Regulation of the
OsNHX1
Gene Expression:
Identification and Characterization of Novel
Transcription Factors
Diego Melo Almeida
Dissertation presented to obtain the Ph.D degree in Biochemistry
Instituto de Tecnologia Química e Biológica António Xavier | Universidade Nova de
Lisboa
Work performed at:
Supervisors:
Dr. Nelson José Madeira Saibo
Head of the Plant Gene Regulation laboratory (ITQB NOVA) Principal Investigator
Prof. Dr. M. Margarida Oliveira
Head of GPlantS Unit (ITQB NOVA)
Associate Professor with Habilitation (Agregação) at ITQB NOVA
Dr. Glenn B. Gregorio
Crop Breeding Manager at Sweet and Waxy Corn. San Rafael, Philippines. Former senior scientist and plant breeder at International Rice Research Institute (IRRI). Los Baños, Philippines.
Genomics of Plant Stress Unit
Instituto de Tecnologia Química e
Biológica António Xavier
Universidade Nova de Lisboa
Av. Da República
2780-157 Oeiras
Portugal
International Rice Research
Institute
Plant Breeding, Genetics, and
Biotechnology Division
Los Baños
Equipped with his five senses, man explores the universe around
him and calls the adventure Science.
Edwin Powell Hubble
IX
ACKNOWLEDGMENTS
To Nelson Saibo for being my supervisor, for taking care of me during these eight
years, for always pushing me to do and be the best, making me grow. For
teaching me when I was a newly-graduated student. Most importantly, for
open my eyes to science and show what science means.
To Prof. Margarida Oliveira for having accepted me at her lab (already crowded at
that time!), for all precious minutes you devoted me, for always teaching me
more than science. Above all, for show me what it means to love what we
do.
To Glenn Gregorio for having accepted me at IRRI, delighting my world, making
understand that our research produces more than scientific papers. We are
able to shape people´s life.
To Ana Paula Santos for being so positive, for helping me with all micro-world and
for all advices.
To Titi for all great lab conversations, for making our lab a better fun place to
work, for helping me solving the challenges of my work.
To Duarte Figueiredo for all wonderful moments we had inside and outside the
lab, for helping me with everything during my first times in lab, for the great
sense of humor.
To Muffy for all our great weekends/afterhours crazy projects, for all great talks at
23 PM, and for helping me with million things during these years.
To Sónia Negrão for helping me enter in the salt stress world, for all the precious
tips and talks.
To Pedro Barros for all support, wonderful talks, advices, for always ask about my
work, and for the huge help into the bioinformatics world.
To Lena for all support with the physiological work, for always taking care of us,
for being so honest, direct, and for having a big heart.
To Miguel Costa for all enormous support with the physiological work, new ideas,
X
To Joana Machado for helping me so much during my last months in the lab, and
for all longs days playing with the IRGA.
To Rita Borba, Paulo Gouveia, Sissi, Nuno, Rita Leal, Bruno, Vanessa, Cátia,
Grosa and Inês for making the GPlantS Lab a great place to learn, work and
growth.
To all Forest Biotech group for all patience needed to be our second home.
To Isabel Abreu for supporting me a lot into my trips into the protein-world, for
teaching me how to think as a scientist, for always motivate me to do the best and more, much more…
To Dedas for being a great friend, for all huge support at work, for being such a
great person.
To Gruska for being a great friend, for helping me a lot with my lab work, for being
so patience with my mess, for all the great talks, for teaching me polish. Dziękuję kobietę.
To Tânia Serra for all the tremendous support. I have no words for thank you for
teaching so much. Thank you for listen me, helping me solving challenges,
inspire me.
Aos meus pais e irmã, por estarem sempre comigo. Por me amarem. Obrigado
pela força, equilíbrio e conselhos que dão a cada dia. Por estarmos sempre
fortemente ligados uns aos outros.
A Tânia Lucas, por todo o carinho, amor e paciência durante esta fase. Por
ensinar-me sempre a ser otimista, por acreditar em nós, por levar-nos a
brincar como crianças, por sempre sorrir. Principalmente, por tornar as
XI
I, Diego Melo Almeida, declare by my honour to have had active participation inthe following papers and book chapter:
- Diego M. Almeida*, M. Cecília Almadanim*, Tiago Lourenço, Isabel A. Abreu,
Nelson J. M. Saibo, M. Margarida Oliveira. “Screening for abiotic stress
tolerance in rice: salt, cold and drought.” Paula Duque (ed). Environmental
Responses in Plants: Methods and protocols, Methods in Molecular Biology,
1398:155-82. DOI: 10.1007/978-1-4939-3356-3_14, Springer Science +
Business Media New York 2016. *First authors
Diego M. Almeida performed part of the review work and manuscript writing.
- Tânia S. Serra, Duarte D. Figueiredo, André M. Cordeiro, Diego M. Almeida,
Tiago Lourenço, Isabel A. Abreu, Alvaro Sebastián, Lisete Fernades, Bruno
Contreras-Moreira, M. Margarida Oliveira, Nelson J.M. Saibo (2013). “OsRMC,
a negative regulator of salt stress response in rice, is regulated by two AP2/ERF transcription factors” Plant Molecular Biology journal. 2013 Jul;82(4-5):439-55. DOI: 10.1007/s11103-013-0073-9.
Diego M. Almeida participated in the experimental work and manuscript writing.
- Diego M. Almeida, Glenn B. Gregorio, M. Margarida Oliveira, Nelson J. M.
Saibo. “Five novel transcriptions factors regulating OsNHX1 expression in a salt tolerant rice genotype” (under re-submission to Plant Molecular Biology journal)
Diego M. Almeida participated in the experimental design, the laboratory work
XII
- Diego M. Almeida, M. Margarida Oliveira, Nelson J. M. Saibo. “Na+ and K+ homeostasis in plants under salt stress” (Review submitted to Genetics and Molecular Biology journal)
Diego M. Almeida performed the review work and manuscript writing. This
manuscript includes part of the work described in Chapter I.
- Diego M. Almeida, Joana Machado, Pedro M. Barros, Helena Sapeta, M.
Margarida Oliveira, Nelson J. M. Saibo. “OsPCF2, a class I TCP transcription factor involved in stomatal aperture in rice” (In preparation)
Diego M. Almeida performed the review work and manuscript writing. This
XIII
LIST OF ABBREVIATIONS
3-AT 3-amino-1,2,4-triazole
a.a. Amino acid
ABA Abscisic acid
ABRE ABA-responsiveelement
At Arabidopsis Thaliana
ATP Adenosine triphosphate
bp Base pair
BLAST Basis Local Alignment Search Tool
Bv Beta vulgaris
CaM Calmodulin
CAMV35S Cauliflower Mosaic Virus 35S promote
cDNA Complementary DNA
CM-His Complete Minimal medium lacking Histidine
CM+His Complete Minimal medium supplemented
with Histidine
CPP Cystein-rich Poly comb-like Protein
CPY CarboxypeptidaseY
Cys Cysteine
ºC Degree Celsius
DNA Deoxyribonucleic Acid
dNTPs Deoxynucleotide Triphosphates
dS DeciSiemens
DTT Dithiothreitol
EC Electrical conductivity
EDTA Ethylene Diamine Tetraacetic Acid
EMSA Electrophoretic Mobility Shift Assays
XIV
g Gram
g Relative centrifugal force
GFP Green fluorescent protein
gs Stomatal Conductance
GUS β-glucuronidase
h Hour
H2O2 Hydrogen peroxide
Hv Hordeum vulgare L.
In Ipomea nil
IRRI International Rice Research Institute
KDa KiloDalton
Kg Kilogram
Km Michaelis–Menten constant
L Litre
LUC Luciferase
M Molarity
m Mass
Mb Mega base pair
µCi MicroCurie
µg Microgram
µL Microliter
µM Micromolar
min Minute
mL Mililitre
mM Milimolar
mRNA Message Ribonucleic Acid
MYC Myelocytomatosis Oncogene
MYB Myeloblastosis Oncogene
XV
NIN Nodule Inception
ng Nanogram
NLS Nuclear Localization Signal
NSCC Non Selective Cation Channels
Os Oryza sativa
PCNA Proliferating Cell Nuclear Antigen
PCR Polymerase Chain Reaction
PEG Polyethylene Glycol
PPM Parts per Million
PM Plasma Membrane
PVC Prevacuolar compartment
QTL Quantitative Trait Locus
RIL Recombinant Inbred Line
RNA Ribonucleic Acid
ROS Reactive Oxygen Species
rpm Rotations per minute
RT Room temperature
RT-PCR Reverse Transcription – PCR
RT-qPCR Quantitative Real-Time RT-PCR
RWC Relative Water Content
s Second
Sc Saccharomyces cerevisiae
SD Standard Deviation
SDS-PAGE Sodium Dodecyl Sulfate –
Polyacrylamide Gel Electrophoresis
SE Standard Erro
SES Standard Evaluation Score
SOS Salt Overly Sensitive
XVI
T-DNA Transfer-DNA
TF Transcription Factor
TGN Trans-Golgi Network
TRX Thioredoxin
Vv Vitis vinifera
XVII
SUMMARY
For half of the world´s population, rice is life. This cereal crop is
considered an important staple food worldwide, and more than three billion people
count on it for 50-80% of their daily calorie intake. Soil salinity is a major
environmental constraint to crop production, resulting in considerable yield losses
around the globe every year. According to the Food and Agriculture Organization
(FAO), in 2008 over 6% of world's total landand over 20% of irrigated land were
affected by high levels of salt. Irrigated land is only 15%of cultivated land, but it produces one third of the world’s food, raising awareness about salinity as a serious problem for crop productivity. Rice like as most crops is very sensitive to
salt, showing salt stress symptoms and reduced yield at relatively low soil salinity
levels (≈ 40 mM NaCl). Among the agronomically important cereals, rice shows
the highest sensitivity to salt. However, some degree of genotype tolerance for
salt stress is available in rice germplasm. To cope with salt stress conditions,
plants evolved several and diverse response mechanisms. One of these
mechanisms is tissue tolerance, in which high salt concentration is found in leaves
but is compartmentalized, especially in the vacuole, reducing the deleterious
effect of Na+ in the cytosol and driving water uptake to cells. Cation/H+ antiporters
mediate the transport of Na+ into the vacuole. This Na+/H+ exchange is mediated
by members of a family of transporters referred to as K+,Na+/H+ antiporters
(NHX-type). Among them, NHX1 is the most abundant and the best characterized
member. Several studies have shown that NHX1 overexpression leads to
improved salt and drought stress tolerance in various plant species. Given that
transcription factors (TFs) can act as master regulators of different cellular
processes, they are promising candidates for modifying complex traits in crop
plants, such as salt stress tolerance. Nevertheless, NHX1 transcriptional
regulation under salt stress is poorly understood.
The main objective of our study was the identification and functional
characterization of TFs regulating OsNHX1 expression under salt stress in a salt
XVIII
tolerant rice genotype in which the regulation of the OsNHX1 gene expression in
response to salt stress could be relevant for the salt stress tolerance at seedling
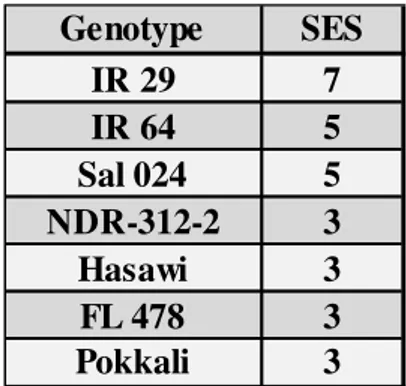
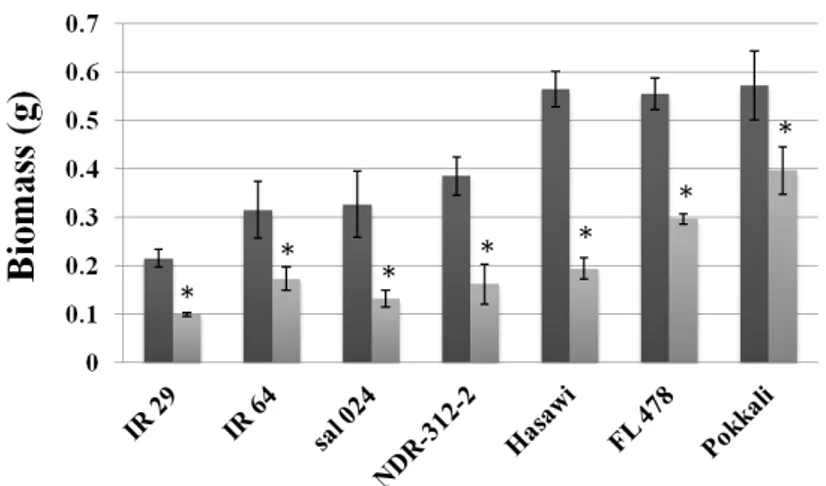
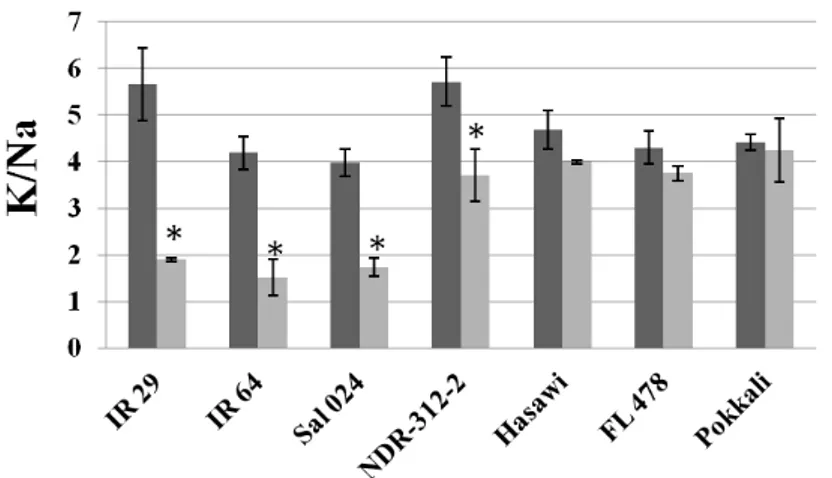
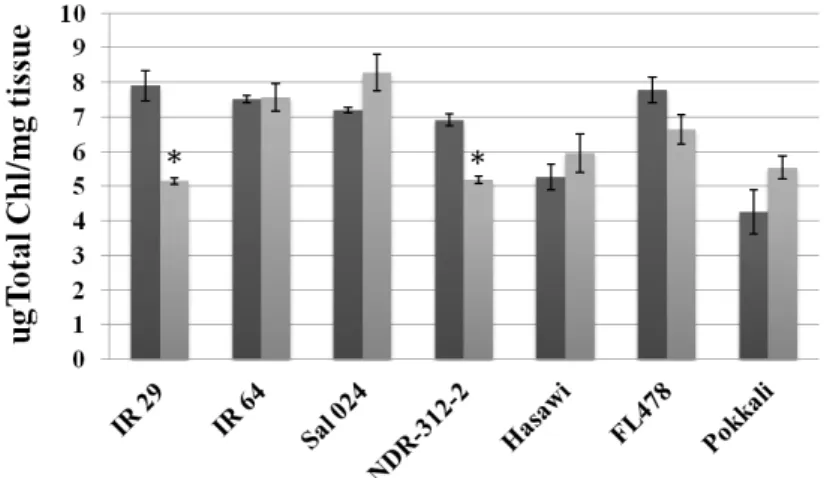
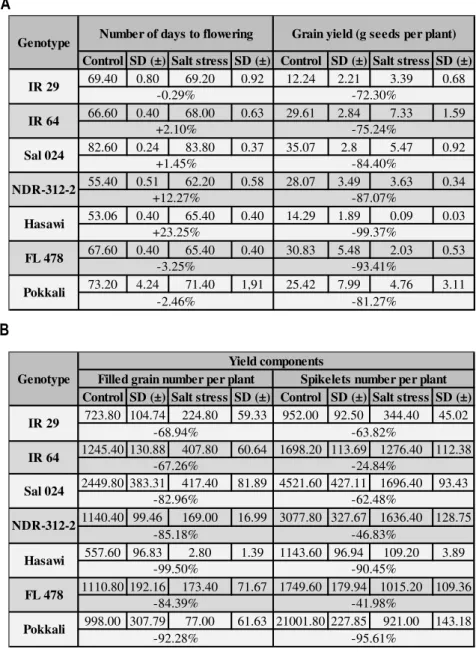
stage. Among the seven rice genotypes analyzed, we have selected Hasawi,
which showed a strong salt stress tolerance and high OsNHX1 responsiveness to
salt stress. Using the Yeast-One-Hybrid (Y1H) system to screen a salt-induced
rice cDNA expression library from Hasawi, five TFs belonging to three distinct
families were identified as binding to OsNHX1 promoter: one TCP (OsPCF2), one
CPP (OsCPP5) and three NIN-likes (OsNIN-like 2, OsNIN-like 3 and OsNIN-like
4). Transactivation activity assays performed in Arabidopsis and rice protoplasts
showed that OsPCF2 and OsNIN-like 4 are activators of the OsNHX1 gene
expression, while OsCPP5 and OsNIN-like 2 act as repressors. The
transactivation activity of OsNIN-like 3 needs to be further investigated.
When we analyzed the transcript levels of these TFs in rice seedlings
subjected to abiotic stress conditions, it was observed that all of them are early
regulated by both salt stress and PEG-simulated drought, especially in roots. The
expression of OsPCF2 in roots under salt stress and the OsNIN-like 4 in roots
subjected to PEG were mainly up-regulated in Hasawi, indicating that these TFs
may be associated with the salt and drought stress tolerance observed in Hasawi.
Analyses of the rice NHX-type gene promoters showed that OsPCF1 and
OsPCF2 (both TFs are TCP class I) binding motifs were over-represented in the
promoter of all OsNHX genes. Using an Electrophoretic Mobility Shift Assay
(EMSA), we showed that both OsPCF1 and OsPCF2 proteins bind to all OsNHX
gene promoters. In addition, a genome-wide search identified TCP class I binding
motifs in the promoter region of 3.089 rice genes. Among these genes, ten
(OsAKT2, OsKAT2, OsKAT3, OsKC1.2, OsALMT1, OsVHA-a1, OsVHA-a2,
OsVHA-a3, OsVHA-F, and OsPIP1;1) are somehow related to stomatal aperture.
We showed that OsPCF2 binds to the respective cis-regulatory elements present
in the promoters of all these genes. In addition, a rice T-DNA insertion line for
OsPCF2 (gene knockout) revealed a general down-regulation for most of the
XIX
conductance to water vapour under light conditions as well as reduced salt stresstolerance. Moreover, we observed that OsPCF2 seems to be posttranslationally
regulated by H2O2, thus modulating its binding to the OsNHX1 promoter.
This work allowed the identification of five novel TFs binding to the
promoter of OsNHX1, which is known to have a role controlling plant cell turgor
and expansion, thus mediating abiotic stress effects on plant development.
Further characterization of these TFs will help us to better understand their
function and it will unveil novel targets for improvement of plant abiotic stress
XXI
SUMÁRIO
Para metade da população mundial, o arroz é vida. Este cereal é
considerado um alimento essencial em todo o mundo. Mais de três mil milhões de
pessoas dependem diretamente do arroz para satisfazer cerca de 50-80% da sua
ingestão diária de calorias. A salinização dos solos é um dos maiores obstáculos
ambientais que limita a produção agrícola, resultando em perdas significativas na
produtividade a nível mundial. De acordo com a Organização das Nações Unidas
para Alimentação e Agricultura (FAO), em 2008 mais de 6% da área cultivada
mundial total e mais de 20% das superfícies irrigadas estavam afetadas pela
salinidade. As superfícies irrigadas representam apenas 15% das terras
cultivadas, mas produzem um terço dos alimentos a nível mundial. Estes
números despertam uma maior sensibilização para o grave problema da
salinidade dos solos na produtividade de diversas culturas. O arroz, assim como
a maioria das culturas cerealíferas, é sensível à salinidade e apresenta sintomas
de stress salino e redução na produção em solos com relativamente baixa salinidade (≈ 40 mM NaCl). Entre os cereais de maior interesse agronómico, o arroz é o mais sensível à salinidade. No entanto, alguns genótipos de arroz
apresentam um maior grau de tolerância à salinidade. Para lidar com a elevada
salinidade, as plantas desenvolveram muitos e variados mecanismos, sendo a
tolerância ao nível dos tecidos um deles. Neste caso, as folhas apresentam uma
elevada concentração de sal, mas este está compartimentado no vacúolo,
reduzindo os efeitos nefastos do Na+ no citosol e promovendo a absorção de
água para as células. Os anti-portadores catião/H+ medeiam o transporte de Na+
para o vacúolo. O transporte Na+/H+ entre o citosole o vacúolo é mediado por
membros da família de transportadores referidos como anti-portadores K+,Na+/H+
(tipo NHX). Entre estes transportadores, o NHX1 é o membro mais abundante e
melhor caracterizado. Além disso, vários estudos demostraram que a
sobre-expressão do NHX1 conduz a um aumento da tolerância aos stresses salino e
hídrico, em várias espécies de plantas. O fato dos fatores de transcrição (FT)
XXII
bons candidatos para regular características complexas em plantas, como por
exemplo a tolerância ao stress salino. No entanto, a regulação transcricional do
NHX1 em resposta ao stress salino está longe de ser bem conhecida.
O objetivo principal deste estudo foi a identificação e caracterização
funcional de FT que se ligam ao promotor do OsNHX1 proveniente de um
genótipo de arroz tolerante à elevada salinidade. Inicialmente, este projecto visou
a identificação de um genótipo de arroz tolerante ao stress salino em que a
resposta do OsNHX1 ao stress salino pudesse ser relevante para a tolerância da
planta. Entre os sete genótipos de arroz analisados selecionámos o genótipo
Hasawi, pois apresentou uma acentuada tolerância ao stress salino assim como
uma elevada indução do gene OsNHX1 pelo mesmo stress. Utilizando o sistema
Yeast-One-Hybrid (Y1H) para fazer a triagem de uma biblioteca de expressão de
cDNA de arroz, genótipo Hasawi, induzido pelo stress salino, identificámos cinco
FT, pertencentes a três famílias distintas, que se ligam ao promotor do OsNHX1
de Hasawi: um TCP (OsPCF2), um CPP (OsCPP5) e três NIN-Like (OsNIN-like 2,
OsNIN-like 3 e OsNIN-like 4). Observámos que alguns destes FT funcionam
como repressores (OsCPP5, OsNIN-like 2) e outros como ativadores da
transcrição (OsPCF2, OsNIN-like 4). A atividade transcricional do OsNIN-like 3
necessita ser mais investigada.
Quando analisámos o nível de transcrição dos FTs em plântulas de arroz
submetidas a condições de stress abiótico, observou-se que a expressão génica
de todos os FT é rapidamente modulada pelo stress salino e hídrico (induzido por
tratamento com PEG), especialmente nas raízes. A expressão de OsPCF2 pelo
stress salino, nas raízes, e do OsNIN-like 4 pelo PEG, nas raízes, foram
principalmente sobre induzidos em Hasawi (tolerante ao stress salino e hídrico),
indicando que estes FTs podem estar associada na tolerância ao sal e seca
observados em Hasawi.
A análise da região promotora dos genes NHX de arroz revelou que os
motivos de ligação para o OsPCF1 e OsPCF2 (FTs TCP classe I), encontram-se
XXIII
Electrophoretic Mobility Shift Assay (EMSA) mostrámos que as proteínas
OsPCF1 e OsPCF2 interagem com todos os motivos identificados. Além disso,
uma análise in silico a todo o genoma do arroz permitiu identificar locais de
ligação para FT da família TCP classe I na região promotora de 3.089 genes.
Entre estes genes, dez (OsAKT2, OsKAT2, OsKAT3, OsKC1.2, OsALMT1,
OsVHA-a1, a2-OsVHA, OsVHA-A3, OsVHA-F e OsPIP1; 1) estão, de alguma
maneira, relacionados com a abertura dos estomas. A ligação da proteína
OsPCF2 aos promotores destes genes foi demostrada por EMSA. A análise de
uma linha mutante de arroz com inserção de T-DNA (knockout para o gene
OsPCF2) revelou uma redução generalizada da expressão génica dos alvos do
OsPCF2, redução do teor de K+ nas partes aéreas e raízes, redução na
condutância estomática em condições de luz e redução da tolerância ao stresse
salino. Para além disso, verificámos que a ligação do OsPCF2 ao promotor do
OsNHX1 parece ser regulada por modificações pós-traducionais induzidas pelo
H2O2.
Este trabalho permitiu a identificação de cinco FT que se ligam ao
promotor do OsNHX1, o qual tem sido descrito como tendo uma função na
regulação da turgescência e expansão celular das plantas, mediando assim o
desenvolvimento das plantas em resposta aos stresses abióticos. Estudos
adicionais de caracterização funcional destes FT, irão revelar possivelmente
novos alvos para o aumento da tolerância das plantas aos stresses abióticos e
XXV
TABLE OF CONTENTS
Acknowledgments ... IX
List of Abbreviations ... XIII
Summary ... XVII
Sumário ... XXI
Chapter I ... 1
General introduction and Research Objective
Chapter II ... 57
Physiological and Molecular Responses of Seven Rice Genotypes to Salt Stress
Chapter III ... 85
Identification and Characterization of Five Novel Transcriptions Factors
Regulating OsNHX1 Expression in a Salt Tolerant Rice Genotype
Chapter IV ... 131
OsPCF2, a class I TCP Transcription Factor Involved in Stomatal Aperture in Rice
Chapter V ... 183
1
Chapter I
2
TABLE OF CONTENTS – CHAPTER I
Importance of rice ... 3
Salt stress effects on plant growth and yield ... 4
Sodium uptake from soil, sensing and signaling mechanism ... 7
Mechanisms of salt tolerance ... 9
Sodium transporters and plants salt stress tolerance ... 9
H+-Pumps and the plant salt stress response ... 11
Plasma membrane H+-ATPase ... 12
Vacuolar H+-ATPase... 13
Plasma membrane and vacuolar H+-PPase ... 16
SOS1 and the plant salt stress response ... 17
HKTs and the plant salt stress response... 21
HKT1 family ... 22
HKT2 family ... 25
NHX1 and the plant salt stress response ... 27
Diversity of plant NHX antiporters ... 28
Plant tissue localization ... 28
NHX gene expression under stress conditions ... 29
NHX structural organization and regulatory properties ... 30
Topology ... 30
Post-translational modifications ... 31
Function of NHX antiporters ... 32
Salt tolerance ... 32
K+ homeostasis ... 33
pH homeostasis ... 34
Vesicular trafficking ... 35
Transcriptional regulation ... 37
Research Objectives ... 38
3
IMPORTANCE OF RICE
For half of the world´s population, rice is life. Globally, this cereal crop is
considered a staple food for more than three billion people, accounting for 50-80%
of their daily calorie intake. Asia produced and consumed more than 90% of the world’s rice, and in many Asian countries rice represents a significant portion of its economy, labor force and even cultural heritage (Papademetriou, 2000). Rice is
cultivated in more than a hundred countries, as far north as (53º N) on the border
between Russia and China, and as far south as central Argentina (40º S) (IRRI,
2013; Papademetriou, 2000); clearly showing rice adaptability to diverse growing
conditions. There are two cultivated rice species: Oryza sativa, grown worldwide,
and Oryza glaberrima, grown in West and Central Africa. Based on the
morphology, physiology, agronomy, genetics and biochemistry features, Oryza
sativa can be divided in two main subspecies adapted to various environmental
conditions, indica and japonica. Indica includes the genotypes grown in tropical
and subtropical regions, whereas japonica genotypes grown in the cooler zones of
the subtropics and in the temperate environments (IRRI, 2013; Papademetriou,
2000).
In Europe, rice has an important sociocultural and ecological role in
several Mediterranean countries. Milled rice consumption in Europe ranges from
3.5-5.5 kg/person/year in northern countries to 6–14.8 kg/person/year in southern
countries. About 80% of the European rice production takes place in Italy and
Spain, and 12% in Portugal and Greece. The period of introduction of rice in
Europe is not concerted among academics, varying from 8th century up to late
10th century. Certainly, the Moors introduced rice to the south of the Iberian
Peninsula. Portugal is by far the largest rice consumer in Europe, consuming 14.8
Kg/person/year. Portuguese rice production is concentrated in three regions:
Mondego, Tejo and Sado rivers beds (IRRI, 2013).
The current world population of 7.3 billion is expected to reach 8.8 billion
in 2035. Most people will be living in Asia and Africa (UN, 2015). Combined, these
4
than 96% of the world’s rice consumption (IRRI, 2013; USDA, 2012). For the majority of the developing countries, rice availability means food security which is
closely connected to political stability (Bradsher, 2008; FAO, 2011). The task of
producing additional rice to meet the expected demands of people poses a major
challenge; for every one billion people added to the world’s population, it is
estimated that more 100 million tons of rice (paddy) have to be produced annually
(IRRI, 2013). This means an effort for overall increase in rice production of 26% in
the next 20 years, which must be achieved in a more efficient and
environmental-friendly system, using fewer resources (land, water, labor, etc.). To meet this goal,
high yield genotypes better adapted to adverse environmental conditions are
needed, while limiting yield losses. This is not possible without a comprehensive
understanding of the mechanisms controlling plant growth, development and
environmental stress adaptation (Bradsher, 2008; FAO, 2011; IRRI, 2013;
Papademetriou, 2000).
Many plant biology studies use Arabidopsis thaliana as model system.
However, Arabidopsis is not the best model for monocots, and our main staple
food crops, such as wheat, rice, and maize, are all monocots. In addition,
dicotyledons (dicots) and monocots are significantly distinct in many aspects of
their development (Izawa and Shimamoto, 1996). In spite of being a crop species,
rice has also emerged as a model organism for plant molecular biology studies,
and the main reasons for this are: it is relatively small, compared to other
monocots, and it has a fully sequenced genome (390 Mb); tools for functional
genomic analysis, T-DNA insertional mutant libraries are available, and the
production of transgenic plants is relatively easy, as compared to other cereals,
due to highly efficient transformation protocols (Nishimura et al., 2006; Shimamoto
and Kyozuka, 2002).
SALT STRESS EFFECTS ON PLANT GROWTH AND YIELD
Soil salinity is a major environmental constrain to crop production,
5
(Munns, 2005; Munns and Tester, 2008; Shabala and Cuin, 2008). Salt stress affects over 6% of the world’s total land area, most of this salt affected land has arisen from natural causes, including rainfall, windblown salt from ocean,tsunamis, and rock weathering. Apart from natural causes, soil salinization is
commonly associated to irrigation practices, such as the use of water with high
salt concentration, or land cleaning by removal of deep rooted vegetation or
replaced with shallow-rooted plants that use less water, leaving more water to
pass through soil to groundwater, raising the water table and bringing salt to the
surface where it can be left behind as the water evaporates (Abrol et al., 1988).
These Man-made actions led to a significantly increase in salt affected agriculture
cultivated land. Currently it is estimated that 20% of the total irrigated land is
salt-affected. Given that irrigated land produces at least twice more than rain-fed land and is responsible for one third of the world’s food production, it raises awareness for salinity as a serious problem for crop productivity (Munns, 2005; Munns and
Tester, 2008).
High soil salinity is a condition characterized by a high concentration of
soluble salts, in which NaCl is the most soluble and widespread salt. Soils are
classified as saline when the electrical conductivity (EC) is 4 dS/m (≈ 40 mM
NaCl) or more, which significantly reduces growth and yield of most crops. Rice as
well as most crop plants are glycophytes and show salt stress symptoms and
reduced yield even when the EC is lower than 4.0 dS/m. Among cereal crops, rice
is the most salt sensitive one (Munns and Tester, 2008). The salinity threshold for
rice is 3.0 dS/m with a 12% reduction in yield, per dS/m, beyond this threshold
(Gao et al., 2007). However, some degree of genotype tolerance for salt stress
tolerance is available in rice germplasm. Among 180.000 rice genotypes screened
by the International Rice Research Institute (IRRI), 17% had acceptable tolerance
at an EC of 10 dS/m at seedling stage (Gregorio et al., 2002).
Salt stress affects plants in two distinct phases. The first phase is the
osmotic effect; independent of the accumulation of salt in the shoot. Salts
6
from roots thermodynamically unfavorable, which induces water deficit (Pardo,
2010; Roy et al., 2014). Water deficit is rapidly transmitted (within minutes) from
roots to shoots causing intracellular turgor reduction and decreased cell
expansion (Munns, 2005; Munns and Tester, 2008). This signal also promotes the
biosynthesis of abscisic acid (ABA), which will induce stomatal closure and
consequently reduction in transpirational water loss (Munns, 2005; Munns and
Tester, 2008; Roy et al., 2014). Lower stomatal conductance leads to a lower
carbon assimilation, biomass production and decreased yield. The second phase
of salinity is ionic specific; this is due to the accumulation to toxic concentrations
of sodium (Na+) and/or chloride (Cl-) ions, especially in the older leaves, inducing
tissue necrosis and early leaf senescence (Roy et al., 2014). For most plant
species Na+ appears to reach a toxic concentration earlier than Cl- (Tester and
Davenport, 2003), and for rice (Chi Lin and Huei Kao, 2001; Tsai et al., 2004) Na+
has been shown to be the primary toxic ion. Furthermore, osmotic and ionic stress
disturb aerobic metabolism and induce the accumulation of reactive oxygen species (ROS) beyond the plant’s capacity for cellular oxidant detoxification, which in turn negatively affects cellular structures and metabolism (Chaves and
Oliveira, 2004; Chaves et al., 2009).
A deleterious effect imposed by salt stress, during the second phase, is
ions imbalance (Munns and Tester, 2008). Potassium (K+) is an essential
macronutrient that plays important functions related to enzyme activation, osmotic
adjustment and turgor generation, regulation of membrane potential, and
cytoplasmatic pH homeostasis (Barragán et al., 2012; PPI, 1998). Due to
similarity in physicochemical properties between Na+ and K+ (i.e., ionic radius and
ion hydration energy), the former competes with K+ for major binding sites in key
metabolic processes in the cytoplasm, such as enzymatic reactions, protein
synthesis and ribosome functions (PPI, 1998; Marschner, 1995). Na+ inhibits
enzyme activity of many of these enzymes that require K+ for functioning
(Duggleby and Dennis, 1973). With over 50 different cytoplasmic enzymes being
7
impairment, both in root and leaf tissues (PPI, 1998; Marschner, 1995). It hasbeen suggested that for plant survival under salt stress, it is essential to maintain
a high K+ concentration while keeping a low concentration of Na+ in the cytosol,
resulting in a high cytosolic K+/Na+ ratio. The restriction of Na+ accumulation in
shoots under salt stress has been reported as correlating with the salt stress
tolerance of rice (Lutts et al., 1996) and maize (Zea mays L.) (Tester and
Davenport, 2003).
SODIUM UPTAKE FROM SOIL, SENSING AND SIGNALING MECHANISMS
The enormous negative membrane potential across the plasma
membrane of plants root cells (negative inside) favor the passive transport of Na+
into the cells, and especially so when the sodium concentration increases in the
soil solution. In contrast, Na+ efflux (i.e., removal from the cell) is not passive and
requires energy expenditure (Maathuis et al., 2014). The major pathway for
passive Na+ entry into root cells at high soil salinity is mediated by a family of Non
Selective Cation Channels (NSCCs family), but their molecular identity remains
unknown (Blumwald et al., 2000; Kronzucker and Britto, 2011) (Fig. 1). In addition
to the Na+ flow across cellular membranes to enter the roots (symplast flow), it has
been reported that, at least in some species, interruptions in the endodermis
(passage cells) allow the movement of water and solutes (i.e., Na+) through the
cell wall and intercellular spaces. This type of transport, to the xylem stream,
without crossing the plasma membrane is referred as “apoplast flow” (Kronzucker
and Britto, 2011; Yeo et al., 1987) (Fig. 1). Casparian strips and suberine layers in
the root endoderm and exodermal layers provide some barrier to apoplast flow
(Yeo et al., 1987). In many plant species, such as rice, the apoplast flow is
considered to be the major port of Na+ entry (≈ 50% of total Na+ uptake) (Yeo et
al., 1987), especially at high salinity levels, and is responsible for a significant
amount of Na+ transported to the shoot (Kronzucker and Britto, 2011; Yeo et al.,
1987). Na+ ions taken up by the roots are then transported to shoots via xylem
8
creating tensions in the root xylem, which provides the major force to move water
from roots to up to the shoots (Nobel, 2009) (Fig. 1).
Sodium has also a strong inhibitory effect on K+ uptake by cells, probably
by inhibiting K+ transporters, such as AKT1 (hyperpolarization-activated
inward‐rectifying K+ channel), a major player in K+ acquisition by plants (Fuchs et
al., 2005; Hirsch et al., 1998), and HAK5 (carrier-type HUP/HAK/KT transport)
(Nieves-Cordones et al., 2010), both present in the plasma membrane of root
cells. Additionally, membrane depolarization caused by large cytosolic Na+ influx
results in increased K+ efflux possible through depolarization-activated
outward‐rectifying K+ channels (e.g.,GORK) (Adams and Shin, 2014) and NSCCs
(Sun et al., 2009).
Very little is known about how Na+ is sensed in most cellular systems. In
theory, Na+ can be sensed either before or after entry the cell, or both.
Extracellular Na+ may be sensed by a membrane receptor, whereas intracellular
Na+ may be sensed either by membrane proteins or by any of the many Na+
sensitive enzymes in the cytoplasm (Conde et al., 2011). The plasma membrane
Na+/H+ antiporter SOS1 (Salt Overly Sensitive 1) has been described as a
possible Na+ sensor (Shi et al., 2000). Its transport activity is essential for Na+
efflux from cells (Quintero et al., 2002), but its unusually long cytoplasmatic tail is
thought to be involved in Na+ sensing (Shi et al., 2000) (Fig. 3). However, this
mechanism it is not fully clear.
In plant cells, Ca2+ acts as a second messenger connecting a wide range
of extracellular stimuli with various intracellular responses (Conde et al., 2011).
Salt stress originates a fast and transient increase in free cytosolic Ca2+, likely
released from the vacuole (Pottosin et al., 2009), that is decoded by Ca2+ sensors
such as calmodulin (CaM), calcineurin B-like proteins (CBLs) and CBL-interacting
protein kinases (CIPKs). When acting as a CBL-CIPK complex, these Ca+ sensors
are often designed as calcium-dependent protein kinases (CDPKs) (Conde et al.,
2011; Yang and Poovaiah, 2003). Cytosolic Ca2+ sensors in turn trigger many
9
(e.g,. NSCCs are strongly blocked by external Ca+2), as well as enzymatic activityand gene transcription, ending up in ion homeostasis (Adams and Shin, 2014;
Conde et al., 2011; Martinez-Atienza et al., 2007; Pardo and Quintero, 2002;
Yamaguchi et al., 2005).
MECHANISMS OF SALT TOLERANCE
Salt stress frequently affects plant habitats and many species evolved
varied mechanisms for dealing with it. These mechanisms for salt tolerance can
be classified into three main categories. The first one is osmotic stress tolerance,
which is regulated by is regulated by long distance signals that reduce shoot
growth (Roy et al., 2014) and involves biosynthesis and accumulation of
compatible solutes to maintain water uptake (Peleg et al., 2011). Another
mechanism is ion exclusion, in which Na+ transport reduces the accumulation of
toxic Na+ within leaves. This system operates by controlling the Na+ loading to the
xylem and Na+ retrieval from the xylem, before reaching the shoot photosynthetic
tissues (Fig. 1). Finally, the third mechanism is tissue tolerance, in which high salt
concentration is found in leaves, but Na+ is compartmentalized at the cellular and
intracellular level(especially in the vacuole) reducing the deleterious effect of Na+
in the cytosol and driving water uptake to cells (Fig. 1) (Munns and Tester, 2008).
In most cases, the plant salt stress tolerance relies on the three mechanisms
together, rather than only one mechanism is particular (Munns and Tester, 2008;
Pires et al., 2015; Roy et al., 2014).
SODIUM TRANSPORTERS AND PLANTS SALT STRESS TOLERANCE
The study of salt stress tolerance in plants usually focuses on the control
of Na+ movement, namely on: Na+ exclusion in roots, Na+ long distance transport,
and Na+ compartmentalization at both cellular and tissue level (Conde et al., 2011;
Munns, 2005; Roy et al., 2014). These processes are mediated by membrane
transporters reason why the manipulation of their activity has an enormous
10
Here, we focus on the specific membrane transporters described as involved in
the above outlined tolerance processes. In contrast to animal cells, higher plants
do not have Na+-ATPases or Na+/K+-ATPases and rely on H+-ATPases and H+
-pyrophosphatases (PPases) to create a proton-motive force necessary to drive
Na+ transport across membranes (Conde et al., 2011). The plasma membrane
localized SOS1 (Ji et al., 2013; Martinez-Atienza et al., 2007) and the vacuole
membrane (tonoplast) localized NHX1 (Fukuda et al., 2011; Jiang et al., 2010) are
two Cation/H+ antiporters involved in Na+ exclusion back to the soil and in K+-Na+
compartmentalization in the vacuole. In addition, members of the HKT1 family of
HKTs (high affinity potassium transporters) are involved in the control of Na+ long
distance transport by reabsorption of Na+ from the xylem sap into the root cells,
preventing the large accumulation of Na+ in the above-ground tissues (Rus et al.,
2004) (Fig. 1). It is noteworthy that HKT1 Na+ exclusion mechanism from the
transpiration stream has been frequently indicated as a strong trait in salt
tolerance of different cereals, such as rice (Ren et al., 2005) and durum wheat
(Triticumturgidum L. subsp. durum) (James et al., 2006).
In the following sections, the role that different Na+ transporters and H+
11
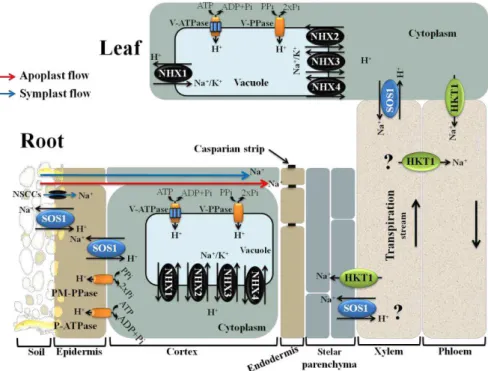
Figure 1. Summary diagram showing key plasma and tonoplast membrane transporters, channels and pumps mediating Na+ and K+ homeostasis in plants under salt stress (adaptedfrom Roy et al. 2014). Na+ ions enter the cells via Non Selective Cation Channels (NSCCs),
likely other cation transporters (not shown) and through the cell wall and intercellular spaces (apoplast flow – red arrow). The Na+/H+ antiporter SOS1 extrudes Na+ at the root soil interface,
thus reducing the Na+ net influx of Na+. At the xylem parenchyma cells, HKT1-like proteins
retrieve Na+ from the xylem sap thereby restricting the amount of Na+ reaching the
photosynthetic tissues. To translocate Na+ back to the root, ions unloaded from xylem may be
transported into phloem via additional HKT1-like protein. In addition, HKT1-like proteins also load Na+ into shoot phloem and then Na+ is transferred into roots via downstream of phloem,
preventing Na+ accumulation in shoots. SOS1, xylem parenchyma cells localized, is also
suggested to mediate Na+ efflux from xylem vessels under high salinity. Incoming Na+, in root
and shoots, is stored in the large central vacuole by tonoplast localized NHX exchangers (NHX1-4). Plasma membrane (PM) H+-ATPase (P-ATPase), PM H+-PPase (PM-PPase),
tonoplast H+-ATPase (V-ATPase) and tonoplast H+-PPase (V-PPase) generate electrochemical
potential gradient for secondary active transport.
H+-Pumps and the plant salt stress response
Proton gradients are crucial for the transport of ions and solutes across
the different plant cell membranes. Three primary proton transport proteins are
found in plant cells: (1) plasma membrane (PM) and (2) vacuolar H+-ATPases,
which couple ATP hydrolysis with proton transport, and (3) PM and vacuolarH+
12
al., 2010; Gaxiola et al., 2007). In plant cells, H+-ATPase and H+-PPase are major
components of the vacuole membrane (Silva and Gerós, 2009). The H+-Pumps
generate an electrochemical potential gradient across membranes, which is the
motive force for a large set of secondary transports.
1- Plasma membrane H+-ATPase
The PM H+-ATPase belongs to a class known as P-type ATPases
(P-ATPases), and is encoded by a large gene family (Fuglsang et al., 2010; Gaxiola
et al., 2007). The pump is formed by a single subunit protein, which contains ten
trans-membrane helices and a large cytoplasmatic domain (Fuglsang et al.,
2010). Arabidopsis and rice genomes encode eleven and ten P-ATPases,
respectively (Arango et al., 2003; Axelsen and Palmgren, 2001).
The proton motive force created by P-ATPases is largely responsible for
an inside negative potential across the plasma membrane, which is essential for
root nutrient uptake, stomatal aperture, phloem loading, and cell growth
(Blumwald et al., 2000; Gaxiola et al., 2007; Mansour, 2014). Besides regulation
of many physiological processes, the P-ATPases have a critical role in plant
adaptation to salt stress conditions. Higher P-ATPases activity under salt stress
conditions repolarizes the NaCl-induced depolarization of PM. This response has
been strongly associated with salt stress tolerance (Mansour, 2014). The
maintenance of the PM potential under salt stress through P-ATPases activity has
a great effect on reduction of Na+ influx via depolarization-activated NSCCs and
K+ efflux via KORs and NSCCs, which help to restore higher K+/Na+ levels (Sun et
al., 2009). Also, P-ATPases higher activity under stress energizes the active
transport that exclude Na+ from root cells, a process dependent on the SOS1
Na+/H+ antiporter (Gaxiola et al., 2007). Furthermore, it was reported that higher
activation of P-ATPases is often found in halophytes and salt tolerant genotypes,
which may correlate with salt stress tolerance (Mansour, 2014). For instance, in
rice callus lines, a higher activation of P-ATPases occurred in salt-tolerant lines as
13
The salt-dependent activation of PM H+-pump is associated withincreased levels of gene expression as well as post-translational modifications of
the enzyme present in a preexisting pool (Gaxiola et al., 2007; Mansour, 2014).
However, it is likely that most regulation of the pump activity occurs at the
post-translational level (Fuglsang et al., 2010; Gaxiola et al., 2007). The pump activity
can be modulated by phosphorylation/dephosphorylation of the penultimate a.a.
residue of the cytoplasmatic C-terminus domain, a threonine residue. The
phosphorylated threonine residue promotes binding of the activating 14-3-3
protein (Fuglsang et al., 2010).
Stomatal aperture involves regulation of osmotic pressure within the
guard cells, a process powered by P-ATPases activity and responsive to a wide
variety of external signals (Gaxiola et al., 2007). Blue light perception in guard
cells is mediated by phototropins, which intitiate a signal transduction signal
pathway that involves an upstream protein phosphatase I and a downstream
protein kinase that phosphorylates the penultimate C-terminus a.a. residue of the
P-ATPase (Gaxiola et al., 2007; Takemiya et al., 2006). Under drought and salt
stress conditions, stomatal closure is induced by ABA through a mechanism that
involves production of hydrogen peroxide (H2O2) and dephosphorylation of the
P-ATPases (Gaxiola et al., 2007; McAinsh et al., 1996; Zhang et al., 2001).
2- Vacuolar H+-ATPase
Among the three proton-pumps found in plant cells, the vacuolar H+
-ATPase (-ATPase) is the most complex one (Gaxiola et al., 2007). The
V-ATPase was first found associated with the endomembrane system where it
acidifies and generates a proton force motive within diverse cell compartments
(e.g.,vacuole, endoplasmic reticulum and trans-Golgi network) (Ratajczak, 2000).
However, V-ATPases have also been associated with cell plasma membrane
(Hanitzsch et al., 2007). The ability of the V-ATPase to maintain the cytosolic pH
homeostasis and to acidify the endomembrane compartments is crucial during
14
Vacuolar H+-ATPases are multisubunit enzymes composed of two
subcomplexes: Three peripheral V1 complexes consisting each of eight subunits
(A, B, C, D, E, F, G and H) responsible for ATP hydrolysis, and the
membrane-integral V0 complex comprising up to six subunits (a, c, c´, c´´, d and e)
responsible for proton translocation (Gaxiola et al., 2007) (Fig. 2). In plants, the
sub-unit c´ is not found and many of the V-ATPase subunits are encoded by gene
families. In Arabidopsis andrice, the 13 subunits which compose the vacuolar H+
-ATPases (A, B, C, D, E, F, G, H, a, c, c´´, d and e) are encoded by a total of 27
genes and 22 genes respectively (known as VHA genes). If all possible isoform
combinations are used, we will have hundreds of different V-ATPase complexes
(Hanitzsch et al., 2007; Sze et al., 2002).
By convention, the subunits of V1 and V0 complexes are distinguished
with capital and case letters, respectively. The V1 complex consists of: (1) a
globular hexameric head with three alternating copies of subunits A and B forming
a ring, (2) a central rotational stalk composed of single copies of subunits D and F,
and (3) a outer stalk made of subunits C, E, G and H. Subunits A and B mediate
the hydrolyses of ATP at three reaction sites associated with subunit A. Both the
central rotational stalk and fixed outer stalk connect the V1 complex to the
membrane inserted V0 complex. The proton transporting V0 complex consists of
six or more c subunits, also forming a ring structure. In addition, each V0 complex
contains one copy of subunits a, d and e (Beyenbach and Wieczorek, 2006;
Hanitzsch et al., 2007) (Fig. 2). It has been reported that structural changes of the
V-ATPase complex or presence/absence of individual protein isoforms could be
correlated with differences in V-ATPase localization and activity between plant
15
The plant vacuole plays a very important role in the maintenance ofcellular metabolism due to its role in long term storage of toxic ions, long or short
term storage of mineral and/or organic acids and in pH and Ca2+ cytoplasmatic
homeostasis. Furthermore, the V-ATPase is the most abundant H+-pump in the
tonoplast, and it has been shown that its activity is modulated to cope with
environmental and metabolic changes (Ratajczak, 2000). For instance, under salt
stress, a general increase of V-ATPase activity has been reported in many plant
species (Matsumoto and Chung, 1988; Silva and Gerós, 2009). The V-ATPase
provides the driving force necessary for K+-Na+ vacuole compartmentalization, a
process linked on the NHX1 antiporter activity (Bassil and Blumwald, 2014; Jiang
et al., 2010).
The ability to respond salt stress via changes in the expression of the
V-ATPase subunits encoding genes might be a prerequisite and a characteristic of
salt stress tolerance in plants. It has been reported that the transcript levels of
some subunits are up-regulated in response to salt stress (Kirsch et al., 1996;
Narasimhan et al., 1991; Silva and Gerós, 2009). However, the expression of
V-ATPase genes does not always involve a fixed stoichiometry of mRNAs for the
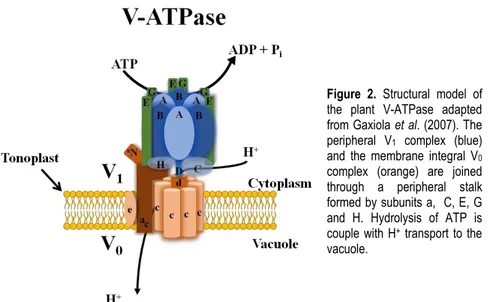
different subunits (Silva and Gerós, 2009). Other factors may also account for the Figure 2. Structural model of the plant V-ATPase adapted from Gaxiola et al. (2007). The peripheral V1 complex (blue)
and the membrane integral V0
complex (orange) are joined through a peripheral stalk formed by subunits a, C, E, G and H. Hydrolysis of ATP is couple with H+ transport to the
16
regulation of V-ATPase activity. Indeed, VHA-A subunit from barley (Hordeum
vulgare L.) was shown to interact to 14-3-3 proteins, well known activators of PM
ATPases, in a phosphorylation-dependent way. That interaction was suggested to
activate V-ATPase activity (Klychnikov et al., 2007).
3- Plasma membrane and vacuolar H+-PPase
H+-pyrophosphatases (H+-PPase) are highly hydrophobic single subunit
proteins that generate proton gradients across the vacuole, Golgi and plasma
membranes using the energy of hydrolysis of pyrophosphate (PPi) molecules
(Gaxiola et al., 2007). Plants have two phylogenetically distinct types of H+
-PPases: type I and type II. Type I H+-PPases depend on cytosolic K+ for their
activity and are moderately sensitive to inhibition by Ca2+, and type II H+-PPases
are K+ insensitive but extremely Ca2+ sensitive.
The Arabidopsis genome encodes two H+-PPases: one type I H+-PPase
(AVP1) and another type II H+-PPase (AVP2) (Drozdowicz et al., 2000). Rice
genome also encodes two H+-PPases: OVP1 and OVP2 (Sakakibara et al., 1996).
However, more isoforms have been proposed (Choura and Rebai, 2005).
Phylogenetic analysis of V-PPase sequences showed that rice H+-PPases are
likely to be type I H+-PPases (Drozdowicz et al., 2000). Type I H+-PPases are
mainly suggested to acidify the vacuole (Gaxiola et al., 2007). However these H+
-pumps were also found in the plasma membrane (Alexandersson et al., 2004;
Ratajczak et al., 1999). Arabidopsis type II H+-PPases, AVP2, has been shown to
localize exclusively to Golgi apparatus (Mitsudaa et al., 2001).
The expression levels of the H+-PPases are strictly regulated at
transcriptional level in response to various environmental conditions or
developmental stages (Fuglsang et al., 2010; Silva and Gerós, 2009). Under salt
stress an increased H+-PPase activity has been reported (Maeshima, 2000). It
has also been shown that the pollen-specific cis-acting region of the AVP1 gene is
involved in the expression of the AVP1 gene during pollen development.
AtCAM15, AtCAMTA 1 (calmoduline-binding transcription factors) (Mitsuda et al.,
17
(Mitsuda et al., 2004) were identified as binding to the cis-acting region of theAVP1 gene. However, a comprehensive mechanism of H+-PPase gene
expression and post-translational regulation is still needed. It is likely that the
protein C-terminus plays an essential role in supporting the physiological function
of H+-PPase activity (Fuglsang et al., 2010).
Given the importance of the pH homeostasis in the cytosol for cell
metabolism, it is likely that the activity of all three H+-pumps (P-ATPase,
V-ATPase and H+-PPase) is regulated by common regulatory mechanisms (e.g.,
14-3-3 proteins).
SOS1 and the plant salt stress response
Comparisons of unidirectional Na+ fluxes and rates of net accumulation of
Na+ in root indicate that 70-99% of the Na+ fluxed into the root is extruded back to
the apoplast (Munns, 2005; Tester and Davenport, 2003). For rice, that value is
indicated as 96% (Munns, 2005), meaning that over time Na+ will be accumulated
into roots and transferred to shoot via the transpiration stream, later accumulating
there. Since it is important to maintain low cytoplasmatic Na+ concentrations for
growth and survival under saline conditions, plants have developed a direct
mechanism to extrude Na+ from cells across the plasma membrane to the soil or
apopoplast. Small differences in Na+ exclusion capacity creates major changes in
Na+ net accumulation (Brini and Khaled., 2012; Munns, 2005; Tester and
Davenport, 2003). However, the role of cellular Na+ efflux is not intuitive in
multicellular plants, as Na+ transport out of one cell would negatively impact the
surrounding neighbor cells. So, the role of Na+ efflux has to be considered in
specific tissues and in the context of whole plants (Zhu, 2003). Sodium efflux is
catalyzed by the plasma membrane Na+/H+ antiporter encoded by SOS1 (Salt
Overly Sensitive1 = AtNHX7) gene, identified in several plants including
Arabidopsis (Wu et al., 1996), rice (Martinez-Atienza et al., 2007), wheat (Xu et
al., 2008), and tomato (Xu et al., 2008). SOS1 uses the proton gradient
18
for H+ across the membrane (Ji et al., 2013; Qiu et al., 2004; Shi et al., 2002).
Activity of the ArabidopsisSOS1 promoter is detected ubiquitously in virtually all
tissues, but it appears to be more active in: (1) root epidermal cells (particularly at
the root tip), suggesting that meristem requires special protection, since the root
tip cells have very small vacuoles and thus incapable of vacuolar Na+
compartmentalization, and (2) root parenchyma cells lining the vasculature
(Kronzucker and Britto, 2011; Shi et al., 2002). The SOS1 gene expression
pattern, together with the results of ion analysis in sos1 mutant plants, suggests
that SOS1 has several roles: (1) Na+ efflux from roots; (2) slow down Na+
accumulation in the cytoplasm in order to gain time for Na+ storage in the vacuole;
and (3) control of long-distance Na+ transport between roots and leaves by loading
and unloading Na+ into and from the xylem (Conde et al., 2011; Zhu, 2003). SOS1
may mediate active loading of Na+ to the xylem under mild salinity (25mM NaCl).
However, at salt stress (100 mM NaCl), expression of SOS1 is induced and SOS1
may function in Na+ retrieval from the xylem (Shi et al., 2002). Such SOS1 role in
long-distance transport is important for coordination between transpiration Na+
flow and Na+ vacuolar sequestration in leaves. However, thermodynamic analysis
by Munns and Tester (2008) indicates the Na+ removal from the xylem is unlikely
to be mediated by a Na+/H+ antiporter such as SOS1, because its operation “in reverse” under high Na+ conditions is thermodynamically unfavorable. Instead, class I HKTs have been shown to be involved in xylem unloading of Na+
(Davenport et al., 2007; James et al., 2006; Ren et al., 2005). Thus, the role of
SOS1 in long-distance Na+ transport remains unclear. Nevertheless, many reports
suggest that SOS1 plays a critical role in Na+ exclusion, thus maintaining cellular
ion homeostasis and allowing plants to survive and grow under salt stress
conditions (Cuin et al., 2011; Shi et al., 2003).
The transcript level of SOS1 is upregulated by salt stress (Shi et al.,
2000). Analysis of the 2 Kb upstream of the SOS1, CIPK24/SOS2 and
CBL4/SOS3 transcription initiation site revealed that the promoter of these genes
19
WRKY, and TCP classes (Ji et al., 2013). However, TF(s) mediating promoteractivity of SOS genes have not yet been identified. Upregulation of SOS1
transcripts under salt stress is suggested to be regulated at post-transcriptional
levels, as SOS1 promoter activity is not upregulated by salt stress but the SOS1
gene expression driven by the constitutive Cauliflower mosaic virus 35S promoter
is (Shi et al., 2003). This may indicate that the SOS1 transcript is unstable in the
absence of salt stress and the salt stress causes a post-transcriptional
stabilization of the transcript (Shi et al., 2003). More recently, it was suggested
that the Na+ stress induced SOS1 mRNA stability is mediated by reactive oxygen
species (ROS) (Chung et al., 2008). In addition, SOS1 upregulation by salt stress
is partly under the control of SOS2 and SOS3 (Shi et al., 2000). CIPK24/SOS2 is
a protein kinase and CBL4/SOS3 is a calcium sensor, that together with SOS1 are
the three key components comprising the Salt Overlay Sensitive (SOS) signaling
pathway identified in Arabidopsis (Wu et al., 1996) and rice (Martinez-Atienza et
al., 2007). At the cellular level, the SOS signaling pathway has been proposed to
mediate cellular signaling under salt stress to maintain the ion homeostasis (Ji et
al., 2013).
Activation of the Na+/H+ antiport activity of SOS1 by salt stress is
controlled by SOS3 and SOS2 (Ji et al., 2013; Zhu, 2003). In response to an
external stimulus, such as high Na+ concentration, transient increases in
cytoplasmatic Ca2+ occur and that is decoded by the calcineurin B and neuronal
Ca2+ sensor-like protein SOS3. Activation of SOS3 requires N-myristoylation and
Ca2+ bound on EF-hand Ca2+ binding sites. Activated SOS3 then physically
interacts with the auto-inhibitory domain of the SOS2, a member of the SnRK
(sucrose non-fermenting-related serine/threonine kinase) family, which activates
the kinase and facilitates the localization of the SOS2-SOS3 complex to the
plasma membrane. The SOS2-SOS3 complex then associates with the Na+/H+
antiporter SOS1, phosphorylating its C-terminal auto-inhibitory domain, which
becomes activated and pumps Na+ out of the cell (Brini and Khaled., 2012;
20
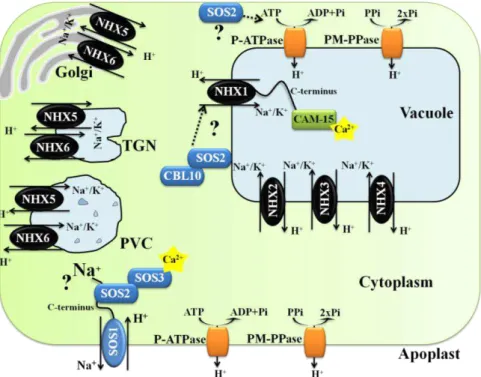
The SOS pathway is not limited to the three main proteins, as it interacts
with other stress related proteins. A SOS3 homolog SOS3-LIKE Calcium Binding
Protein8 (SCABP8/CBL10) interacts with SOS2 to form an alternative protein
kinase complex that regulates SOS1 activity in the plasma membrane in response
to salt stress, mainly in shoots; while SOS3 functions primarily in the root (Quan et
al., 2007) (Fig. 3). SOS2 phosphorylates CBL10 in a Ca2+ independent manner
upon salt stress, and this phosphorylation stabilizes the SOS2-CBL10 complex
association with the plasma membrane and increases SOS1 antiporter activity
(Hasegawa, 2013; Kim et al., 2007; Quan et al., 2007). Abscisic acid insensitive
(ABI2) interacts with SOS2 to prevent SOS3 binding to SOS2 and kinase
activation. Such ABI2-SOS2 interaction may represent an integrating node
between salt stress and ABA signaling (Hasegawa, 2013; Ohta et al., 2003).
The SOS pathway may also regulate the Na+ vacuolar
compartmentalization. Interaction of SOS2-CBL10 may result in localization of the
kinase complex at the vacuolar membrane where it is possibly involved in the
regulation of Na+/H+ exchange at the tonoplast, presumably by regulation of NHX
antiporter activity (Kim et al., 2007; Qiu et al., 2004). However, to date no NHX
antiporter has been shown to be directly regulated by SOS2 and/or by the
SOS2-complex. In addition, SOS2 has been suggested to regulate the V-ATPase
activity. SOS2 was found to interact with the B1 and B2 subunits of the V-ATPase
in the absence of CBL proteins, and tonoplast vesicles from the Arabidopsis
sos2-2 mutant showed reduced ATPase and H+-translocation activities (Batelli et al.,
2007).
Potassium homeostasis has also been shown to be modulated by the
SOS signaling pathway. The CIPK23 directly phosphorylates and activates the
AKT1 K+ channel at the plasma membrane, significantly increasing the K+ uptake
under low-K+ stress (Pardo, 2010; Ren et al., 2013). Furthermore, the protein
CBL10 has been indicated to directly interact with AKT1 channel, negatively
regulating its activity in roots in a CIPK-independent way (Ren et al., 2013). It is
21
high K+/Na+ cytosolic ratio under stress (Tester and Davenport, 2003). Thepossibility that CBL10 functions as an interconnecting regulator of SOS1 and
AKT1 may indicate that CBL10 plays a crucial role in ion homeostasis (K+/Na+)
under salt stress by regulating both Na+ and K+ uptake/exclusion (Ren et al.,
2013).
HKTs and the plant salt stress response
Another important determinant of salt stress tolerance in plants is the
activity of the HKT (high affinity potassium transporter) proteins (Munns and
Tester, 2008; Roy et al., 2014). The HKT family is quite diverse, and this diversity
reflects their large amplitude of functions (Almeida et al., 2013; Munns and Tester,
2008; Roy et al., 2014). The HKT family is divided in two distinct classes
according to their transport characteristics. The main distinguishing feature is the
a.a. sequence that constitutes the first pore domain (PD) (Platten et al., 2006).
Members of class I transporters (HKT1) have a serine (S), forming an S-G-G-G
motif, where most of the members of class II (HKT2) have a G in the position
occupied by the S in class I transporters, forming a G-G-G-G motif (Maser et al.,
2002). The presence of either S or G at this position is critical for K+ specificity of
the transporter. The presence of a S is characterized by a preference for Na+
conductance over other cations (HKT1), whereas the presence of a G is
characterized by transport of Na+ and/or K+ depending on the external
concentrations of these two ions (HKT2) (Kronzucker and Britto, 2011; Platten et
al., 2006). However, there are notable exceptions, in particular HKT2;1 from
cereals, in which the G has reverted to S (Kronzucker and Britto, 2011), but it has
been clearly shown to be involved in mediating Na+ and K+ entry into roots
(Kronzucker and Britto, 2011; Munns and Tester, 2008). The main role of HKT1 is
believed to be Na+ retrieval from the transpiration stream avoiding the over