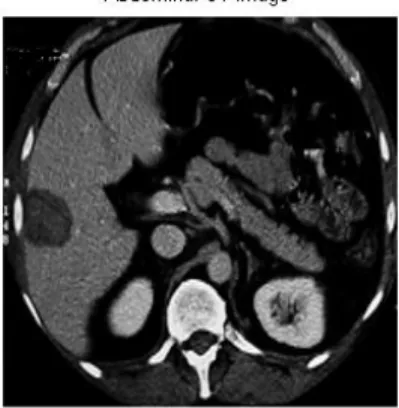
Diagnosis of Liver Tumor from CT Images using Curvelet Transform
Texto
Imagem
Documentos relacionados
Em outras palavras, o cenário atual de pesquisa apresenta uma criança que pensa e, mais que isso, que pensa bem. Desde cedo, então, a criança dispõe de recursos cognitivos
This technique employs Discrete Wave Transform (DWT) method for pre-processing, Approximate Entropy (ApEn) to extract features and Artificial Neural Network (ANN) for
Clustering results as well as the information obtained from clusters may vary with the use of different image features and parameters settings of clustering algorithms.. CT images
fibro scan measures liver fibrosis and CAP (Controlled Attenuation Parameter) which is equal to percentage of liver fat saturation .Materials and Methods: This
In this paper, a scooter fault diagnosis system based on continuous wavelet transform technique and faults classification using artificial neural network for the purpose of the
Methods: Various resected tumor samples (33 from lymph nodes, 42 from liver, six from lung, and 68 primary tumors) were assessed via a collagen gel droplet-embedded culture
For this, Wavelet Transform is applied as a method to extract relevant features of signal and KNN is used as a classification technique.. The results showed that
However, despite the increasing use of laparoscopy in colorectal and liver resections, combined purely laparoscopic resection of the primary colorectal tumor and synchronous LM