Identification of photoperception and light signal transduction pathways
in citrus
Vera Quecini
Instituto Agronômico de Campinas, Centro de Pesquisa e Desenvolvimento de Recursos Genéticos,
Campinas, SP, Brazil.
Abstract
Studies employing model species have elucidated several aspects of photoperception and light signal transduction that control plant development. However, the information available for economically important crops is scarce. Citrus genome databases of expressed sequence tags (EST) were investigated in order to identify genes coding for func-tionally characterized proteins responsible for light-regulated developmental control in model plants. Approximately 176,200 EST sequences from 53 libraries were queried and allbona fide and putative photoreceptor gene families were found in citrus species. We have identified 53 orthologs for several families of transcriptional regulators and cy-toplasmic proteins mediating photoreceptor-induced responses although some importantArabidopsis phytochrome-and cryptochrome-signaling components are absent from citrus sequence databases. The main gene families re-sponsible for phototropin-mediated signal transduction were present in citrus transcriptome, including general regu-latory factors (14-3-3 proteins), scaffolding elements and auxin-responsive transcription factors and transporters. A working model of light perception, signal transduction and response-eliciting in citrus is proposed based on the iden-tified key components. These results demonstrate the power of comparative genomics between model systems and economically important crop species to elucidate several aspects of plant physiology and metabolism.
Key words:cryptochrome, data mining, light signaling, phototropin, phytochrome.
Received: July 21, 2006; Accepted: February 8, 2007.
Introduction
Plant development is highly plastic, allowing the envi-ronment to exert tight control over the transitions between genetic developmental programs in order to maximize growth and reproduction (Meyerowitz, 2002). Light pro-vides spatial and temporal information to regulate plant de-velopment throughout its life cycle: from germination and seedling establishment to the onset of the reproductive stage (Schäfer and Nagy, 2006). Plants are able to detect environ-mental light direction, duration, fluency and wavelength due to a complex system of photoreceptor molecules: the blue (B) and ultraviolet-A (UV-A) 320-500 nm light-sensing cryptochromes (cry) and phototropins (phot) (Banerjee and Batschauer, 2005) and the red (R)/far red (FR) 600-750 nm phytochrome (phy) receptors (Schepenset al., 2004). Re-cently, a novel family of putative B photoreceptors has been described inArabidopsis thaliana: the ZEITLUPE (ZTL)/ Flavin-binding Kelch repeat F-box protein (FKF1)/LOV Kelch Protein (LKP2) family (Nelsonet al., 2000; Schultzet
al., 2001; Somerset al., 2000). ZTL/ FKF1/LKP2 are in-volved in the circadian clock mechanism and photoperiodic flowering response (Imaizumiet al., 2005).
In higher plants, photoreceptor co-action is a com-mon theme and the pathways may function synergistically, antagonistically and additively to control several develop-mental responses (Schäfer and Nagy, 2006). Moreover, plant photoreceptors operate in concert with numerous other signaling systems; including phytohormones, carbo-hydrate-mediated, temperature, gravity and the endoge-nous clock transduction pathways (Halliday and Fankhauser, 2003; Chenet al., 2004; Heggie and Halliday, 2005). The molecular mechanisms involved in light-induced signal transduction include the following: light-re-gulated sub-cellular localization of the photoreceptors (Guoet al., 1999; Nagy and Schäfer, 2002; Chen et al., 2005; Kong et al., 2006); a large reorganization of the transcriptional program (Casal and Yanovsky, 2005; Fran-klinet al., 2005) and light-regulated proteolytic degrada-tion of several photoreceptors and signaling components (Höcker, 2005; Huq, 2006).
Studies employing model plant species have demon-strated that photoperception, the signal transduction path-www.sbg.org.br
Send correspondence to Vera Quecini. Centro de Pesquisa e Desenvolvimento de Recursos Genéticos, Instituto Agronômico de Campinas, Caixa Postal 28, 13001-970 Campinas, SP, Brazil. E-mail: [email protected].
ways and the responses elicited by light form a complex interconnected network rather than a linear pathway (Chen et al., 2004; Quecini and Liscum, 2006). The diversity of light responsiveness observed in plants has arisen from the elaboration, combination and re-arrangement of a basic repertoire of mechanisms responsible for light-mediated developmental regulation, allowing adaptation to a wide range of climatic and latitudinal regions. Comparative genomics has provided tools to access the genetic bases of this diversity in non-model species using bioinformatics, which increases the fundamental knowledge of gene inter-actions and permits analyses of the functional significance of proteinsin silico(e.g.Santelli and Siviero, 2001; Souza et al., 2001; Hechtet al., 2005).
Virtually all information about light-regulated devel-opment in woody plants comes from studies withPopulus, a temperate deciduous perennial (Zhu and Coleman, 2001; Olsen and Juntilla, 2002). Adaptive traits in temperate peren-nial woody plants involve an integrated physiological re-sponse directed at plant survival and nutrient storage over the winter period and are greatly dependent of photoreceptor-mediated perception of seasonal progression (Thomas and Vince-Prue, 1997). Surprisingly, recent evidence has dem-onstrated that light is also the main factor triggering the tran-sition between vegetative to reproductive developmental stages of trees in Equatorial regions (Borchertet al., 2005). The effects of light on developmental processes in citrus and other neotropical tree species have been described in several situations, although without approaching the molecular as-pects of the metabolism (Steppeet al., 2006; Chen L-Set al., 2005; Ravehet al., 2003; Tornéet al., 2001). Mutant studies employing transgenically generated plants have demon-strated variable extents of functional conservation in the genes responsible for developmental control between citrus and model species (Penaet al., 2001; Pillitteriet al., 2004). However, the detailed functional characterization of individ-ual genes is a limiting factor in the study of tree species, and new strategies should be devised for the study of gene func-tion (Groover and Robischon, 2006).
Based on evidence of extensive conservation in photo-perception and light signal transduction in angiosperms, this work aimed to identify the characterized components of these pathways in citrus. Our results have demonstrated that a large portion of the genes involved in light responses from model species are present in citrus and that they share extensive pro-tein sequence conservation in several regions, including func-tionally characterized domains. These results demonstrate the potential use of comparative genomic tools to elucidate physi-ological and metabolic processes in crop species.
Material and Methods
Database searches and alignments
Homologs ofArabidopsis thaliana and other model species photoperception and light signal transduction genes
were identified in BLAST searches (Altschulet al., 1997) against EST sequences from the citrus index databases at CitEST, consisting of approximately 176,200 ESTs ob-tained from the sequencing of 53 libraries. Data validation was performed by tBLASTx and tBLASTn searches using BLOSUM80 scoring matrix of the retrieved hits against the databases at NCBI (National Center for Biotechnology In-formation) built inside the CitEST project. Sequences fail-ing to retrieve the original bait sequence were eliminated from the projects. The resulting alignments were filtered by a threshold e-value of 1e-15 and the validated hits were fur-ther analyzed according to functional domain description. Validated sequences were translated and protein (deduced amino acid) alignments were performed using ClustalX (Thompsonet al., 1997). When necessary, alignments were manually adjusted using Lasergene MegAlign (DNASTAR, Madison, WI, USA).
Motif analysis andin silicocharacterization
The identified homologs were investigated for the presence and sequence conservation of recognizable func-tional domains described in several protein analysis and gene function databases (European Bioinformatics Insti-tute-European Molecular Biology Laboratory - EMBL-EBI; Expert Protein Analysis System - ExPaSy from the Swiss Institute of Bioinformatics - SIB; Protein Families database - Pfam).
Phylogenetic analysis
The functionality of citrus genes in comparison to the characterized homologs was assessed by genetic distance and phylogenetic studies. Phylogenetic analyses were per-formed using distance and parsimony methods in the soft-ware PAUP* 4.0b10, using the softsoft-ware default parame-ters. Re-sampling bootstrap trees containing 1000 random samples were constructed using PSIGNFIT software. Mod-ular functional domains were employed for genetic dis-tance studies for genes previously described as having di-vergent regions and conserved blocks.
Results and Discussion
transduction components and light-regulated transcription factors, whereas the remaining 22 are similar to phot signal-ing partners (Figure 1). Homologs from the transcriptional regulators PAT1 (PHYTOCHROME A SIGNAL TRANS-DUCTION 1, AT5G48150) (Bolle et al., 2000), HRB1
(HYPERSENSITIVE TO RED AND BLUE 1,
AT1G02340) (Kanget al., 2005), OBP3 (OBF4-BINDING PROTEIN 3, AT3G55370) (Wardet al., 2005) and from the novel signaling components FHL (FHY1-LIKE, AT5G02200) (Zhou et al., 2005) and SRR1 (SENSITI-VITY TO RED LIGHT REDUCED 1, AT5G59560) (Staigeret al., 2003) were absent from CitEST databases. However, the existence of citrus orthologs cannot be ruled out at this point due to the restricted coverage of the
trans-criptome analysis, the expression levels and patterns of these genes and the post-translational modifications re-quired to generate functional components. InArabidopsis and rice, several genes involved in light signaling have been identified by biochemical, forward and reverse ge-netic assays (Zhouet al., 2005; Keveiet al., 2006)
Photoreceptor-related genes
Citrus EST database contains homologs of members of all families of plant photoreceptors: namely, the phyto-chrome, cryptochrome and phototropin families (Table S1, Figure 2, Figure 3, Figure 4). In higher plants, phy are re-sponsible for the control of major developmental processes, such as seed, germination, de-etiolation, shade avoidance, Figure 1- Functional classification of citrus transcripts associated to photoperception and light signal transduction based on gene ontology (GO)
Figure 2- Domain structure, phylogenetic analyses and alignment of the predicted amino acid sequence of phytochrome family in citrus.
Neigh-bor-joining trees for citrus and tree species deduced amino acid andArabidopsisfull length sequences aligned with ClustalX are shown. Bootstrap values are indicated above each branch. Dark and light gray shading indicate sequence identity and similarity, respectively. At,Arabidopsis thaliana; C Number, contig number; Cit, citrus; CS, Citrus sinensis; CR, Citrus reticulata; FAD, flavin adenosine diphosphate; FR, far-red light; HKRD, histidine
Figure 3- Domain structure, phylogenetic analyses and alignment of the predicted amino acid sequence of the cryptochrome family in citrus. Neigh-bor-joining trees for citrus deduced amino acid andArabidopsisfull length sequences aligned with ClustalX are shown. Bootstrap values are indicated
floral induction and entrainment of the circadian clock (Chenet al., 2004). Four EST singlets sharing sequence similarities to phy family genes were identified; three inC. sinensisand one inC. reticulatagenome (Table S1). Two ofC. sinensisESTs show higher levels of sequence similar-ity toPHYAgenes, whereas the remaining ones are related to PHYB-type of sequences. Interestingly, the PHYB homologs in citrus appear to be more distantly related from thePopulus PHYB1andPHYB2sequences than from theA. thaliana PHYEgene (Figure 2). The branching in thePHYB
genes is a relatively recent event and is absent from many plant species, includingArabidopsis(Mathews, 2006). The partial nature of the sequences prevent us from speculating whether citrus genome has a singlePHYBgene or two, like the current model woody plantPopulus.
The cryptochrome family is represented in the citrus genome by two ESTs corresponding toArabidopsis CRY1 and five sequences similar to CRY-DASH (Table S1, Fig-ure 3). Although, all the identified sequences share similar-ities to higher plant cry sequences, for the majority of them Figure 4- Domain structure, phylogenetic analyses and alignment of the predicted amino acid sequence of phototropin and zeitlupe in citrus.A.
phototropin family.B.zeitlupe family. Neighbor-joining trees for citrus deduced amino acid andArabidopsisfull length sequences aligned with ClustalX are shown. Bootstrap values are indicated above each branch. Dark and light gray shading indicate sequence identity and similarity, respectively. At,
Arabidopsis thaliana; B, blue light; C Number, contig number; CA,Citrus aurantium;Cit, citrus; CS,Citrus sinensis; CR,Citrus reticulata; FAD, flavin adenosine diphosphate; LOV, light, oxygen, voltage subtype of PAS; PT,Poncirus trifoliata; R, red light; STKD, serine/threonine kinase domain; UV-A,
(five), the deduced amino acid sequence identity is re-stricted to the photolyase-like domain (Figure 3), indicating that these genes may function as photolyases rather than bona fideB photoreceptors. The C-terminal extension, es-sential for cry1 function inArabidopsis, is conserved inC. sinensisEST and in C. reticulataEST contig (Table S1, Figure 3). Cryptochromes are mainly responsible for de-etiolation under blue light inArabidopsis, including control of transcriptional regulation, inhibition of hypocotyls growth, promotion of cotyledons expansion, and synthesis of several non-photoreceptor pigments, such as chlorophyll and anthocyanins (Li and Yang, 2006). In addition, this class of photoreceptor acts in coordination with phy to reset the circadian clock and to control the transition to flowering (Yanovsky and Kay, 2002). At least two of the identified sequences (CS00-C1-100-038-A12.CT and C13-CA) are likely to code for functional cry family members in citrus.
InArabidopsis, the phototropin photoreceptor family consists of two closely related members that share almost 60% protein identity. In the genome of citrus species, an EST contig whose deduced amino acid sequence shows 71% identity to PHOT2 and four cDNAs with sequences approximately 50% identical to PHOT1 and PHOT2, were identified (Figure 4A, Table S1). The overall sequence con-servation between citrus and other species PHOT proteins is high, including at the N-terminal LOV domains, essential for chromofore binding and protein function in Arabidopsis,reviewed in Quecini and Liscum (2006). The remaining identified ESTs and EST contigs present high levels of sequence identity restricted to the C-terminal serine/threonine kinase and thus, may not perform photo-perception-related functions. InArabidopsisand rice, phot family controls a specific sub-set of physiological pro-cesses, including phototropic stem curvature, stomata opening control and chloroplast relocation (Quecini and Liscum, 2006). Only recently, phots have been demon-strated to be involved in B-mediated seedling de-etiolation (Foltaet al., 2003; Takemyiaet al., 2005). InArabidopsis, phot1 and phot2 have specialized and overlapping roles, phot1 being the most important photoreceptor sensing di-rectional B under low fluence rates and phot2 responsible for high light responses (Briggs and Christie, 2002). The presence of multiple PHOT-like sequences in citrus ge-nome suggests that such a fluence rate-specific role might occur.
Recently, a three-member family of putative photo-receptors has been characterized in Arabidopsis; the ZTL/FKF1/LKP2 family (Somers, 2001). It is represented by three distinct sequences in citrus genome databases: one highly similar to FKF1 with lower homology to ZTL and two sharing sequence similarity to FKF1 and LKP2 (Table S1, Figure 4B).ArabidopsisZTL/FKF1/LKP2 proteins are characterized by the presence of a flavin-binding LOV do-main at the protein N-terminal, an F-box dodo-main and a stretch of Kelch repeats, providing a direct link between
light perception and ubiquitin-mediated protein degrada-tion. The family has been functionally associated to the endogenous time-keeping mechanism and the control of photoperiodic flowering time (Imaizumi et al., 2003; Imaizumiet al., 2005). InArabidopsis, ZTL and FKF1 are thought to be components of an Skp1-Cullin-F-box (SCF) E3 ubiquitin ligase complex (Vierstra, 2003). ZTL has been implicated in the ubiquitin-mediated proteolytic degrada-tion of the clock component TOC1 (Máset al., 2003) and in the regulation of developmental responses to R, possibly through its interaction with phyB (Keveiet al., 2006), while FKF1 has been demonstrated to control the levels of the photoperiod-sensingCONSTANS(CO) gene via degrada-tion of its transcripdegrada-tional repressor, a DOF type transcrip-tion factor, CDF1 (Imaizumiet al., 2003; Imaizumiet al., 2005).Thus, ZTL post-translationally regulates TOC1 lev-els and FKF1 controls dailyCOexpression in part by de-grading CDF1. Citrus FKF1/ZTL- and LKP2-like EST contigs are highly conserved at the F-box and Kelch repeats domains (Figure 4B), suggesting a function in the prote-lolytic degradation of circadian-clock associated factors.
Phy and cry signal transduction components
Phytochrome responses are associated with changes in gene expression (Casal and Yanovsky, 2005) and mem-bers of several transcription factor families are required for phy signaling or are early targets of phy-mediated re-sponses. In citrus, 22 EST contigs corresponding to the Arabidopsistranscriptional regulators and nuclear factors involved in phy and cry light signaling were identified, along with 17 transcripts associated to light-mediated pro-teolysis and seven transcripts similar to several signaling events of light signaling, such as Ca2+-binding and post-translational protein modification (Figure 1, Table S2).
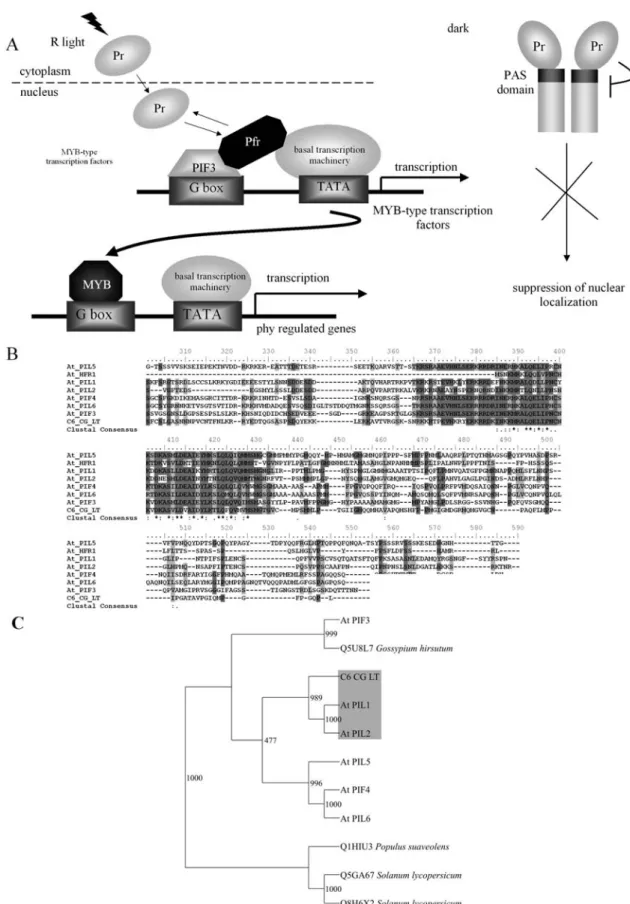
In Arabidopsis, the PIF (PHYTOCHROME-INTE-RACTING FACTOR) and PIL (PIF-LIKE) family of bHLH (basic Helix-Loop-Helix) transcriptional regulators, which includes HFR1/REP1/RSF1 (LONG HYPOCOTYL
IN FAR RED 1/REDUCED PHYTOCHROME
un-identified factors to regulate transcription of a master set of regulators, such asCCA1(Wang and Tobin, 1998),LHY1 (Schafferet al., 1998),TOC1andCO(Harmeret al., 2000; Teppermanet al., 2001); and (ii) these regulators then con-trol the transcription of genes encoding functions necessary for the terminal steps of the signaling cascade. Interest-ingly, in citrus genome databases, thePIF/PIL/HFRfamily is represented by a single EST contig fromC. aurantifolia and C. latifolia, displaying higher identity to PIF4 and HFR1 protein (Table S2, Figure 5). Another two highly similar EST contigs (55.5% deduced amino acid sequence identity) showed moderate (15.0 to 17.5%) and low (7.5 to 2.9%) identity to HFR1 and PIF/PIL gene products, respec-tively. The functional significance of these transcripts as PIF/PIL/HFR-like light-induced transcriptional regulators remains unclear. The absence of PIF-like transcripts inC. sinensisandC. reticulata transcriptomes, which together correspond to approximately 72% of CitEST database, is noteworthy given their importance in light-mediated re-sponses inArabidopsis.
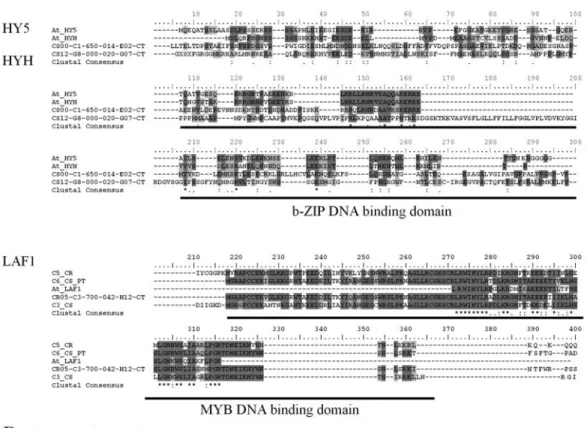
Several basic domain/zinc finger (Zn finger) and MYB-type factors function as downstream convergent tar-gets of phy and cry signaling inArabidopsis, independently of G-box photoreceptor binding (Oyama et al., 1997; Chattopadhyayet al., 1998; Ballesteroset al., 2001). In cit-rus species transcriptome, six cDNAs corresponding to this class of transcriptional regulators were present: namely, one HY5 (LONG HYPOCOTYL 5), one HYH (HY5-HOMOLOGOUS) and four LAF1 (LONG AFTER FAR RED 1) homologs (Table S2, Figure 6A). Down-regulation of these signaling pathways occurs when phyA and the transcription factors, HY5 and LAF1, are degraded in a light-dependent fashion by the proteasome in a mecha-nism that involves COP1 (CONSTITUTIVELY PHOTO-MORPHOGENIC 1) (Saijoet al., 2003; Seoet al., 2004; Janget al., 2005). The COP9 signalosome (CSN) is also in-volved in HY5 degradation (Penget al., 2001). Moreover, SPA1 (SUPRESSOR OF PHYA-105 MUTATION) and the other members of the SPA family regulate the ubi-quitin-ligase activity of COP1 (Saijoet al., 2003; Seoet al., 2003). In citrus EST databases, COP1, other components of the CSN and a small group of SPA-like transcripts were identified, suggesting the existence of a similar mechanism of phy-mediated signal desensitization route (Table S2, Figure 6B).
Photoreceptor-initiated signaling pathways also con-sist of cytosolic components in Arabidopsis and other model species. General transduction pathways, such as G-protein, Ca+2-calmodulin and protein phosphorylation cascades have been demonstrated to take part in light-triggered signaling (Bowleret al., 1994). Homologs of sev-eral Ca+2-binding and protein phosphorylation factors in phy- and cry-initiated signal transduction were identified in citrus transcriptome (Table S2). Phosphorylation may be
responsible for the fine tuning of light signal transduction at several checkpoints, including the degradation of active phyA mediated by the ubiquitin/26S proteasome pathway; the interaction of light-signaling positive factors with COP1, an E3 ubiquitin-protein ligase functioning as a nega-tive regulator of photomorphogenesis (Seoet al., 2004), re-ducing the affinity of phosphorylated phyA to its signaling partners (Kim et al., 2004) and controlling the phospho-rylation level and, consequently, the signaling activity, of phy (Ryuet al., 2005). Extensive functional conservation in angiosperms phosphorylation and post-translational pro-tein modification suggests that the transcripts identified in citrus are involved in the proteolytic degradation of light-signaling components.
Phot signal transduction components
Figure 5- Light-signaling associated bHLH transcriptional regulator family in citrus.A.schematic representation of phytochrome-regulated PIF3 transcriptional activation.B.alignment of the bHLH DNA-binding domainPIF/PIL family inArabidopsisand citrus. C. phylogenetic analysis of plant
Figure 6- Proteolysis-mediated photomorphogenesis control pathway in citrus.A.alignment of positive photomorphogenesis regulator families
HY5/HYH and LAF1.B.alignment of negative photormorphogenesis regulator families EID1 and SPA1. Dark and light gray shading indicate sequence identity and similarity, respectively. At,Arabidopsis thaliana; bZIP, basic leucine zipper; C, contig; CS,Citrus sinensis; CR,Citrus reticulata; PT,
Concluding Remarks
This preliminary survey of citrus photoperception-associated genes has provided useful information for fur-ther studies of light developmental control in these species. It has allowed the identification of conserved members of light-triggered signaling in a non-model species and the elaboration of a work model frame for light perception and signaling in citrus (Figure 7). These prospects are particu-larly attractive considering the range of economically im-portant physiological processes of citrus that are regulated by light, including secondary metabolism regulation and shading responses. An immediate goal of plant genomics is to transfer knowledge between model and crop species, al-lowing a better understanding of the mechanisms underly-ing several aspects of plant physiology. Thus, genomic and functional information can be integrated into the accumu-lated knowledge of citrus genetics and physiology to ad-vance basic and applied research. These studies will help to elucidate the molecular basis of developmental plasticity and to understand how environmental factors modulate plant development and the expression of phenotypic char-acters. The results obtained provide a new perspective on several aspects of light-regulated physiological processes
in citrus, such as de-etiolation, seedling establishment and shade-avoidance response.
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W and Lipman DJ (1997) Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nu-cleic Acids Res 25:3389-3402.
Ballesteros ML, Bolle C, Lois LM, Moore JM, Vielle-Calzada JP, Grossniklaus U and Chua NH (2001) LAF1, a MYB tran-scription activator for phytochrome A signaling. Genes Dev 15:2613-2625.
Banerjee R and Batschauer A (2005) Plant blue-light receptors. Planta 220:498-502.
Bolle C, Koncz C and Chua N-H (2000) PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Genes Dev 14:1269-1278.
Borchert R, Renner SS, Calle Z, Navarrete D, Tye A, Gautier L, Spichiger R and von Hildebrand P (2005) Photoperiodic in-duction of synchronous flowering near the Equator Nature 433:627-629.
Bowler C, Neuhaus G, Yamagata H and Chua N-H (1994). Cyclic GMP and calcium mediate phytochrome phototransduction. Cell 77:73-81.
Briggs WR and Christie JM (2002) Phototropins 1 and 2: Versa-tile plant blue-light receptors. Trends Plant Sci 7:204-210. Casal JJ and Yanovsky MJ (2005) Regulation of gene expression
by light. Int J Dev Biol 49:501-511.
Chattopadhyay S, Ang LH, Puente P, Deng XW and Wei N (1998)
ArabidopsisbZIP protein HY5 directly interacts with light-responsive promoters in mediating light control of gene ex-pression. Plant Cell 10:673-683.
Chen L-S, Qi Y-P and Liu H-C (2005) Effects of aluminum on light energy utilization and photoprotective systems in citrus leaves. Ann Bot 96:35-41.
Chen M, Chory J and Fankhauser C (2004) Light signal trans-duction in higher plants. Annu Rev Genet 38:87-117. Chen M, Tao Y, Lim J, Shaw A and Chory J (2005) Regulation of
phytochrome B nuclear localization through light-depen-dent unmasking of nuclear-localization signals. Curr Biol 15:637-642.
Fairchild CD, Schumaker MA and Quail PH (2000)HFR1 en-codes an atypical bHLH protein that acts in phytochrome A signal transduction. Genes Dev 14:2377-2391.
Folta KM, Lieg EJ, Durham T and Spalding EP (2003) Primary in-hibition of hypocotyl growth and phototropism depend dif-ferently on phototropin-mediated increases in cytoplasmic calcium induced by blue light. Plant Physiol 133:1464-1470.
Franklin KA, Larner VS and Whitelam GC (2005) The signal transducing photoreceptors of plants. Int J Dev Biol 49:653-664.
Friml J (2003) Auxin transport - Shaping the plant. Cur Op Plant Biol 6:7-12.
Friml J, Wisniewska J, Benkova E, Mendgen K and Palme K (2002). Lateral relocation of auxin efflux regulator PIN3 mediates tropism inArabidopsis. Nature 415:806-809.
Fuglsang AT, Borch J, Bych K, Jahn TP, Roepstorff P and Palmgren MG (2003) The binding site for regulatory 14-3-3 protein in plant plasma membrane H+-ATPase: Involvement Figure 7- Schematic overview of the photosensory signaling pathways in
of a region promoting phosphorylation-independent interac-tion in addiinterac-tion to the phosphorylainterac-tion-dependent C terminal end. J Biol Chem 278:42266-42272.
Groover A and Robischon M (2006) Developmental mechanisms regulating secondary growth in woody plants. Curr Opin Plant Biol 9:55-58.
Guo H, Duong H, Ma N and Lin C (1999) TheArabidopsisblue light receptor cryptochrome 2 is a nuclear protein regulated by a blue light-dependent post-transcriptional mechanism. Plant J 19:279-287.
Halliday KJ and Fankhauser C (2003) Phytochrome-hormonal signalling networks. Plant Phytologist 157:449-463. Harmer SL, Hogenesch JB, Straume M, Chang HS, Han B, Zhu T,
Wang X, Kreps JA and Kay SA (2000) Orchestrated tran-scription of key pathways inArabidopsisby the circadian clock. Science 290:2110-2113.
Harper RM, Stowe-Evans EL, Luesse DR, Muto H, Tatematsu K, Watahiki MK, Yamamoto K and Liscum E (2000) The
NPH4locus encodes the auxin response factor ARF7, a con-ditional regulator of differential growth in aerial
Arabidopsistissue. Plant Cell 12:757-770.
Hecht V, Foucher F, Ferrándiz C, Macknight R, Navarro C, Vardy ME, Ellis N, Beltrán JP, Rameau C and Weller JL (2005) Conservation ofArabidopsisflowering genes in model le-gumes. Plant Physiol 137:1420-1434.
Heggie L and Halliday KJ (2005) The highs and lows of plant life: Temperature and light interactions in development. Int J Dev Biol 49:675-687.
Höcker U (2005) Regulated proteolysis in light signaling. Curr Opin Plant Biol 8:469-476.
Huq E (2006) Degradation of negative regulators: A common theme in hormone and light signaling networks? Trends Plant Sci 11:4-7.
Huq E and Quail PH (2002) PIF4, a phytochrome-interacting bHLH factor, functions as a negative regulator of phyto-chrome B signaling inArabidopsis. EMBO J 21:2441-2450.
Imaizumi T, Schultz TF, Harmon FG, Ho LA and Kay SA (2005) FKF1 F-box protein mediates cyclic degradation of a repres-sor ofCONSTANSinArabidopsis. Science 309:293-297. Imaizumi T, Tran HG, Swartz TE, Briggs WR and Kay SA (2003)
FKF1 is essential for photoperiodic-specific light signalling inArabidopsis. Nature 426:302-306.
Jang IC, Yang JY, Seo HS and Chua N-H (2005) HFR1 is targeted by COP1 E3 ligase for post-translational proteolysis during phytochrome A signaling. Genes Dev 19:593-602. Kang X, Chong J and Ni M (2005) HYPERSENSITIVE TO RED
AND BLUE 1, a ZZ-type zinc finger protein, regulates phytochrome B-mediated red and cryptochrome-mediated blue light responses. Plant Cell 17:822-835.
Kevei E, Gyula P, Hall A, Kozma-Bognar L, Kim WY, Eriksson ME, Toth R, Hanano S, Feher B, Southern MM,et al.(2006)
Forward genetic analysis of the circadian clock separates the multiple functions of ZEITLUPE. Plant Physiol 140:933-945.
Kim JI, Shen Y, Han YJ, Park JE, Kirchenbauer D, Soh MS, Nagy F, Schafer E and Song PS (2004) Phytochrome phosphoryl-ation modulates light signaling by influencing the protein-protein interaction. Plant Cell 16:2629-2640.
Kinoshita T and Shimazaki K (2002) Biochemical evidence for the requirement of 14-3-3 protein binding in activation of
the guard-cell plasma membrane H+-ATPase by blue light. Plant Cell Phys 43:1359-1365.
Kong SG, Suzuki T, Tamura K, Mochizuki N, Hara-Nishimura I and Nagatani A (2006) Blue light-induced association of phototropin 2 with the Golgi apparatus. Plant J 45:994-1005. Li QH and Yang HQ (2006) Cryptochrome signaling in plants.
Photochem Photobiol 83:94-101.
Martinéz-Garcia JF, Huq E and Quail PH (2000) Direct targeting of light signals to a promoter element-bound transcription factor. Science 288:859-863.
Más P, Kim WY, Somers DE and Kay SA (2003) Targeted degra-dation of TOC1 by ZTL modulates circadian function in
Arabidopsis thaliana. Nature 426:567-570.
Mathews S (2006) Phytochrome-mediated development in land plants: Red light sensing evolves to meet the challenges of changing light environments. Mol Ecol 15:3483-3503. Meyerowitz EM (2002) Plants compared to animals: The broadest
comparative study of development. Science 295:1482-1485. Nagy F and Schäfer E (2002) Phytochromes control photomor-phogenesis by differentially regulated, interacting signaling pathways in higher plants. Annu Rev Plant Biol 53:329-355. Nelson DC, Lasswell J, Rogg LE, Cohen MA and Bartel B (2000).
FKF1, a clock-controlled gene that regulates the transition to flowering inArabidopsis. Cell 101:331-340.
Ni M, Tepperman JM and Quail PH (1999) Binding of phyto-chrome B to its nuclear signalling partner PIF3 is reversibly induced by light. Nature 400:781-784.
Olsen JE and Juntilla O (2002) Far red end-of-day treatment re-stores wild type-like plant length in hybrid aspen over-expressing phytochrome A. Physiol Plant 115:448-457. Oyama T, Shimura Y and Okada K (1997) TheArabidopsisHY5
gene encodes a bZIP protein that regulates stimulus-induced development of root and hypocotyl. Genes Dev 11:2983-2995.
Pena L, Martin-Trillo M, Juarez J, Pina JA, Navarro L and Marti-nez-Zapater JM (2001) Constitutive expression of
ArabidopsisLEAFY or APETALA1 genes in citrus reduces their generation time. Nature Biotechnol 19:263-267. Peng Z, Serino G and Deng XW (2001) Molecular
characteriza-tion of subunit 6 of the COP9 signalosome and its role in multifaceted developmental processes inArabidopsis. Plant
Cell 13:2393-2407.
Pillitteri LJ, Lovatt CJ and Walling LL (2004) Isolation and char-acterization of a TERMINAL FLOWER homolog and its correlation with juvenility in citrus. Plant Physiol 135:1540-1551.
Quail PH (2002) Photosensory perception and signalling in plant cells: New paradigms? Curr Opin Cell Biol 14:180-188. Quecini V and Liscum E (2006) Signal transduction in blue
light-mediated responses. In: Schäfer E and Nagy F (eds) Photomorphogenesis in Plants and Bacteria: Function and Signal Transduction Mechanisms. 3rd edition. Springer Ac-ademic Publishers, Dordrecht, pp 305-327.
Raveh E, Cohen S, Raz T, Yakir D, Grava A and Goldschmidt EE (2003) Increased growth of young citrus trees under reduced radiation load in a semi-arid climate. J Exp Bot 54:365-373. Ryu JS, Kim J-I, Kunkel T, Kim BC, Cho DS, Hong SH, Kim S-H,
Saijo Y, Sullivan JA, Wang H, Yang J, Shen Y, Rubio V, Ma L, Hoecker U and Deng X-W (2003) The COP1-SPA1 interac-tion defines a critical step in phytochrome A-mediated regu-lation of HY5 activity. Genes Dev 17:2642-2647.
Santelli RV and Siviero F (2001) A search for homologues of plant photoreceptor genes and their signaling partners in the sugarcane expressed sequence tag (Sucest) database. Genet-ics Mol Biol 24:49-53.
Schäfer E and Nagy F (2006) Photomorphogenesis in Plants and Bacteria: Function and Signal Transduction Mechanisms. Springer Academic Publishers, Dordrecht, 662 pp. Schaffer R, Ramsay N, Samach A, Corden S, Putterill J, Carré IA
and Coupland G (1998) Thelate elongated hypocotyl
muta-tion of Arabidopsis disrupts circadian rhythms and the
photoperiodic control of flowering. Cell 93:1219-1229. Schepens I, Duek P and Fankhauser C (2004)
Phytochrome-mediated light signaling in Arabidopsis. Cur Biol 7:564-569.
Schultz TF, Kiyosue T, Yanovsky M, Wada M and Kay SA (2001) A role for LKP2 in the circadian clock of
Arabidopsis. Plant Cell 13:2659-2670.
Seo HS, Watanabe E, Tokutomi S, Nagatani A and Chua N-H (2004) Photoreceptor ubiquitination by COP1 E3 ligase de-sensitizes phytochrome A signaling. Genes Dev 18:617-622.
Seo HS, Yang JY, Ishikawa M, Bolle C, Ballesteros ML and Chua N-H (2003) LAF1 ubiquitination by COP1 controls photo-morphogenesis and is stimulated by SPA1. Nature 423:995-999.
Somers DE (2001) Clock-associated genes in Arabidopsis: A family affair. Philos Trans R Soc Lond B Biol Sci 356:1745-1753.
Somers DE, Schultz TF, Milnamow M and Kay SA (2000) ZEITLUPE encodes a novel clock-associated PAS protein fromArabidopsis. Cell 101:319-329.
Souza GM, Simões ACQ, Oliveira KC, Garay HM, Fiorini LC, Gomes F dos S, Nishiyama-Junior MY and Silva AM da (2001) The sugarcane signal transduction (SUCAST) cata-logue: Prospecting signal transduction in sugarcane. Genet Mol Biol 24:25-34.
Staiger D, Allenbach L, Salathia N, Fiechter V, Davis SJ, Millar AJ, Chory J and Fankhauser C (2003) The Arabidopsis SRR1gene mediates phyB signaling and is required for nor-mal circadian clock function. Genes Dev 17:256-268. Steppe K, Dzikiti S, Lemeur R and Milford JA (2006) Stomatal
oscillations in orange trees under natural climatic condi-tions. Ann Bot 97:831-835.
Takemyia A, Inoue S, Doi M, Kinoshita T and Shimazaki K (2005) Phototropins promote plant growth in response to blue light in low light environments. Plant Cell 17:1120-1127.
Tatematsu K, Kumagai S, Muto H, Sato A, Watahiki MK, Harper RM, Liscum E and Yamamoto KT (2004)MASSUGU2
en-codes Aux/IAA19, an auxin-regulated protein that functions together with the transcriptional activator NPH4/ARF7 to regulate differential growth responses of hypocotyl and for-mation of lateral roots inArabidopsis thaliana. Plant Cell 16:379-393.
Tepperman JM, Zhu T, Chang HS, Wang X and Quail PH (2001) Multiple transcription-factor genes are early targets of
phytochrome A signaling. Proc Natl Acad Sci USA 98:9437-9442.
Thomas B and Vince-Prue D (1997) Photoperiodism in plants. 2nd edition. Academic Press, San Diego, 428 pp.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F and Higgins DG (1997) The CLUSTALX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876-4882.
Torné JM, Moysset L, Santos M and Simon E (2001) Effects of light quality on somatic embryogenesis in Araujia sericifera.Phys Plant 111:405-411.
Vierstra RD (2003) The ubiquitin/26S proteasome pathway, the complex last chapter in the life of many plant proteins. Trends Plant Sci 8:135-142.
Wang ZY and Tobin EM (1998) Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene dis-rupts circadian rhythms and suppresses its own expression. Cell 93:1207-1217.
Ward JM, Cufr CA, Denzel MA and Neff MM (2005) The DOF transcription factor OBP3 modulates phytochrome and cryptochrome signaling in Arabidopsis. Plant Cell 17:475-485.
Wilmoth JC, Wang S, Tiwari SB, Joshi AD, Hagen G, Guilfoyle TJ, Alonso JM, Ecker JR and Reed JW (2005) NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lat-eral root formation. Plant J 43:118-130.
Yanovsky MJ and Kay SA (2002) Molecular basis of seasonal time measurement inArabidopsis. Nature 419:308-312. Zhou Q, Hare PD, Yang SW, Zeidler M, Huang LF and Chua N-H
(2005) FHL is required for full phytochrome A signaling and shares overlapping functions with FHY1. Plant J 43:356-370.
Zhu B and Coleman GD (2001) Phytochrome-mediated photo-period perception, shoot growth, glutamine, calcium, and protein phosphorylation influence the activity of the poplar bark storage protein gene promoter (bspa). Plant Physiol 126:342-351.
Internet Resources
Citrus Biotechnology Laboratory, http://citest.centrodecitricultu ra.br (September 13, 2006)
Cluster v.2.11 Software, http://rana.lbl.gov/EisenSoftware.htm. DNASTAR Lasergene Software, http://www.dnastar.com/web/
index.php.
European Bioinformatics Institute-European Molecular Biology Laboratory (EMBL-EBI), www.ebi.ac.uk/interpro/ (Sep-tember 04, 2006).
Expert Protein Analysis System (ExPaSy), http://www.expasy. org/prosite/ and http://www.us.expasy.org/sprot/ (October 05, 2006).
Gene Ontology (GO), http://www.godatabase.org/cgi-bin/amigo/ go.cgi (October 23, 2006).
PAUP* 4.0b10 Software, http://paup.csit.fsu.edu/.
Protein Families (Pfam), http://www.sanger.ac.uk/Software/ Pfam/ (October 15, 2006).
PSIGNFIT Software, http://www.bootstrap-software.org/. The Institute for Genomic Research (TIGR)Arabidopsis thaliana
Tree View v.1.6 Software, http://rana.lbl.gov/EisenSoftware.htm.
Supplementary Material
The following online material is available for this ar-ticle:
Table S1
Table S2 Table S3
Supplemental References Figure S1
This material is available as part of the online article from http://www.scielo.br/gmb.
Namea Gene ESTb %c e value biological process References
CRY1 AT4G08920 C10-CR (2)
CS00-C1-100-038-A12-CT
23.3 74.6
3e-91 1e-106
FAD binding domain, DNA photolyase, B photoreceptor, photomorphogenesis,
circadian clock entrainment, photoperiodic responses
Cashmore et al., 1999
CRY-DASH AT5G24850 C2-CR/CS (13)
C3-CS (4) C11-CS (3) C13-CA (2) PT11-C1-901-056-H09-CT 23.5 22.2 21.2 64.1 12.7 1e-93 1e-89 2e-87 1e-106 1e-62
Cryptochrome family, putative B photoreceptor, transcriptional regulator in
Synechocystis
Brudler et al., 2003, Kleine et al., 2003
PHOT1 AT3G45780 C8- CS (5)
CS00-C3-701-104-B03-CT CS00-C3-702-030-D01-CT CR05-C3-701-091-E08-CT CS00-C1-100-062-A04-UV PT11-C1-900-012-G11-CT 19.8 50.0 50.0 18.0 37.8 17.0 1e-56 4e-84 7e-54 6e-53 1e-53 2e-45
LOV1 and LOV2 domain, serine-threonine kinase domain, B photoreceptor, phototropism, chloroplast
movement, stomata opening control
Huala et al., 1997
PHOT2 C12-CS/PT (17)
C7-CS (7) C9-CS (2) C1-CR/CS (2) C4-CR/CS (2) CS00-C3-701-104-B03-CT CS00-C1-100-124-G03-CT 71.1 33.4 38.1 27.4 17.5 47.5 42.1 7e-94 5e-69 3e-63 4e-59 3e-38 4e-84 6e-74
phototropin family Jarillo et al., 2001
PHYA AT1G09570 CS00-C3-701-101-C11-CT
CS00-C3-705-056-G06-CT
58.3 34.7
6e-86 4e-81
PAS1, PAS2, chromophore binding domain, HKL domain, R/FR
photoreceptor
Sharrock and Quail, 1989
PHYB AT2G18790 CR05-C1-102-036-H07-CT
CS12-G8-000-003-D03-CT
15.3 8.1
3e-75 3e-55
phytochrome family Reed et al., 1993
FKF1 AT1G68050 C5-CS (2)
C6-CR/CS (3)
76.3 19.3
1e-142 1e-46
Kelch repeats, F-box domain, LOV domain, putative photoreceptor, photoperiodic flowering control,
circadian clock
Nelson et al., 2000
LKP2 AT2G18915 CS00-C3-702-030-D01-CT 22.0 1e-76 putative photoreceptor, circadian clock Schultz et al., 2001
ZTL AT5G57360 C5-CS 29.7 8e-71 putative photoreceptor, circadian clock Somers et al., 2000 a
Gene name abbreviations: CRY: cryptochrome; DASH: Drosophila, Arabidopsis, Synechocistis, human; PHOT: phototropin; PHY: phytochrome, FKF1: F-box, Kelch repeat, Flavin-binding protein1; LKP2: LOV domain, Kelch repeat protein2; ZTL: zeitlupe.
b
C: contig, CA: Citrus aurantium, CG: Citrus aurantifolia, CR: Citrus reticulata, CS: Citrus sinensis, LT: Citrus latifolia, PT: Poncirus trifoliata, (number of reads). c
Identity percentage at the amino acid level. d
Name Gene EST % e value biological process Reference ATHB2 AT4G16780 C6-CS/PT (23)
CS00-C2-003-056-F03-CT
40.1 27.5
3e-51 2e-34
homeobox leucine zipper, shade avoidance response
Carabelli et al., 1996
COP1 AT2G32950 C5-CS/CG (5) C2-CS (2) CA26-C1-002-001-E08-CT 81.9 65.1 40.4 1e-89 3e-78 9e-61
E3 ubiquitin ligase, Zn finger and RING finger domains, proteolysis
Osterlund et al., 2000, Seo et al., 2004
COP8/FUS4 /FUS8
AT5G42970 CS00-C3-702-101-D09-CT CR05-C1-102-033-G01-CT CA26-C1-002-037-H04-CT
2e-75 2e-72 7e-69
subunit 4 of COP9 signalosome complex, subunit of the 19S regulatory particle of the
26S proteasome
Serino et al., 1999
COP9/FUS7 AT4G14110 C1-CS (3) LT33-C1-003-023-F05-CT
21.9 28.0
3e-66 2e-56
COP9 signalosome subunit, identical to cDNA CSN complex subunit 8 (CSN8)
Dohmann et al., 2005
COP10/FUS9 AT3G13550 C2-CR/CS (6) C3-CS/PT (23) C4-CR/CS (8) CR05-C1-102-060-B12-CT 64.8 39.0 39.0 38.5 5e-69 5e-41 1e-40 1e-39
ubiquitin-conjugating enzyme (COP10), proteolysis
Yanagawa et al., 2004
COP11/FUS6 AT3G61140 C1-CS/PT(2) 16.9 2e-86 COP9 signalosome complex subunit 1 / CSN complex subunit 1 (CSN1) / COP11 protein
(COP11) / FUSCA protein (FUS6)
Kang et al., 2000
EID1 AT4G02440 C1-CS/PT (5) 22.0 7e-51 Cyclin-like F-box protein, protein degradation, photomorphogenesis
Marroco et al., 2006
FAR1 AT4G15090 C9-PT (2) CS00-C1-100-058-F05-CT
37.3 32.7
1e-29 5e-35
FAR1 family, transposase-like domain Hudson et al., 1999
FHY3 AT3G22170 C9-PT (2) CS00-C1-100-058-F05-CT
34.2 24.6
1e-29 1e-33
FAR1 family, transpose-like domain Wang and Deng, 2002, Lin and Wang, 2004
FHY1 AT2G37680 CR05-C1-100-082-A02-CT CS00-C1-100-124-B05-CT CR05-C3-700-106-G10-CT CS00-C1-102-029-H04-CT 68.3 58.7 58.3 18.3 2e-71 2e-71 2e-71 2e-70
no recognizable domain, phyA-mediated photomorphogenesis
Shen et al., 2005a
HAF2 AT3G19040 C3-CA/CR (2) 47.5 2e-56 TATA-binding protein-associated factor TAF1 (TAFII250), bromodomain, ubiquitin
domain, histone acetyltransferase activity
Bertrand et al., 2005
HFR1 AT1G02340 C6-CG/LT (2) 29.4 4e-32 transcriptional regulator, bHLH domain, de-etiolation
Duek and Fankhauser, 2003
HY5 AT5G11260 CS00-C1-650-014-E02-CT 17.9 1e-35 transcription regulator, bZIP DNA binding domain, photomorphogenesis
Chattopadhyay et al., 1998
HYH AT3G17609 CS12-G8-000-020-G07-CT 16.1 2e-42 transcriptional regulator, bZIP DNA binding motif
Holm et al., 2002
LAF1 AT4G25560 C6-CS/PT (2) C3-CS (2) C5-CR (2) 73.9 67.4 63.0 2e-78 1e-68 6e-62
MYB transcription factor, R2R3 group, de-etiolation
C2-CS (2 45.3 1e-46 protein, histidine kinase, phy- and auxin-mediated signal transduction
Choi et al, 2005
PAP1 AT1G56650 C3-CS (2) CR05-C3-700-042-H12-CT CR05-C3-700-004-E04-EU PT11-C1-900-084-F06-CT 28.6 28.2 25.4 24.6 5e-56 2e-58 1e-55 1e-49
transcriptional regulator, auxin responsive, anthocyanin biosynthesis
Teng et al., 2005
PFT1 AT1G25540 C1-CG/CS (2) C2-CS/CR (6)
28.4 65.4
1e-97 1e-148
von Willebrand factor type A (VWF-A), glutamine-rich C-terminal, flowering time
Cerdán and Chory, 2003
PIF3 AT1G09530 C6-CG/LT (2) 25.5 4e-27 transcriptional regulator, bHLH domain, photomorphogenesis
Ni et al., 1998, Ni et al., 1999
PIF4 AT2G43010 C6-CG/LT (2) 24.5 1e-22 PIF family, transcriptional regulator, bHLH domain, de-etiolation (cell expansion)
Huq and Quail, 2002
PKS1 AT2G02950 C1-CR/CS (2) 20.2 1e-18 no recognizable domain, phytochrome kinase substrate,
Lariguet et al., 2003
PP7 AT5G63870 C3-CS/CR/PT (3)
C12-CS (11) CS13-C1-001-008-C12-CT PT11-C9-005-041-C05-CT 29.6 23.5 25.3 23.9 2e-55 7e-45 9e-46 1e-42 metallo-phosphoesterase motif, serine/threonine specific protein phosphatases
signature, de-etiolation
Møller et al., 2003
RAP1 / ATMYC2 AT1G32640 C2-CR/CS (2) CR05-C3-701-055-H07-CT
25.8 39.5
1e-39 4e-42
MYC-related transcriptional activator, bHLH leucine zipper motif, photomorphogenesis.
Heim et al., 2003
RFI2 AT2G47700 C1-CS (3) 34.5 4e-39 zinc finger (C3HC4-type RING finger) family protein, photomorphogenesis
Chen and Ni, 2006
SPA1 AT2G46340 C7-CR (3) C5-CR/CS/PT (3) CS13-C1-001-017-G02-CT 32.2 25.6 26.8 1e-114 2e-31 4e-31
proteolysis targeting, WD-repeat domain, light-regulated proteolysis
Höcker et al., 1999, Laubinger et al., 2004
SHB1 AT4G25350 C1-LT/CS (3) PT11-C1-900-042-H02-CT
27.8 17.1
2e-53 2e-25
EXS domain, SPX domain, photomorphogenesis under B
Kang and Ni, 2006
SUB1 AT4G08810 C3-CG/CS/PT (11) PT11-C1-901-057-B05-CT
37.0 22.4
2e-64 5e-44
Ca+2-binding protein, de-etiolation Guo et al., 2001
aGene name abbreviations: ATHB: Arabidopsis thaliana homeobox; ATMYC: Arabidopsis thaliana MYC-type; COP: constitutively photomorphogenic ; EID: Eimpfindlicher Im Dunkelroten
Licht; FAR: far-red impaired response; FHY : far-red elongated hypocotyl ; FUS: Fusca; HAF: histone acetylation factor; HFR : long hypocotyl in FR light ; HY: long hypocotyl; HYH :
HY5-homologue; LAF: long after far red; NDPK: nucleotide diphosphate protein kinase; PAP: production of anthocyanin pigment; PAT: phytochrome A-signal transduction ; PFT:
phytochrome and flowering time; PIF: phytochrome-interacting factor; PP: protein phosphatase ; RAP:ethylene response factor subfamily B-4 of ERF/AP2 transcription factor family; RFI:red and far red insensitive ; SPA: suppressor of phytochrome A-105; SHB: short hypocotyl under blue; SUB: short under blue.
bC: contig, CA: Citrus aurantium, CG: Citrus aurantifolia, CR: Citrus reticulata, CS: Citrus sinensis, LT: Citrus latifolia, PT: Poncirus trifoliata, (number of reads). cIdentity percentage at the amino acid level;
dFunctional domains abbreviations: ATP: adenosine triphosphate; bHLH: basic helix-loop-helix; bZIP: basic Zipper; FAD: flavina adenosine dinucleotide; HKL: histidine kinase-like; LOV:
Name
Gene
EST
%
e value
biological process
Reference
ARF7/
NPH4
AT5G20730
C4-CR
dC1-CS/CR
eCS00-C3-705-071-D01-CT
55.1
41.5
47.8
1e-89
3e-62
5e-93
B3 DNA binding domain,
AUX/IAA family, auxin-regulated
transcription
Harper
et al.
,
2000
GRF1
and
GRF
family
AT4G09000
C2-CS
fC4-CS
gC9-CA/PT
hC10-CA/CS
iCS00-C1-101-018-E05-CT
CS00-C3-700-041-F04-CT
71.1
64.3
51.6
52.2
39.5
29.9
1e-113
1e-104
4e-99
1e-102
1e-101
2e-97
regulatory factor1-G-box factor
14-3-3 homolog isoform family,
signal transduction (scaffolding)
Ferl, 2004
NPH3
and
NRL
family
AT5G64330
C5-CS
jC6-CR
kC8-CR/CS
lCS00-C3-701-013-G03-CT
CS00-C3-701-060-E12-CT
PT11-C1-900-042-G09-CT
PT11-C1-900-043-E10-CT
47.1
21.6
19.3
25.2
35.4
39.3
33.3
9e-71
3e-59
4e-59
6e-82
5e-96
1e-78
4e-67
plant-specific NPH3 domain,
BTB/POZ domain, signal
transduction (scaffolding)
Motchoulski and
Liscum, 1999,
Haga
et al.
, 2005
PIN1
and
PIN
family
AT1G73590
C3-CG/CS
mC11-CA/CS
nCS00-C3-700-106-C03-CT
23.4
24.6
38.7
2e-75
3e-89
4e-97
auxin efflux carrier, tropic
responses
Blakeslee
et al.
,
2004
RPT2
AT2G30520
C1-CS
oC7-CR
pCS00-C3-702-027-G06-CT
27.6
70.8
18.3
2e-65
1e-132
4e-42
plant-specific NPH3 domain,
BTB/POZ domain, signal
transduction (scaffolding)
Inada
et al.
, 2004
a
Gene name abbreviations: ARF: auxin-responsive factor ; GRF:general regulatory factor (14-3-3 protein) ; NPH: non-phototropic hypocotyl ; PIN: pin-formed ; RPT: root phototropism.
b
C: contig, CA: Citrus aurantium, CG: Citrus aurantifolia, CR: Citrus reticulata, CS: Citrus sinensis, LT: Citrus latifolia, PT: Poncirus trifoliata, (number of reads).
c
Identity percentage at the amino acid level.
d