a special kind of small RNAs: the Piwi‐interacting RNAs (piRNAs) 19. The main currently known function of piRNAs is to silence transposable elements (TEs) in the germline, and this role is highly conserved across animal species. Association of piRNAs with PIWI proteins forms an active piRNA‐Induced Silencing Complex (piRISC) that can recognize and
Figure 3 ‐ Schematic representation of siRNA‐guided mRNA
cleavage. The siRNA (yellow) binds with its 3’ end in the PAZ cleft and the 5’ is predicted to bind near the other end of the cleft. The mRNA (brown) comes in between the N‐terminal and PAZ domains and out between the PAZ and middle domain. The active site in the PIWI domain (shown as scissors) cleaves the mRNA opposite the middle of the siRNA guide18
silence complementary mRNA targets. Genetic evidence has demonstrated that, in addition to piRNAs, PIWI proteins are necessary for TE silencing and have crucial roles in gonadal development 20,21.
Unraveling PIWI proteins in germ cells throughout species
Two Piwi family members in Drosophila, Aubergine and Piwi, were found to bind with piRNAs in ovaries 22,23. Transposition of telomeric retroelements is enhanced in aubergine mutants whereas piwi mutants mobilized the endogenous retrovirus gypsy 24. Both mutants demonstrated derepression of retrotransposons 23. Mutations in piwi and aubergine lead to severe defects in gametogenesis and piwi is essential in the asymmetrical division essential for the renewal of germline stem cells (GSCs). Indeed adult piwi mutant gonads lack any GSCs and gametes 25.
The Xenopus PIWI protein Xiwi is expressed in germ cells and piRNAs are the predominant class of small RNAs in Xenopus sperm and oocytes. These piRNAs can target retrotransposons in a similar manner as they do in Drosophila 26. In Zebrafish, the two PIWI homologs Ziwi and Zili were also characterized 27. Loss of Ziwi leads to germ cell loss, an increase in apoptosis of pre‐meiotic cells and abnormal sex determination. Similar to Ziwi, expression of Zili is also specific to testis and ovary and Zili mutants are agametic. However, the observed germ cell loss of is not caused by an increase in apoptosis but rather due to an inability to differentiate, probably instigated by defects in meiosis.
In mice, 3 Piwi‐like proteins have been identified, coded by 3 genes: Miwi (or Piwi‐like 1), Mili (Piwi‐like 2) and Miwi2 (Piwi‐like 4) 28. Interestingly, mutations in any of these 3 mouse PIWI genes lead to degradation of only the male germline, whereas in Drosophila and Zebrafish both the female and the male germline are affected after PIWI inactivation. Mili and Miwi2 mutants lead to the activation of retrotransposons in the male germline, arrest of gametogenesis and complete sterility in males 20,29,30. Miwi2 mutants showed increased
retrotransposon expression in testes, accompanied by decreased DNA methylation machinery. Only Mili expression has been identified in PGCs, despite the mutants’ apparent null effect in female germline. This gene was recently proposed to mediate DNA repair through chromatic relaxation in mice 31. These evidences, together with the implication of piRNA in de novo methylation of TE sequences during spermatogenesis 20, suggest that the role of PIWI proteins and piRNAs extends beyond post‐transcriptional silencing.
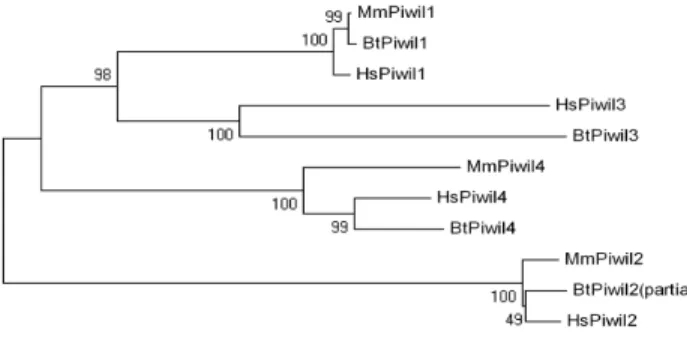
Sasaki and colleagues in 2003 identified a Piwi‐like gene in human cells 32 , Piwi‐like 3 (PIWIL3), which does not have a murine homolog, but is present in other mammals such as cattle (Fig. 4).
PIWIL3’s absence in the rodent genome jeopardizes the identification of its function by gene knockout technology. A proper model to study the functions of this Piwi‐like 3 gene is lacking in mammals, however the bovine reveals an easy and straightforward model to approach this challenge.
Disclosures in the PIWIL3 former research
Some aspects of PIWIL3 protein were addressed recently in bovine oocytes which provided new understandings into the function of this protein in mammalian germ cells. Bernard Roelen’s lab developed a custom‐made rabbit‐anti‐PIWIL3 polyclonal antibody and one year later we came to knowledge that it was not recognizing the protein (will be further discussed). Still, a piRNA‐like population is present in bovine oocytes (Ketting & Roelen, unpublished results) suggesting a unique function of PIWIL3 in oogenesis.
Preliminary results identified PIWIL3 mRNA in bovine ovaries, testes, cumulus cells and oocytes 33 and also in bovine oocytes during oocyte maturation but not during pre‐ implantation development (Fig. 5, A), suggesting a protein maternal supply already documented in Drosophila 33,34. With the PIWIL3 custom‐made antibody, co‐localization with the meiotic spindle was suggested, after GVBD stage (Fig. 5, B) which ultimately lead to the hypothesis that PIWIL3 has a role in oocyte maturation, especially during meiosis.
Figure 4 ‐ Homology between
mammalian PIWI proteins. Mm= Mus musculus (mouse), Hs= Homo sapiens (human); Bt= Bos taurus (cattle). (Ketting, R. Roelen, B., unpublished results).
The sea urchin PIWI homolog Seawi presents a similar localization pattern 35 and it has been suggested that Seawi associates with a complex of microtubule‐associated ribonucleoproteins (MT‐RNP), providing a route for the spatial segregation of factors that are important to establish morphogenetic gradients. Also in Xenopus oocytes, Xiwi protein and piRNAs associate with microtubules of the meiotic spindle 26. However no defects in spindle assembly were detected after depletion of Xiwi, suggesting that it is a mere passenger of the microtubule network.
The idea of PIWIL3 localization in the spindle was an unexpected and interesting result because no function of PIWI proteins in the meiotic spindle of mammalian oocytes had been documented. So far, Piwi‐like 3 seemed to have a particular purpose, especially during meiosis and we could speculate about the possibility of the protein and the piRNA being maternally accumulated, suggesting their active participation in the transport and localization of mRNA.
In the meantime, whilst studying PIWIL3, Transforming Acidic Coiled‐Coil protein 3 (TACC3) was identified in metaphase II oocytes (will be further discussed). Since the PIWIL3 results were compromised, we became interested in TACC3’s function during bovine oocyte development. A
B
Figure 5 ‐ mRNA quantification of PIWIl3 and PIWIL3 expression pattern. (A): Relative mRNA expression levels of PIWIL3 during oocyte maturation and early embryo development. X‐axis: developmental stages. Y‐axis: relative PIWIL3 amounts normalized to the reference gene SDHA. Error bars represent standard deviation. (B): merge channel of DAPI and Piwi‐like 3 protein on a metaphase II oocyte, with metaphase plate magnified. Scale bar = 20 μM 33Transforming Acidic Coiled‐Coil (TACC) Proteins
While trying to protect the genome against adversities that could endanger the upcoming generations, cell division is also one of the most important processes the cell needs to face. In order to accomplish it, the spindle needs to recruit microtubule fibers that are tightly regulated by a main microtubule‐organizing center (MTOC), the centrosome in animal cells
36. A family group of proteins known to interact with centrosomal activity are the TACC
proteins.
TACC proteins were initially identified as a group of proteins implicated in cancer and the first protein member was discovered whilst searching for amplified genomic regions in breast cancer 37. These highly acidic proteins have, in contrast to a very diverse N‐terminus, a predicted coiled‐coil domain on its C‐terminus, known as the TACC domain, the signature of this protein family that is thought to carry most of the functional properties 38,39. In Drosophila, the TACC domain provides the protein its correct localization on the spindle 39,40. Being present in different organisms such as Schizosaccharomyces pombe, Drosophila melanogaster, Caenorhabditis elegans, Xenopus laevis and mammals, TACC proteins have different names. In some species there is only one TACC protein such as in Schizosaccharomyces pombe (Alp7 also known as Mia1p), Drosophila melanogaster (D‐ TACC), Caenorhabditis elegans (TAC‐1) and Xenopus laevis (Maskin or xTACC3). On the other hand, mammals have three TACC proteins: TACC1, TACC2 (also known as AZU‐1 and ECTACC) and TACC3 (also known as AINT and ERIC1)39,41–43. TACC1 and TACC2 protein are implicated in breast cancer 37 while TACC3 is associated with multiple myeloma and several cancer lines 44,45 (http://www.proteinatlas.org/ENSG00000013810). TACC3 is expressed in relatively few adult tissues but it shows high levels in testis, ovary and in hematopoietic tissues. Together with TACC3‐deficient mice displaying embryonic lethality, a reduced cell number and mitotic defects, these data suggest an important role of TACC3 during early cell division 44,46,47.
Concerning the three mammalian TACC proteins, its expression pattern depends on the cell phase cycle. At the interphase stage, TACC2 associates with centrosomes whereas TACC1 and TACC3 are diffused both in the cytoplasm and in the nucleus, with TACC3 being to some extent concentrated in the nucleus 38,39. During mitosis, TACC1 and TACC2 are present in centrosomes while TACC3 is present in a more diffuse region around the centrosomes 38,48.
Besides its centrosomal localization, TACC proteins are also present on microtubules during cell division 49. C. elegans’ TAC‐1 and X. laevis’ Maskin are present in the spindle and Drosophila’s D‐TACC is associated with both spindle and astral microtubules (fibers not connected to kinetochores) (Fig. 6) 39. Regarding the protein’s role in the cell, all mutant phenotypes described to date seem to indicate microtubule rearrangement and instability. In C. elegans, embryos without TAC‐1 exhibit defects in pronuclear migration and a phenotype of both shorter and defective spindle in anaphase. However, despite the spindle‐positioning defects, embryos are still able to assembly microtubules, suggesting that TAC‐1 protein is not required for its formation 41.
Also in Drosophila melanogaster, mutants in D‐TACC show incomplete pronuclear separation 39. Most of the mutant embryos exhibit an arrest in the first mitotic division and those that can divide have abnormally short centrosomal microtubules, culminating in mitotic defects that lead to embryo lethality. In Xenopus, Maskin is not required for microtubule stability but is required for its anchoring to the centrosome. In embryos depleted with Maskin (xTACC3), the microtubule content was reduced in size and number and cells exhibited an abnormal spindle (Fig. 7) 40,50,51. It was also demonstrated that RNAi downregulation of TACC3 in HeLa cells leads to arrest at G1 checkpoint, prior to anaphase, with aberrant spindle morphology and severely misaligned chromosomes 40,52.
Overexpression studies of D‐TACC, Maskin and TACC3 show protein accumulation on the spindle poles leading also to an increase in microtubule length and number38. C. elegans TAC‐1 D. melanogaster D‐TACC X. laevis Maskin H. sapiens TACC3
Figure 6 ‐ Centrosome and spindle localization of TACC proteins in C. elegans, D. melanogaster, X. laevis
and Human. The upper row shows, in the first three organisms, overlay images with microtubules in green, DNA in blue and the corresponding TACC proteins in red. The lower row shows the distribution of each TACC protein alone49. H. sapiens column exhibits TACC protein in green and α‐tubulin in red, bar: 10 µm 48.
The rationale is that flaws in kinetochore‐microtubule attachment can lead to spindle checkpoint activation and TACC3 regulates chromosome alignment by ensuring both proper kinetochore microtubule attachment and spindle assembly checkpoint. 53
Phosphorylation and activation of TACC3 by Aurora‐A kinase
Recruitment of TACC3 proteins to microtubules is regulated by Aurora‐A kinase‐mediated phosphorylation and this function appears to be conserved in Human, Xenopus, C. elegans and Drosophila 41,48,54–56.
It has been proposed that Aurora‐A is vital during mitosis through phosphorylation of a series of substrates that promote diverse critical events to maintain cell integrity 57. During mitosis, Aurora‐A kinase associates with centrosomes and spindle, independently of microtubules and D‐TACC and mutations of Aurora‐A lead to metaphase arrest and decreased length of astral microtubules as well as prevention of D‐TACC centrosomal association 54,56. Aurora A interacts with TACC3, phosphorylating it and allowing its targeting to centrosomes 55. For Xenopus and Human TACC3, the exact phosphorylation sites have already been identified 40,48,55. Suppression of Aurora‐A by siRNA caused mitosis failure, with
incorrect separation of centriole pairs, chromosome misalignment on the metaphase plate and incomplete cytokinesis. Despite spindle abnormalities and unaligned chromosomes, some cells lacking functional Aurora‐A are still able to divide, however segregation defects in anaphase and chromatin bridges further develop into aneuploidies or tetraploidies which ultimately lead to cell death 57.
Figure 7 ‐ Depletion of Xenopus’ TACC3 (xTACC3): effects on the spindle. The upper row shows the control
situation and the lower one shows abnormal spindle and chromosome abnormalities in result if xTACC3 depletion with RNAi40.
Besides TACC3 vital interaction with Aurora‐A kinase, also its involvement with two other other proteins, ch‐TOG and clathrin, is essential to a proper complex assembly and function.
TACC3/ch‐TOG/clathrin complex stabilizes kinetochore fibers by inter‐
microtubule bridging
In higher organisms, the chromosome’s kinetochores are attached to the spindle via parallel microtubules, named kinetochore fibers (K‐fibres) 58. The microtubules of K‐fibers are connected by inter‐microtubule bridges that are thought to stabilize the fiber during chromosome movement 59.Clathrin interacts with these inter‐microtubule bridges in order to stabilize K‐fibers
60,61. Apart from being involved in coating vesicles during interphase, during cell division it
seems to target microtubule spindles 62. It acts as part of a complex involving two other proteins: TACC3 and Colonic‐Hepatic Tumour Overexpressed Gene (ch‐TOG), forming the TACC3/ch‐TOG/clathrin complex 61,63,64. ch‐TOG, or XMAP215, was identified as having an essential role in spindle organization in Human during mitosis 65. Recently, clathrin was identified as a binding protein for TACC3, and ch‐TOG was found to be associated with both clathrin and TACC3 66. Depletion of clathrin, ch‐TOG or TACC3 result always in a mitotic defected phenotype in different species, suggesting a consensus interaction across species
40,61,63,65.
Additionally, TACC3/ch‐TOG/clathrin bridges also protect the fiber from microtubule loss 67 and promote growth by antagonizing the mitotic centromere‐associated kinesin
(MCAK) that promotes microtubule depolymerization 55,68.
Despite recent progresses in TACC3 research during mitosis, its role in mammalian meiosis is relatively unknown. In mouse oocytes, TACC3 was identified during meiotic oocyte maturation and the phosphorylated form of TACC3 accumulated from GVBD to the metaphase II arrest. The phosphorylated protein was restricted to the spindle poles. The effect of TACC3 depletion by siRNA was (i) inhibition of polar body extrusion (ii) arrested meiosis I with spindle defects and (iii) lack of phosphorylated TACC3 labeling at the poles 69.
TACC3/ch‐TOG/clathrin complex activation, two hypotheses
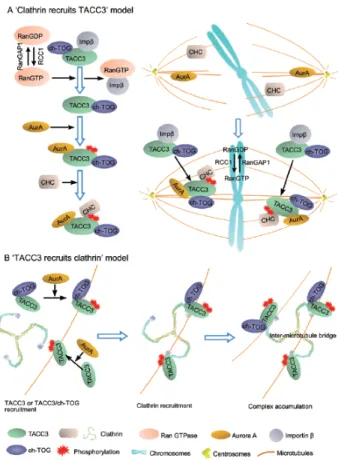
The correct order of recruitment of these three proteins to the complex is yet unknown, however two major ones were proposed during the last three years (Fig. 8).
In 2010, three teams 40,63,66 claimed that clathrin recruits phosphorylated TACC3 to spindle poles (Fig. 8, A), however other findings point to a different direction. In 2011, Booth et al. demonstrated that TACC3 and ch‐TOG bind to the spindle microtubules under Aurora‐ A regulation, followed by recruitment of clathrin to the microtubules forming complex with TACC3 or TACC3/ch‐TOG subcomplex (Fig. 8, B). Furthermore, it was also proposed that the complex is spindle‐specific 61. According to Booth et al., clathrin cannot bind microtubules while maskin (TACC3) and ch‐TOG can, therefore, clathrin would require an additional factor to bind these fibers, however no such factor was identified. Also, the majority of clathrin in mitotic cells is not restricted to the spindle whereas TACC3 and ch‐TOG are predominantly spindle‐associated. In agreement with Booth’s theory, (i) overexpression of GFP‐CHC (Clathrin Heavy Chain) did not cause more clathrin to accumulate at the spindle and did not influence TACC3 nor ch‐
Figure 8 ‐ Two models representing the recruitment of TACC3/ch‐TOG/clathrin complex to the spindle. (A) Clathrin recruits
TACC3: Aurora‐A kinase activates
TACC3 by phosphorylating it in two sites (Ser620 and Ser626 in Xenopus) and enables it to bind with clathrin to form the clathrin/TACC3 complex. Finally, the clathrin‐associated TACC3 is targeted to spindle poles and spindle microtubules for proper spindle assembly. Along with TACC3, ch‐TOG also targets to spindles to regulate spindle stability. (B) TACC3 recruits clathrin. First, TACC3 alone or
complexed with ch‐TOG is recruited to the spindle microtubules in a phosphorylation‐ dependent manner regulated by Aurora‐A. Then, clathrin is recruited to the spindle by TACC3 or TACC3/ch‐TOG. Finally, clathrin forms an inter‐microtubule bridge by interacting with additional TACC3 or TACC3/ch‐TOG on adjacent microtubules to stabilize the spindle microtubules. 89
TOG spindle localization (ii) overexpressing TACC3 increased the spindle location of ch‐TOG and clathrin and (iii) inhibiting TACC3 spindle recruitment via inhibition of Aurora‐A kinase or TACC3 mutation inhibited recruitment of clathrin to the spindle.
Taken together, recruitment of clathrin and TACC3 to the mitotic spindle remains controversial, however in unanimity it is suggested that this complex stabilizes kinetochore fibers by physically bridging between adjacent microtubules. In the light of these results, TACC3 has an important role in mitosis as a controlled adaptor that can integrate several other mitotic proteins into a complex on microtubules which holds kinetochore fibers together.
TACC3 function during mammalian meiosis has already been documented 69; however its role during bovine oocyte development is still blurred. Besides having clear implication in human cancer lines, understanding its role during cell‐division is crucial when the matter of mammalian oocyte developmental competence is addressed. The aim of this project will be to get insights into TACC3’s function during meiosis and early embryo development and ultimately propose a model for TACC3/ch‐TOG/clathrin complex regulation and recruitment in cattle, which may mimic the Human situation.
Material and methods
Oocyte collection and in vitro maturation
Bovine ovaries were obtained from a local slaughterhouse and transported in thermal flasks. Excess tissue was cut off and the ovaries were collected in flasks containing NaCl at 30ºC supplemented with penicillin/streptomycin (1ml/L). Cumulus oocyte complexes (COCs) were removed from the ovaries by suction of follicular fluid from antral follicles ranging 2‐8mm and only those with reasonable cumulus investment were selected under a dissecting microsope. The COCs were rinsed with medium B (Table 1) to avoid agglomeration. COCs were transferred to 4‐well plates, each containing 500µL of maturation medium (Table 1) or different concentrations of the inhibitor MLN8054 (by selleckchem®, 25 mM in DMSO) in maturation medium. Between the wells, 1mL of sterile water was added to prevent evaporation. After 22‐24 H of incubation at 39ºC with 5% CO2, the cultured oocytes were
either selected for in‐vitro fertilization or fixed in 4% paraformaldehyde.
In vitro fertilization and embryo culture
A straw of cryopreserved bull sperm was thawed in a 37ºC waterbath and the content layered on a Percoll gradient and centrifuged for 30 minutes at 700G. The sperm’s correct
Medium Ingredients
B MilliQ water, M199 with Earle's salts, HEPES and glutamine
C LAL water, M199 with Earle's salts and glutamine, NaHCO3
PHE Penicillamine, hypotaurine and epinephrine
Maturation Medium C, Fetal Calf Serum, Pen/Strep, recombinant FSH
Fertilization LAL water, NaCl, KCl, NaHCO3, Na2HPO4, Na pyruvate, phenolred, CaCl2.2H2O,
MgCl2.6H2O, Pen/Strep, BSA
RD MilliQ water NaCl, KCl, NaHCO3, Na2HPO4, Na Lactate, HEPES, phenolred,
CaCl2.2H2O, MgCl2.6H2O, Na Pyruvate, Pen/Strep, BSA
SOF A LAL water, NaCL, KCl, KH2PO4, Na Lactate, MgSO4.7H2O, NaHCO3, CaCl2.2H2O,
phenolred, Non‐essential aminoacid solution
SOF B LAL water, Pen/Strep, Na‐pyruvate, L‐glutamine, BSA
SOF 8ml SOF A : 2ml SOF B
concentration was determined by sampling a small volume of the sperm solution and counting the number of sperm heads in a Bürker Turk chamber. Taking into account which bull was used, the dilution factor was calculated and sperm was added to the fertilization medium (Table 1). The COCs and sperm were incubated for 22 hr at 39ºC, 5% CO2 after
which the presumptive zygotes were stripped of cumulus cells by vortexing for 3 min in RD medium (Table 1). Following denudation, the presumptive zygotes were transferred to a new 4‐well plate with Synthetic Oviductal Fluid (SOF medium, Table 1) and incubated at 39ºC, in a 5% CO2 and 7% O2 environment70. At day 5 after culture, cleaved embryos were
scored and transferred to fresh SOF medium and at day 8, embryo development was determined.
Immunocytochemistry
Sections of bovine testis, bovine ovaries and human testis (provided by University Medical Center Utrecht) were used. To de‐paraffinise sections, slides were immersed 2x5min in xylene and 2x5min in 100% ethanol followed by 20 min incubation in methanol‐3% H2O2 to
block endogenous peroxidase and reduce non‐specific background staining. Samples were then rehydrated in 2 minutes series of 96/80/70% ethanol, rinsed shortly in distilled H2O.
For antigen retrieval, sodium‐citrate buffer (0.01M, pH 6.0) was heated until boiling point and samples were boiled for 3 min. After a 30 min cool‐down, sections were washed in TBST (0.05% Tween) for 2x5min, blocked with 5% NGS (Normal Goat Serum) in TBS for 30 minutes at 37 ºC and left to incubate overnight with the first antibody (1:100) in 2% NGS in TBS at room temperature. The three samples were each stained with Human anti‐PIWIL3 antibody (ab77088), anti‐PIWIL3 antibody (ab93709) and IgG for control purposes (Table 2). On the second day, after washing in TBS (0.05 M, pH 7.6) on the shaking table, incubation with the second antibody (1:200) took place for 40 min at 37ºC followed by another washing step. The samples were then incubated with ABC complex reagent (Vector elite kit 10µl A + 10µl B + 980 µl TBS) for 30 min at RT (ABC made 30 min prior to use). Sections were then washed in TBS, incubated with DAB (diaminobenzidinetetrahydrochloride, mutagenic, photosensitive) in TBS for 5‐10 min and washed again in distilled H2O. For counterstaining, slides were
stained with haematoxylin for 10 seconds and then washed thoroughly with H2O. Series of
70/80/96% ethanol were executed (2x 10 sec each) followed by xylene (2x2min). Lastly, the samples were embedded in DEPEX.
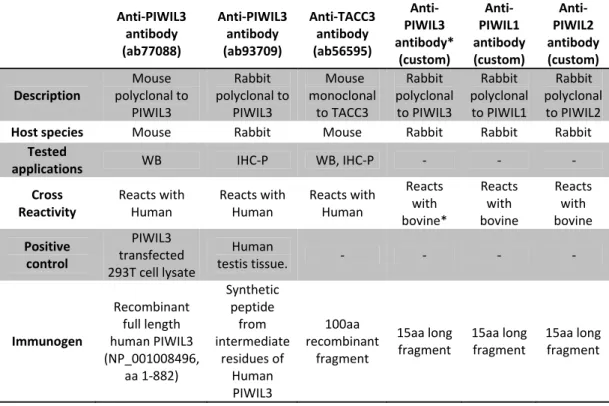
Anti‐PIWIL3 antibody (ab77088) Anti‐PIWIL3 antibody (ab93709) Anti‐TACC3 antibody (ab56595) Anti‐ PIWIL3 antibody* (custom) Anti‐ PIWIL1 antibody (custom) Anti‐ PIWIL2 antibody (custom) Description Mouse polyclonal to PIWIL3 Rabbit polyclonal to PIWIL3 Mouse monoclonal to TACC3 Rabbit polyclonal to PIWIL3 Rabbit polyclonal to PIWIL1 Rabbit polyclonal to PIWIL2
Host species Mouse Rabbit Mouse Rabbit Rabbit Rabbit
Tested
applications WB IHC‐P WB, IHC‐P ‐ ‐ ‐
Cross Reactivity Reacts with Human Reacts with Human Reacts with Human Reacts with bovine* Reacts with bovine Reacts with bovine Positive control PIWIL3 transfected 293T cell lysate Human testis tissue. ‐ ‐ ‐ ‐ Immunogen Recombinant full length human PIWIL3 (NP_001008496, aa 1‐882) Synthetic peptide from intermediate residues of Human PIWIL3 100aa recombinant fragment 15aa long fragment 15aa long fragment 15aa long fragment
Immunofluorescence
Oocytes were stripped of cumulus cells before staining and fixed for a minimum of 15 min in 4% paraformaldehyde (PFA). After fixation the oocytes were briefly washed with 0.1% Triton X‐100 and 10% FCS in PBS (PBST) and permeabilized for 30 min using 0.5% Triton X‐100 and PBST. The oocytes were subsequently blocked for 1h in PBST and incubated with the primary antibody overnight at 4ºC. For the immunofluorescence, all antibodies described in Table 2 were used. The oocytes were washed three times in PBST (15 min) followed by incubation in the secondary antibody for 1 h in the dark. After several washes, the oocytes were stained with 4',6‐diamidino‐2‐phenylindole (DAPI, 5min) and mounted on a slide with Vectashield and isolated with Vaseline.For α‐tubulin staining oocytes were incubated first for 30‐60 min in microtubule stabilizing solution71 at 37 ºC and then fixed in 4% PFA. After fixation, oocytes were washed in PBS with 0.1% (v/v) Tween‐20, incubated 5min in PBS with 2% (v/v) goat serum (Sigma‐ G6767) followed by 60 min incubation with mouse‐anti‐tubulin primary antibody at 37 ºC. After washing, the oocytes were incubated in PBS + serum for 1 hour at 37 ºC, washed and
Table 2 ‐ Detailed description of the antibodies used. The (*) regarding the bovine PIWIL3 symbolizes the predicted details on the
onset of this project. All the custom antibodies are thought not to be specific and further tests have to be done to address this question.
incubated with the second antibody for 1 h. For microfilament staining, an additional incubation of 30 min with phalloidin was performed. After DNA staining with DAPI, the oocytes were mounted as mentioned before. Slides were examined with Leyca® Confocal laser microscopy. Images were analyzed with ImageJ® software and Adobe Illustrator®.
Western blotting
Cells were collected and frozen in ‐80ºC after which they were lysed with sample buffer 4x diluted in RIPA buffer (table 3) and 1% protease inhibitor cocktail (table 3). Afterwards the lysates were boiled at 100 ºC for 5 min and briefly centrifuged. The samples were resolved by an SDS‐PAGE 8% running gel and 5% stacking gel (Table 3), while sponges, filter paper and membrane were left for 10 min at 4 ºC in Blotting buffer. The proteins were then blotted on nitrocellulose membranes (Trans‐Blot®, Bio‐Rad Laboratories) at 100V (±180mA) during 1H. The membranes were incubated with blocking buffer (Table 3) for 1h under gentle shaking. The membranes were incubated with mouse‐anti‐TACC3 (Table 2) diluted in blocking buffer (1:1000) O/N at 4 ºC. After washing in TBS‐Tween (TBS with 0.05% Tween‐20) (3x5min) the membranes were incubated with goat‐anti‐Mouse (sc‐2005, Santa Cruz Biotechnology Inc) (1:5000) for 1H at RT; washed with TBS‐Tween and lastly with TBS. Finally the membranes were placed between a plastic sheet under the SuperSignal® Chemiluminescent Substrate (West Dura Extended Duration Substrate, Thermo scientific) consisting of 600µL of Horseradish peroxidase (HRP) and phosphatase (PO) (1:1). The membrane was exposed to X‐ray film (Fuji®) and the film was developed.After developing the film, the membrane was stripped in stripping buffer for 5 min at RT and the blot was incubated for 1H at 37 ºC with Goat‐anti‐β‐actin (1:5000, Santa Cruz Biotechnology®, sc‐47778) and for 1H at RT with Rabbit‐anti‐Goat (1:10000, Santa Cruz Biotechnology®,sc‐2768) just as described above.
Aurora‐A inhibition experiment
Oocytes were matured as described in table 4 and the small molecule MLN8054 (Selleckchem®, dissolved in DMSO) was added to the medium with the purpose if inhibiting Aurora‐A function 72. MLN8054 stock (250 µM) and DMSO (2µl/200µl maturation medium) were used to achieve a maturation medium with an increased drug concentration. Because the drug was dissolved in DMSO, a control with no DMSO neither drug (normal IVM) was added.
Different groups were used in different sets of experiments. When the percentage of blastocyst formation was evaluated, it was scored on the 8th day of maturation, being relative to the amount of embryos cleaved on day five. This percentage is thus referring to the blastocysts that were both properly matured and fertilized, providing embryo development information. Buffers Composition Running gel SDS‐ PAGE 8% gel (for 10ml) 4.6ml H2O; 2.7ml 30% acryl‐bisacrylamide mix; 2.5ml 1.5 M Tris (PH=8.8); 0.1ml 10% SDS, 0.1ml 10% ammonium persulfate; 0.008ml TEMED Stocking gel 5% (for 10 ml) 6.8ml H2O; 1.7ml 30% acryl‐bisacrylamide mix; 1.25ml 1.5 M Tris (PH=8.8); 0.1ml 10% SDS, 0.1ml 10% ammonium persulfate; 0.01 TEMED Blocking buffer 5% (w/v) non‐fat dry milk powder in TBS‐Tween RIPA buffer 1% Nonidet P‐40, 0.5% sodium deoxycholate, 0.1% SDS, 0.004% sodium azide, 1% PMSF solution, 1% protease inhibitor solution, 1% sodium orthovanadate solution in TBS (Santa Cruz Biotechnology, Santa Cruz, CA, USA) Halt Protease Inhibitor Cocktail Protease and Phosphatase Inhibitor Cocktail (100X), EDTA Solution, 0.5M (100X), 1mL
Oocytes Maturation medium MLN8054 DMSO
Matured with 0 µM MLN8054 490 µl 0 10 µl Matured with 1.25 µM MLN8054 490 µl 2.5 µl 7.5 µl Matured with 2.5 µM MLN8054 490 µl 5 µl 5 µl Matured with 5 µM MLN8054 490 µl 10 µl 0 Matured under normal IVM procedure 500 µl 0 0 Denuded and matured under normal IVM 500 µl 0 0 Denuded and matured with 2.5 µM MLN8054 490 µl 5 µl 5 µl Table 3 ‐ Detailed description of the Buffers used on the Western Blot. Table 4 ‐ Experimental design for the MLN8054 experiment. Maturation medium was previously described in table 1.