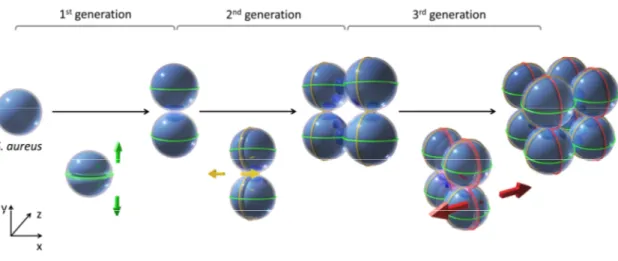
Cell division and chromosome segregation in
Texto
Imagem

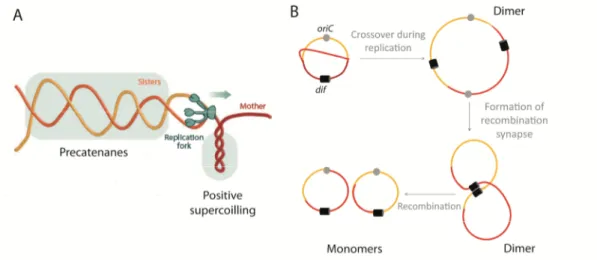
Documentos relacionados
We suggest that protein interactions of PBP2 that occur via the N-terminal domain must be important for retaining PBP2 and/or other cell wall synthetic proteins in
This protein contains a serine protease domain in the N-terminal region that requires the formation of a non-co- valent complex with the membrane-bound cofactor NS2B for activity,
In its native background, the NifA activity is regulated by the prevailing ammonium levels and the N-terminal GAF domain of NifA has been implicated in
The sequence analyses of these proteins show that the Pso2p catalytic core is localized within the N-terminal part of the protein, while a DNA ligase I domain can be detected in the
In this work, we report the expression of the N-terminal domain of TPL in Pichia pastoris to study the effect of the C-terminal domain deletion on the enzyme activity, and to verify
To date, no crystal structure of laforin has been determined and thus the specific properties and relative positions of the N-terminal CBM and the C-terminal DSP domain
Through quantitative real-time PCR (QRT-PCR) and immunoblotting with Abs against both N-terminal and C-terminal regions of ZNF750, we showed that ZNF750 mRNA and protein
The present work has identified a new microtubule binding region in MAP7D3, formed by the conserved MAP7 domain and the C-terminal tail, which can bind to microtubules and promote