w w w . r b h h . o r g
Revista
Brasileira
de
Hematologia
e
Hemoterapia
Brazilian
Journal
of
Hematology
and
Hemotherapy
Original
article
Comparison
of
microRNA
expression
in
high-count
monoclonal
B-cell
lymphocytosis
and
Binet
A
chronic
lymphocytic
leukemia
Felipe
Magalhães
Furtado
a,∗,
Priscila
Santos
Scheucher
a,
Bárbara
Amélia
Santana
a,
Dalila
Lucíola
Zanette
b,
Rodrigo
do
Tocantins
Calado
a,
Eduardo
Magalhães
Rego
a,
Daniel
Mazza
Matos
c,
Roberto
Passetto
Falcão
aaFaculdadedeMedicinadeRibeirãoPreto(FMRP),RibeirãoPreto,SP,Brazil
bFundac¸ãoOswaldoCruz(FIOCRUZ),Salvador,BA,Brazil
cHospitalUniversitárioWalterCantídio(HUWC),Fortaleza,CE,Brazil
a
r
t
i
c
l
e
i
n
f
o
Articlehistory:
Received30November2016 Accepted24March2017 Availableonline22April2017
Keywords:
MonoclonalB-celllymphocytosis Chroniclymphocyticleukemia microRNA
a
b
s
t
r
a
c
t
Background:EvidencesuggeststhatmonoclonalB-celllymphocytosisprecedesallchronic
lymphocyticleukemiacases,althoughthemolecularmechanismsresponsiblefordisease progressionarenotunderstood.AberrantmiRNAexpressionmaycontributetothe patho-genesisofchroniclymphocyticleukemia.TheobjectiveofthisstudywastocomparemiRNA expressionprofilesofpatientswithBinetAchroniclymphocyticleukemiawiththoseof sub-jectswithhigh-countmonoclonalB-celllymphocytosisandhealthyvolunteers(controls).
Methods:Twenty-onechroniclymphocyticleukemiapatients,12subjectswithmonoclonal
B-celllymphocytosisandtenhealthyvolunteerswereenrolledinthisstudy.Flowcytometry CD19+CD5+-basedcellsortingwasperformedforthechroniclymphocyticleukemiaand
monoclonalB-celllymphocytosisgroupsandCD19+cellsweresortedtoanalyzethecontrol
group.TheexpressionsofmiRNAs(miR-15a,miR-16-1,miR-29b,miR-34a,miR-181a, miR-181bandmiR-155)weredeterminedbyquantitativereversetranscriptasepolymerasechain reaction(qRT-PCR).
Results:Significant differences between the expressions in the chronic lymphocytic
leukemiaandmonoclonalB-celllymphocytosisgroupswererestrictedtotheexpression ofmiR-155,whichwashigherintheformergroup.Acomparisonbetweenhealthy con-trolsandmonoclonalB-celllymphocytosis/chroniclymphocyticleukemiapatientsrevealed highermiR-155andmiR-34alevelsandlowermiR-15a,miR-16-1,miR-181aandmiR-181b inthelattergroup.
∗ Correspondingauthorat:LaboratoriodeHematologia–HospitaldasClinicasdeRibeiraoPreto,BandeirantesAvenue,3900,MonteAlegre,
14051-140RibeirãoPreto,SP,Brazil.
E-mailaddress:[email protected](F.M.Furtado). http://dx.doi.org/10.1016/j.bjhh.2017.03.006
Conclusions: OurresultsshowaprogressiveincreaseofmiR-155expressionfromcontrols tomonoclonalB-celllymphocytosistochroniclymphocyticleukemia.TheroleofmiR-155 inthedevelopmentofovertchroniclymphocyticleukemiainindividualswithmonoclonal B-celllymphocytosismustbefurtheranalyzed.
©2017Associac¸ ˜aoBrasileiradeHematologia,HemoterapiaeTerapiaCelular.Published byElsevierEditoraLtda.ThisisanopenaccessarticleundertheCCBY-NC-NDlicense (http://creativecommons.org/licenses/by-nc-nd/4.0/).
Introduction
Chroniclymphocyticleukemia(CLL)isamalignantneoplasm characterizedbyanexcess ofmonoclonalBlymphocytesin peripheralblood,bonemarrow,thespleenandlymphnodes.1 CLListhemostcommonleukemiaintheWesternworld,with anannualincidenceof5.1cases/100,000people.2
Over the last years, advances in multi-parameter flow cytometryallowedtheidentificationofsmallpopulationsof monoclonalBlymphocytesinthebloodofapparentlyhealthy subjects.3ThepresenceofmonoclonalB-cellsinthe periph-eralbloodofupto5×109/LisclassifiedasmonoclonalB-cell
lymphocytosis(MBL)intheabsenceofotherlymphomatous features.4,5 This condition is found in 0.6–12% of healthy individuals.3,6,7
MBL is classified as high-count or low-count MBL. The first is usually diagnosed in asymptomatic subjects with mildlymphocytosis and the latteroccurs inasymptomatic subjectswithnormalbloodcountswho havebeen submit-tedtoflowcytometryscreening.8–10 ThemostrecentWorld HealthOrganization(WHO)classificationdefines low-count MBLasaperipheralbloodCLL countof<0.5×109/Landno
extramedullarydisease.4AsinCLL,MBLismorecommonin menandinrelativesofCLLpatients11–13 withitsfrequency increasingwithage.11,12,14–17
Virtually,allCLLcasesareprecededbyMBL.18Eventhough themechanismsunderlyingCLLpathogenesisandMBL pro-gressiontoCLLarestillnotwellunderstood,itisspeculated that the initial genetic lesion in MBL may occur in the immaturebonemarrowBcellcompartmentandafterwards repetitiveantigenicstimulationmayinduceadditionalgenetic lesions,eventuallyleadingtoneoplastictransformation.19
MicroRNAs (miRNA) are 19–25 nucleotide single strand RNAsresponsibleforgeneexpressionandcellularmetabolism regulation.20,21ChangesinmiRNAexpressionhavebeen asso-ciatedwithsolidandhematologictumors.21SomemicroRNAs havebeenshowntobeabnormallyexpressedinCLL.22Studies addressingthemolecularandgeneticbasisofMBLmayhelp toelucidateinitialstepsofCLLpathogenesisand,therefore, increaseknowledgeabout CLLorigin.Moreover,onlyafew studieshaveinvestigatedmiRNAsinMBL.9,23Herewe hypoth-esizedthatabnormalitiesinsomemiRNAexpressionsmaybe presentinMBLandmayhavearoleininitialmonoclonalB-cell expansion.
Methods
Cellsamplesfromnormalcontrols,individualswith high-countMBLandCLLpatients
Samples from 12 individuals with high-count MBL and 21 patients withstageACLL accordingtotheBinet classifica-tionwereanalyzed.Allthepatientswerebeingmonitoredin auniversityhospitalinRibeiraoPreto,SaoPaulo,Brazil.Ten healthyindividualswerestudiedasacontrolgroup.Thestudy wasapprovedbytheInstitutionEthicsResearchReviewBoard andwritteninformedconsentwasobtainedfromall partici-pantsinaccordancewiththeDeclarationofHelsinkiandits revisions.
Usingcellsortingbyflowcytometry,CD19+CD5+
lympho-cyteswereisolatedfromtheperipheralbloodofindividuals inthe twostudy groups, andCD19+ lymphocytesfrom the
peripheral blood ofcontrolsas previouslydescribed.24 The meanpercentageofthedesiredcellpopulationafterisolation was89.48%(±8.48%).
The clinical characteristics of the enrolled subjects are summarizedinTable1.
RNAisolationandmiRNAexpression
RNAwasextractedfromCD5+ Blymphocytesisolated from
thestudygroupsandBlymphocytesfromthecontrolgroup, usingTrizol(LifeTechnologies,California,USA).Seven miR-NAsknowntohaveabnormalexpressionsinCLLwereselected foranalysis(miR-15a,miR-16-1,miR-29b,miR-34a,miR-181a, miR-181b and miR-155). miRNA expression was quantified by TaqMan miRNAquantitative reverse-transcription poly-merasechainreaction(qRT-PCR)aspreviouslydescribed.25,26 Briefly, 5ng of total RNA were reverse transcribed using themicroRNAreverse-transcriptionkit(AppliedBio-systems, California, USA) with specific stem-loop primers. qRT-PCR analysis was performed using the miRNA-specific Taqman assay(Applied Bio-systems).Expressionlevelsofthe nucle-olar RNAs RNU24, RNU44 and RNU48 were similar for all groupsandthegeometricmeanoftheirexpressionwasused to normalize the expression of the miRNAs, as previously described27usingthe2−Ctformula.Thecoefficientsof
Table1–Characteristicsoftheenrolledsubjects.
Characteristic Control High-count
monoclonalB-cell lymphocytosis
Chroniclymphocytic leukemia
Numberofsubjects 10 12 21
Mediumage 38 78 70
Range(years) 25–60 57–97 58–81
Gender(n)
Male 2 9 11
Female 8 3 10
Mediumnumberoflymphocytes 2.064 3.933 41.714
Range(L) 1.360–3.300 2.900–5.900 7.500–147.000
%CD5+B-Lymphocytesoftotallymphocytes – 34.9 78.02
Range(%) – 5.97–61.01 58.63–91.09
itsvaluewasconsideredunavailable.AllmiRNAswerestudied induplicate.
Statisticalanalysis
The Kolmogorov–Smirnov normality test was initially per-formedtocomparemiRNAexpressions.Theexpressionsof themiRNAs,miR-15a,miR-16-1,miR-29b,miR-181band miR-155,didnotdeviatefromthenormaldistributionandvariance analysis(ANOVA)wasusedfollowedbytheBonferoniposthoc
test.TheKruskal–Wallisnon-parametrictest(non-parametric ANOVA) was used followed by the Dunn post hoc test for the miRNAs miR-34a and miR-181a that had non-normal distributions.Alltestswithp-values≤0.05 wereconsidered statisticallysignificant.
Results
miRNAexpression
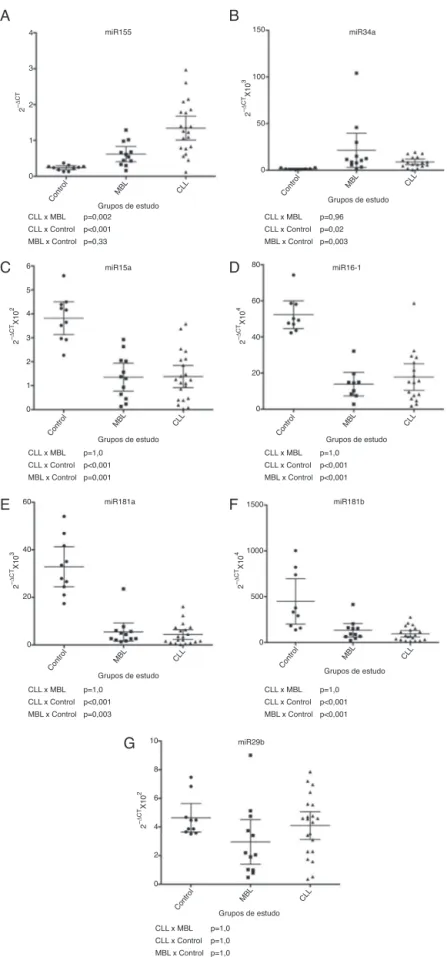
Ofthe sevenmiRNAsdeemed relevantforCLL pathogene-sis,miRNA-155andmiR-34aweredifferentiallyexpressedin theCLL and high-count MBLgroups comparedtothe con-trols(p-value=0.33andp-value=0.003forMBLandcontrols and p-value<0.001 and p-value=0.02 forCLL and controls, respectively)withthehighestvaluesdetectedinCLL(Figure1). miRNA-155expressioninhigh-countMBLhadatendencyto begreater thaninhealthysubjects,but thisdifferencewas notstatisticallysignificant,maybeduetothesmallnumberof samplesstudied(Figure1).miR-34aexpressionwasnot sta-tisticallydifferentbetweenCLLandMBL(Figure1).miR-15a, miR-16-1,miR-181aandmiR-181bweredown-regulatedinCLL andMBL.TheirexpressionwaslowerinCLL patientswhen comparedtocontrols(p-value<0.001forallmiRNAs). High-countMBLsubjectsalsohadlowerexpressionthancontrols formiR-15a,miR-16-1andmiR-181b(p-value=0.003)andfor miR-181a(p-value<0.001)(Figure1).
Expression of miR-29b was similar for all three groups (Figure1).
Discussion
This study with a Binet A CLL group and a high-count MBLgroupdemonstratednewmoleculardifferencesbetween thesetwoentities.
Statisticalsimilaritieshavebeenidentifiedregardingthe frequenciesofimmunoglobulinheavychainvariable(IGHV) genesbetweenhigh-countMBLandtheinitialstagesofCLL, althoughfindingsfromlow-countMBLweredifferentbetween thesegroups.28,29Otherauthorsalsofoundbiological similari-tiesbetweenthesethreeentities.Notonlyhigh-countMBLbut alsolow-countMBLbearcytogeneticabnormalitiescommon inCLL,including13q-,17p-andtrisomy12.6,30–32
Aiming to betterunderstand genetic alterations inCLL, this study comparedtheexpressionsofmiRNAspreviously describedasalteredinthisdiseasewiththeirexpressionin MBL.KnowledgeofmiRNAexpressionindifferentmonoclonal proliferationstagesmayimprovethecomprehensionofCLL pathophysiology.
Severalstudiesusingdifferentmethodologieshave iden-tified overexpressionofmiR-155 inCLL whencomparedto normal controls.23,33–37 Our results confirmed the seminal findingsofFerrajolietal.whoshowedthattheexpressionof miR-155isgreaterinBinetACLLthaninhigh-countMBLand thatitsexpressionintheseindividualsisgreaterthanin nor-malcontrols.23Thus,ourfindingssuggestthatmiR-155may beusedasaprogressionmarkerforMBLindividuals,butthis mustbeconfirmedbyfurtherspecificstudies.
A
miR155
miR15a
miR181a miR181b
miR29b
miR16-1 miR34a
2
–
∆
CT
2
–
∆
CT
X10
3
2
–
∆
CT
X10
2
2
–
∆
CT
X10
3
2
–
∆
CT
X10
4
2
–
∆
CT
X10
2
2
–
∆
CT
X10
4
4 150
100
50
0 3
2
1
0
6 80
60
40
20
0 5
4
3
2
1
0
60 1500
1000
500
0
10
8
6
4
2
0 40
20
0
Grupos de estudo
Grupos de estudo
Grupos de estudo Grupos de estudo
Grupos de estudo
Grupos de estudo Grupos de estudo
CLL x MBL p=0,002 CLL x Control p<0,001 MBL x Control p=0,33
CLL x MBL p=1,0 CLL x Control p<0,001 MBL x Control p=0,001
CLL x MBL p=1,0 CLL x Control p<0,001 MBL x Control p=0,003
CLL x MBL p=1,0 CLL x Control p<0,001 MBL x Control p<0,001
CLL x MBL p=1,0 CLL x Control p=1,0 MBL x Control p=1,0
CLL x MBL p=1,0 CLL x Control p<0,001 MBL x Control p<0,001 CLL x MBL p=0,96 CLL x Control p=0,02 MBL x Control p=0,003
Control Cont
rol
MBL CLL MBL
Control Control
MBL CLL MBL
Cont rol
Cont rol
MBL CL MBL
L
CLL
Control
MBL CLL
CLL CLL
B
C
D
E
F
G
ThisstudyalsoidentifiedmiR-34atobeoverexpressedin CLLwhencomparedtohealthycontrols.35,40,41 ThismiRNA hasbeenassociatedtothe regulationofthe tumorprotein p53(TP53) pathway41–45 and may berelated tofludarabine resistanceinCLL.44
The study that first demonstrated down-regulation of miR-15aand miR-16-1inCLL patientswas alsothefirst to demonstrateabnormal expressionsofmiRNAs incancer.46 Subsequent studies using different techniques have con-firmedthisfinding,particularlyinpatientswithdel13q14.47–49 OtherauthorsfoundsimilarexpressionsofthesemiRNAsin CLLandnormalcontrolsusingmicroarrays35andRT-qPCR.33 Kleinetal.demonstratedthatmousemodelsthathavethe chromosomearea(13q14)responsibleforthetranscriptionof thesemiRNAs deleted developedmonoclonalexpansion of lymphocytesinblood.50
Inthis study,the miRNAsmiR-181aand miR-181b were down-regulatedinCLL whencomparedtonormalcontrols. Other authors found similar results in studies using RT-qPCR,36,48 Northern Blot34 and microarrays.35 Visone et al. demonstratedthat down-regulationofmiR-181b isabetter markerforworseprognosisinCLLthan theIGHVmutation statusandZAP-70expression,andsuggestthatitsexpression shouldbemonitoredinCLLpatients.51
The pattern of miR-29b expression in CLL is not well understood.HeretheresultsforCLLBinetA,high-countMBL andnormalcontrolsweresimilar.Sampathetal.foundthat 70%oftheCLLpatientshaveasimilarexpressiontonormal controls.49Fulcietal.alsohadresultssimilartothisstudy.33 However,Santanam et al. foundthis miRNA tobe overex-pressedinCLL,andZhuetal.foundittobedown-regulated inCLL.48,52 All thesestudieshave heterogeneousgroupsof CLL patientsregarding the stageofthe disease.These dif-ferentresultssuggestthatmiR-29bexpressionmaychange accordingtodiseasestage.However,thishypothesisshould beconfirmedinfuturestudies.
ApartfromthestudyofFerrajolietal.onmiR-155,onlyone othergrouphasevaluatedmiRNAexpressioninhigh-count MBL.23Thus,Morabitoetal.usedmicroarraystostudy indi-vidualspreviouslydiagnosedwithCLLbeforethechangeof diagnosiscriteriaproposedbytheInternationalWorkshopon ChronicLymphocyticLeukemiain2008.9,53Theyfound miR-130awastheonlyonewithdifferentexpressionsbetweenCLL and high-countMBL.Using another techniqueand a more heterogeneoushigh-countMBLpopulation,thisstudyfound theexpressionsofthemiR-34a,miR-15a,miR-16-1,miR-181a, miR-181bandmiR-29btobesimilarinCLLandhigh-count MBL.
Theresultsreportedhereshouldbeconfirmedbyfurther studies, since the groups were small and more individ-ualsmustbestudiedtoallowrobustconclusions.Moreover, althoughthe CLL and MBL groups were composed mainly of older individuals and the MBL group mainly of males, ourcontrolgroupwascomposedmainlyoffemalesandthe medium age for this group was lower than for the study groups,althoughtherearenodatasuggestingthatthelevels ofthesemiRNAschangewithsexandage.
Inconclusion,thefindingthatsomemiRNAshave abnor-malexpressionsinMBLandthatthisconditionisalsopresent inCLLsuggeststhatthesegeneticchangesmaybepartofthe
initialeventsresponsibleformonoclonalCD5+B-cell
prolifer-ation.ThedifferentialexpressionofmiR-155inMBLandCLL suggestsitsroleinsignalingpathwaysthatareimportantto thedevelopmentofthedisease.
Author
contributions
FMF,DLZ,RTCSR,EMR,DMMandRPFdesignedtheresearch protocols.FMF,PSSandBSperformedtheresearch.FMF,BS, DMMandRPFanalyzedthedata.FMF,RTCSR,EMR,DMMand RPFwrotethemanuscript.
Conflict
of
interests
Theauthorsdeclarenoconflictsofinterest.
Acknowledgements
This research was supported by the Conselho Nacional de Desenvolvimento Cientifico e Tecnologico(CNPQ) grant 573.754/2008-0.
r
e
f
e
r
e
n
c
e
s
1.Müller-HermelinkHK,MontserratE,CatovskyD,CampoE, HarrisNL,SteinH.Choniclymphocyticleukaemia/small lymphocyticlymphoma.In:SwerdlowSH,CampoE,Harris NL,JaffeES,PileriSA,SteinH,etal.,editors.WHO classificationoftumorsofhaematopoieticandlymphoid tissues.4thedLyon:WorldHealthOrganizationClassification ofTumors.InternationalAgencyforResearchonCancer; 2008.p.180–2.
2.DoresGM,AndersonWF,CurtisRE,LandgrenO,Ostroumova E,BluhmEC,etal.Chroniclymphocyticleukaemiaandsmall lymphocyticlymphoma:overviewofthedescriptive epidemiology.BrJHaematol.2007;139(5):809–19.
3.RawstronAC,GreenMJ,KuzmickiA,KennedyB,FentonJA, EvansPA,etal.MonoclonalBlymphocyteswiththe characteristicsofindolentchroniclymphocyticleukemiaare presentin3.5%ofadultswithnormalbloodcounts.Blood. 2002;100(2):635–9.
4.SwerdlowSH,CampoE,PileriSA,HarrisNL,SteinH,Siebert R,etal.The2016revisionoftheWorldHealthOrganization classificationoflymphoidneoplasms.Blood.
2016;127(20):2375–90.
5.MartiGE,RawstronAC,GhiaP,HillmenP,HoulstonRS,KayN, etal.DiagnosticcriteriaformonoclonalB-celllymphocytosis. BrJHaematol.2005;130(3):325–32.
6.RawstronAC,BennettFL,O’ConnorSJ,KwokM,FentonJA, PlummerM,etal.MonoclonalB-celllymphocytosisand chroniclymphocyticleukemia.NEnglJMed.
2008;359(6):575–83.
7.D’ArenaG,MustoP.MonoclonalB-celllymphocytosis.Transl MedUniSa.2014;8:75–9.
8.GhiaP,Caligaris-CappioF.MonoclonalB-celllymphocytosis: righttrackorredherring?Blood.2012;119(19):4358–62. 9.MorabitoF,MoscaL,CutronaG,AgnelliL,TuanaG,Ferracin
10.RawstronAC.MonoclonalBcelllymphocytosis–whatdoesit reallymean?CurrHematolMaligRep.2013;8(1):52–9. 11.RawstronAC,YuilleMR,FullerJ,CullenM,KennedyB,
RichardsSJ,etal.InheritedpredispositiontoCLLisdetectable assubclinicalmonoclonalB-lymphocyteexpansion.Blood. 2002;100(7):2289–90.
12.MatosDM,IsmaelSJ,ScrideliCA,deOliveiraFM,RegoEM, FalcaoRP.MonoclonalB-celllymphocytosisinfirst-degree relativesofpatientswithsporadic(non-familial)chronic lymphocyticleukaemia.BrJHaematol.2009;147(3):339–46. 13.GoldinLR,LanasaMC,SlagerSL,CerhanJR,VachonCM,
StromSS,etal.CommonoccurrenceofmonoclonalB-cell lymphocytosisamongmembersofhigh-riskCLLfamilies.BrJ Haematol.2010;151(2):152–8.
14.ShanafeltTD,KayNE,JenkinsG,CallTG,ZentCS,JelinekDF, etal.B-cellcountandsurvival:differentiatingchronic lymphocyticleukemiafrommonoclonalB-celllymphocytosis basedonclinicaloutcome.Blood.2009;113(18):4188–96. 15.GhiaP,PratoG,ScielzoC,StellaS,GeunaM,GuidaG,etal.
MonoclonalCD5+andCD5−B-lymphocyteexpansionsare frequentintheperipheralbloodoftheelderly.Blood. 2004;103(6):2337–42.
16.MulliganCS,ThomasME,MulliganSP.Monoclonal
B-lymphocytosis:demographics,natureandsubclassification in414communitypatients.LeukLymphoma.
2011;52(12):2293–8.
17.KernW,BacherU,HaferlachC,DickerF,AlpermannT, SchnittgerS,etal.MonoclonalB-celllymphocytosisisclosely relatedtochroniclymphocyticleukaemiaandmaybebetter classifiedasearly-stageCLL.BrJHaematol.2012;157(1):86–96. 18.LandgrenO,AlbitarM,MaW,AbbasiF,HayesRB,GhiaP,etal.
B-cellclonesasearlymarkersforchroniclymphocytic leukemia.NEnglJMed.2009;360(7):659–67.
19.ChiorazziN,RaiKR,FerrariniM.Chroniclymphocytic leukemia.NEnglJMed.2005;352(8):804–15.
20.Alvarez-GarciaI,MiskaEA.MicroRNAfunctionsinanimal developmentandhumandisease.Development.
2005;132(21):4653–62.
21.ChenB,LiH,ZengX,YangP,LiuX,ZhaoX,etal.Rolesof microRNAoncancercellmetabolism.JTranslMed. 2012;10:228.
22.WardBP,TsongalisGJ,KaurP.MicroRNAsinchronic lymphocyticleukemia.ExpMolPathol.2011;90(2):173–8. 23.FerrajoliA,ShanafeltTD,IvanC,ShimizuM,RabeKG,
NouraeeN,etal.PrognosticvalueofmiR-155inindividuals withmonoclonalB-celllymphocytosisandpatientswithB chroniclymphocyticleukemia.Blood.2013;122(11):1891–9. 24.BasuS,CampbellHM,DittelBN,RayA.Purificationofspecific
cellpopulationbyfluorescenceactivatedcellsorting(FACS).J VisExp.2010;41.
25.ChenC,RidzonDA,BroomerAJ,ZhouZ,LeeDH,NguyenJT, etal.Real-timequantificationofmicroRNAsbystem-loop RT-PCR.NucleicAcidsRes.2005;33(20):e179.
26.SchmittgenTD,LeeEJ,JiangJ,SarkarA,YangL,EltonTS,etal. Real-timePCRquantificationofprecursorandmature microRNA.Methods.2008;44(1):31–8.
27.VandesompeleJ,DePreterK,PattynF,PoppeB,VanRoyN,De PaepeA,etal.Accuratenormalizationofreal-time
quantitativeRT-PCRdatabygeometricaveragingofmultiple internalcontrolgenes.GenomeBiol.2002;3(7).Research0034. 28.DagklisA,FaziC,SalaC,CantarelliV,ScielzoC,MassacaneR,
etal.Theimmunoglobulingenerepertoireoflow-count chroniclymphocyticleukemia(CLL)-likemonoclonalB lymphocytosisisdifferentfromCLL:diagnosticimplications forclinicalmonitoring.Blood.2009;114(1):26–32.
29.VardiA,DagklisA,ScarfoL,JelinekD,NewtonD,BennettF, etal.ImmunogeneticsshowsthatnotallMBLareequal:the
largertheclone,themoresimilartoCLL.Blood. 2013;121(22):4521–8.
30.NietoWG,AlmeidaJ,RomeroA,TeodosioC,LopezA, HenriquesAF,etal.Increasedfrequency(12%)ofcirculating chroniclymphocyticleukemia-likeB-cellclonesinhealthy subjectsusingahighlysensitivemulticolorflowcytometry approach.Blood.2009;114(1):33–7.
31.RossiD,SozziE,PumaA,DePaoliL,RasiS,SpinaV,etal.The prognosisofclinicalmonoclonalBcelllymphocytosisdiffers fromprognosisofRai0chroniclymphocyticleukaemiaandis recapitulatedbybiologicalriskfactors.BrJHaematol. 2009;146(1):64–75.
32.FaziC,ScarfoL,PecciariniL,CottiniF,DagklisA,JanusA,etal. Generalpopulationlow-countCLL-likeMBLpersistsover timewithoutclinicalprogression,althoughcarryingthesame cytogeneticabnormalitiesofCLL.Blood.2011;118(25):6618–25. 33.FulciV,ChiarettiS,GoldoniM,AzzalinG,CarucciN,Tavolaro
S,etal.QuantitativetechnologiesestablishanovelmicroRNA profileofchroniclymphocyticleukemia.Blood.
2007;109(11):4944–51.
34.MartonS,GarciaMR,RobelloC,PerssonH,TrajtenbergF, PritschO,etal.SmallRNAsanalysisinCLLrevealsa deregulationofmiRNAexpressionandnovelmiRNA candidatesofputativerelevanceinCLLpathogenesis. Leukemia.2008;22(2):330–8.
35.PallaschCP,PatzM,ParkYJ,HagistS,EggleD,ClausR,etal. miRNAderegulationbyepigeneticsilencingdisrupts suppressionoftheoncogenePLAG1inchroniclymphocytic leukemia.Blood.2009;114(15):3255–64.
36.LiS,MoffettHF,LuJ,WernerL,ZhangH,RitzJ,etal.MicroRNA expressionprofilingidentifiesactivatedBcellstatusin chroniclymphocyticleukemiacells.PLoSOne.2011;6(3). 37.VargovaK,CurikN,BurdaP,BasovaP,KulvaitV,PospisilV,
etal.MYBtranscriptionallyregulatesthemiR-155hostgene inchroniclymphocyticleukemia.Blood.2011;117(14):3816–25. 38.DangerR,BrazaF,GiralM,SoulillouJP,BrouardS.MicroRNAs
majorplayersinBcellshomeostasisandfunction.Front Immunol.2014;5:98.
39.SchottJ,StoecklinG.NetworkscontrollingmRNAdecayinthe immunesystem.WileyInterdiscipRevRNA.2010;1(3):432–56. 40.ZanetteDL,RivadaviaF,MolfettaGA,BarbuzanoFG,
Proto-SiqueiraR,Silva-JrWA,etal.miRNAexpressionprofiles inchroniclymphocyticandacutelymphocyticleukemia.Braz JMedBiolRes.2007;40(11):1435–40.
41.AsslaberD,PinonJD,SeyfriedI,DeschP,StocherM,TinhoferI, etal.microRNA-34aexpressioncorrelateswithMDM2 SNP309polymorphismandtreatment-freesurvivalinchronic lymphocyticleukemia.Blood.2010;115(21):4191–7.
42.BommerGT,GerinI,FengY,KaczorowskiAJ,KuickR,LoveRE, etal.p53-mediatedactivationofmiRNA34candidate tumor-suppressorgenes.CurrBiol.2007;17(15):1298–307. 43.HermekingH.ThemiR-34familyincancerandapoptosis.
CellDeathDiffer.2010;17(2):193–9.
44.MerkelO,AsslaberD,PinonJD,EgleA,GreilR.Interdependent regulationofp53andmiR-34ainchroniclymphocytic leukemia.CellCycle.2010;9(14):2764–8.
45.DufourA,PalermoG,ZellmeierE,MellertG,
Duchateau-NguyenG,SchneiderS,etal.InactivationofTP53 correlateswithdiseaseprogressionandlowmiR-34a expressioninpreviouslytreatedchroniclymphocytic leukemiapatients.Blood.2013;121(18):3650–7.
46.CalinGA,DumitruCD,ShimizuM,BichiR,ZupoS,NochE, etal.Frequentdeletionsanddown-regulationofmicro-RNA genesmiR15andmiR16at13q14inchroniclymphocytic leukemia.ProcNatlAcadSciUSA.2002;99(24):15524–9. 47.CalinGA,LiuCG,SevignaniC,FerracinM,FelliN,DumitruCD,
chroniclymphocyticleukemias.ProcNatlAcadSciUSA. 2004;101(32):11755–60.
48.ZhuDX,MiaoKR,FangC,FanL,ZhuW,ZhuHY,etal. AberrantmicroRNAexpressioninChinesepatientswith chroniclymphocyticleukemia.LeukRes.2011;35(6):730–4. 49.SampathD,LiuC,VasanK,SuldaM,PuduvalliVK,Wierda WG,etal.Histonedeacetylasesmediatethesilencingof miR-15a,miR-16,andmiR-29binchroniclymphocytic leukemia.Blood.2012;119(5):1162–72.
50.KleinU,LiaM,CrespoM,SiegelR,ShenQ,MoT,etal.The DLEU2/miR-15a/16-1clustercontrolsBcellproliferationand itsdeletionleadstochroniclymphocyticleukemia.Cancer Cell.2010;17(1):28–40.
51.VisoneR,VeroneseA,RassentiLZ,BalattiV,PearlDK,Acunzo M,etal.miR-181bisabiomarkerofdiseaseprogressionin chroniclymphocyticleukemia.Blood.2011;118(11):3072–9. 52.SantanamU,ZanesiN,EfanovA,CostineanS,Palamarchuk
A,HaganJP,etal.Chroniclymphocyticleukemiamodeledin mousebytargetedmiR-29expression.ProcNatlAcadSciUS A.2010;107(27):12210–5.
53.HallekM,ChesonBD,CatovskyD,Caligaris-CappioF,Dighiero G,DohnerH,etal.Guidelinesforthediagnosisandtreatment ofchroniclymphocyticleukemia:areportfromthe