47
O
r
IG
IN
A
l
A
r
T
IC
lE
ABSTRACT
Leprosy is still a worldwide public health problem. Brazil and India show the highest prevalence ratesofthedisease.NaturalinfectionofarmadillosDasypus novemcinctuswithMycobacterium leprae
hasbeenreportedinsomeregionsoftheUnitedStates.Identificationofbacilliisdifficult,particu-larlyduetoitsinabilitytogrowinvitro.Theuseofmoleculartoolsrepresentsafastandsensitive alternativemethodfordiagnosisofmycobacteriosis.Inthepresentstudy,thediagnosticmethods usedwerebacilloscopy,histopathology,microbiology,andPCRusingspecificprimersforM. leprae
repetitivesequences.PCRwereperformedusinggenomicDNAextractedfrom138samplesofliver, spleen,lymphnodes,andskinof44D. novemcinctus, Euphractus sexcinctus, Cabassous unicinctus,
andC. tatouayarmadillosfromtheMiddleWesternregionofthestateofSãoPauloandfromthe experimentalstationofEmbrapaPantanal,locatedinPantanaldaNhecolândiaofMatoGrossodo Sulstate.Also,themolecularanalysisof19samplesfrominternalorgansofotherroadkilledspecies ofwildanimals,suchasNasua nasua(ring-tailedcoati),Procyon cancrivoros(hand-skinned), Cer-docyon thous(dog-pity-bush),Cavia aperea(restlesscavy),Didelphis albiventris(skunk),Sphigurrus spinosus(hedgehog),andGallictis vittata (ferret)showedPCRnegativedata.Noneofthe157ana- lyzedsampleshadshownnaturalmycobacterialinfection.Onlythearmadilloinoculatedwithmate-rialcollectedfromuntreatedmultibacillaryleprosypatientpresentedPCRpositiveanditsgenomic sequencingrevealed100%identitywithM. leprae.Accordingtothesepreliminarystudies,basedon theusedmethodology,itispossibletoconcludethatwildmammalsseemnottoplayanimportant roleintheepidemiologyofleprosyintheMiddleWesternregionoftheSãoPaulostateandinthe PantanalofMatoGrossodoSulstate.
Keywords:Dasypus novemcinctus, Euphractus sexcinctus, Cabassous tatouay, Mycobacterium leprae,
eco-epidemiology,wildmammals.
[Braz J Infect Dis 2010;14(1):47-53]©ElsevierEditoraLtda.
Search for
Mycobacterium leprae
in wild mammals
Authors
SílviaCristinaBarboza Pedrini,MD1
PatríciaSammarcoRosa, MD1
ÍsisMeriMedri,MD2
GuilhermeMourão,MD2
EduardoBagagli, Professor3
CarlosAlbertode MagalhãesLopes, Professor3
1InstitutoLaurodeSouza Lima,Bauru–SãoPaulo, Brazil
2EmbrapaPantanal-
Corumbá,MS
3Microbiologyand
ImmunologyDepartment, InstitutodeBiociências, UniversidadeEstadual Paulista(UNESP)- Botucatu,SãoPaulo,Brazil.
Submittedon:08/03/2009 Approvedon:11/22/2009
Correspondence to:
SílviaCristinaBarboza Pedrini
RodoviaComandante JoãoRibeirodeBarros, km225/226
POBox3021 Bauru,SP,Brasil CEP:17034-971 E-mail:silviapedrini@ yahoo.com.br
Wedeclarenoconflict ofinterest.
INTRODUCTION
Leprosy,amillenarianillnessdescribedinChi-na,IndiaandEgyptat600aC,1isstillconsidered
aseriousPublicHealthproblem.M. leprae,the etiologicalagent,differentlyfromotherin vitro
culturedMycobacteria whose ecological niches are very well defined, has still important un-knownpointsinitseco-epidemiology.
Generally,Mycobacteriaaremicroorganisms highlyadaptedtotheenvironmentwherethey live. ParticularlyM. leprae, which is a bacillus highly adapted to the human being to whom theyrelyforitssurvivalandperpetuation.
Inter-human leprosy transmission is known
BesidesfailuresinisolatingM. leprae fromtheenviron- ment,thelackofepidemicoutbreaksandthelowreproduc-ibilityofnaturalinfectioncasesmakeitdifficulttoelucidate thehabitatofthisorganism.Theexactsourceofinfection isnotknown,whichseemstoreflectanextremelyrestricted ecologicalnicheofthismicroorganism.
With the recent progress of molecular biology tech-niques, some aspects of the biology and ecology ofM. leprae have been cleared. Molecular markers have been used thoroughly in leprosy diagnosis and research. PCR productscanbeanalyzedbyagarosegelelectrophoresisor slot-blot hybridization, using specific probes, such as the 212bpone,whichbindstoaninternalsegmentofthe360 bpregionofM. leprae18kDagene.3 Analysisofpolymor-phisms of PCR fragments after enzyme digestion (PRA) for the hsp65 gene, present in allmycobacteria4 was used
alsoforthecharacterizationofM. leprae.5WoodsandCole (1989),basedtheirPCRonthespecificrepetitiveelement ofM. leprae(RLEP)demonstratedbyvisualizationofa372 bpproduct.6 ThismarkerwaslaterusedforDNAhybrid-izationbySantoset al.(1993).7 Thesearchforthepatho-geninwildanddomesticanimalsispracticallyunexplored. Exceptfornon-humanprimates,theuniqueanimalgroup inwhichM. leprae growssuccessfullyisinarmadillos,es-peciallythenine-bandedD. novemcinctus.8,9
Experimental studies involving armadillosD. novem-cinctus show that these animals develop a disseminated form of disease. This animal model is valuable because wheninoculatedwithM. leprae,armadillosproducelarge amountsofbacillithatcanbeusedinthedevelopmentof diagnostictestsforpreparationofMitsudaantigenandan-tigensfractionsproduction.10-12
ArmadillosbelongtotheOrderCingulata,13whichhad its evolutionary development centered in South America sincethePaleoceneera,around65millionyearsago,when theSouthAmericancontinentwasalreadyseparatedfrom the African continent, but still disconnected from North America.14,15Thenine-bandedarmadillowasintroducedin NorthAmericaabout17014,16 andindigenousinfectioninar-madilloswasobservedmorerecently.17,18Significantprogress inM. leprae global eco-epidemiology knowledge was also possibleafterstudiesofmolecularandgenomiccharacter-izationofseveralstrainscomingfromdifferentpartsofthe world. Through single nucleotides polymorphism analysis (SNPs),itwasverifiedthattheoriginofM. lepraeremounts totheAfricancontinent,havingbeendisseminatedfromAf-ricatoAsiaandlatertoEurope.Thearrivalofthebacillusin Americamusthaveoccurredrecently,about500yearsago, duringtheNewWorldcolonization.19
Inthepresentstudy,usingmoleculartoolsindifferent speciesofwildanimalsandinfourspeciesofarmadillos(D. novemcinctus,Euphractus sexcinctus,Cabassous tatouayand
C.unicinctus),weaimedatsearchingforM. lepraecarriers anddiscusstheroleoftheseanimalhostsintheepidemiol-ogy of leprosy in the Middle-west area of São Paulo state, where leprosy is endemic, and also in Pantanal of Mato GrossodoSulstate,whereleprosyisstillhiperendemic.
MATERIAL AND METHODS
Animals
Liver fragment from an M. leprae experimentally infected ar-madillo.AspositivecontrolforPCRreaction,aM. leprae
infectedliverfragmentwasusedforDNAextraction.AD. novemcinctusarmadillowaspreviouslyinoculatedwithM. lepraesuspensionbysubcutaneousandintravenousroute (108 bacilli/mL). This animal was maintained captive for 20monthsandevaluatedbimonthlyuntiltheappearance ofdisseminateddisease.Afternecropsytheanimalshowed granulomasintheliver,spleen(3.06x109bacilli/g),lymph nodes,lungs,adrenalsglands,andskin.20,21
Wild armadillos. A total of 44 wild armadillos of four different species (D. novemcinctus n = 18; E.sexcintus
n=22;C.tatouay n=02;C.unicinctusn=02),youngadults, males and females, weights varying from 3.5 kg to 6.5 kg were studied. The animals from São Paulo were captured intheMiddleWesternareaofthestateofSãoPaulo,inthe municipalitiesofBotucatu(22nd56’15”S,48th26’15”W), Pardinho, São Manuel (23rd 03’ 45” S, 48th. 18’, 45” W), Manduri(23rd03’45”S,49th18’45”W)andBauru(22nd 18’41”S,49th03’45”W).Besides,wehadcapturedarma-dillosfromtheNhumirimranch,anexperimentalstation of Embrapa Pantanal, located in the Pantanal da Nhe-colândiaoftheMatoGrossodoSulstate(18º59’S;56º 39’W).Fromtheseanimalsatotalof138sampleswere obtained:26earfragments,32feces,21nostrilswab,20 blood,and39internalorgans(livern=15;spleenn=7; lymphnodesn=10;kidneyn=1;adrenalglandsn=1, andlungsn=2).
Other wild animal species. Ten road killed animals werealsoanalyzed(Ring-TailedCoatiNasuanasuan= 02; skunkDidelphis albiventris n = 1; hedgehog Sphig-urrus spinosusn=01;hand-skinnedProcyon cancrivoros
n=01;restlesscavyCavia aperean=1;ferretsGallictis vittatan=2,anddog-pity-bushCerdocyon thousn=2), beingatotalof19samplescollected.
Amount of animals .Theapparentlysmallsamplecol-lection is justified by the Ambient Protection Lays of Brazilianwildanimal’sbiodiversity.Soitisn’tpermitted tousealotofspecimens,accordingtotheBrazilianIn-stituteofEnvironmentandRenewableNaturalResources (IBAMA)andtheEthicPrinciplesCommittee.
withtiletamineandzolazepam(5.0mg/kg/I.M)andsub-49 Braz J Infect Dis 2010;14(1):47-53
Pedrini,Rosa,Medri et al.
mitted to subclavian vein puncture for the total blood collection.Allanimalswerecapturedundersupervision oftheBrazilianInstituteofEnvironmentandRenewable NaturalResources(IBAMA),licensenumber187/05and 006/2007B.
Necropsy and sampling. From the wild armadillos, samples of blood, nostril swab, liver, spleen, mesenter-iclymphnodes,lungs,adrenalglands,kidneys,andear fragmentswerecollected.DNAsamplesoffecesofarma-dilloweresuppliedfromtheDNABankoftheMicology LaboratoryoftheDepartmentofMicrobiologyandIm-munology (Institute of Biosciences, UNESP, Botucatu, SP,Brazil).Fragmentsofliver,spleen,lymphnodes,and skin were collected also for histopathological examina-tionusingtheFaraco-Fitestainingmethod.
Microbiological methods
Organ decontamination and digestion.Organsfragments wereweightedandgroundin2,0mLofsteriledistilled water. Samples were decontaminated by the Petroff Method22 and the sediment was used for culture and bacilloscopy.
Semi-quantitative bacilloscopy.Smearof10μLofthe sediment obtained after processing of organ samples werecoldstainedbyZiehl-Neelsenstainingmethod.One hundred fields of the slides were examined under light microscope (100x magnification). The results were ex- pressedbythesemi-quantitativemethodasfortubercu-losisdiagnosisandforothermycobacteriosis,according tothefollowingcriteria:
• (-)negativeforacidfastbacilli(AFB)in100ex- aminedfields;
• (+)lessthanaAFB/fieldin100examinedfields; • (++)from1to10AFB/fieldin50examined fields;
• (+++)morethan10AFB/fieldin20examined fields.
Culture. Liver, spleen and mesenteric lymph nodes were inoculated in LJ culture medium with 2,5% ferric citrateinduplicatetubeswithtightlids.Thetubeswere incubated at 37ºC and observed daily in the first week and then weekly up to 90 days until the appearance of colonies. Cultures were considered negative in the ab-senceofgrowth.
Molecular methods
DNA extraction. After criofracture, the organ samples (about 300 mg) were submitted to a pre-treatment,23,24 before digestion with Proteinase-K. Thus, after mac-eration with liquid nitrogen (N2 liq), the material was transferred to 1.5 mL micro tube with 600 μL of Tris-HCl 10mM; EDTA 1mM; pH8,0 (TE). Samples were
homogenized in vortex and centrifuged at 14,000 rpm/ 25ºC/5 min. The supernatants were discarded and the sedimentssuspendedin600μLofTEfollowingnewcen-trifugationinthesameconditions.Thesedimentwasthen suspended in 600 μL of lisis buffer (Tris-HCl 100 mM, EDTA0,125mM,SDS1,0%,2-mercaptoetanol0,2%and waterMilli-Qq.s.p.)andincubatedat56ºCfor1h.After thisperiodsampleswereincubatedinwaterbathat95ºC/ 10 min. Soon after, 20 μL of Proteinase-K (20 mg/mL) were added to each microtube, which was incubated at 56ºC overnight. Afterwards, 500 μL of phenol, chloro-form,andisoamilicalcoholmixturewereadded(25:24:1) to tubes, they were homogenized and centrifuged at 25º C/13.000 rpm/20 min. The supernatants were care-fullycollectedandtransferredintonewmicrotubes,and the phenol/chloroform/isoamilic alcohol extraction was repeatedoncemore.DNAwasprecipitatedwithisopro-panoland10μLofsodiumacetate3M,at-20ºCfor30 minutes. The material obtained was centrifuged at
4ºC/13.000 rpm/20 min and the “pellet” washed twice
withethanol70%.Afterdryingat37ºCfor1h,DNAwas eluted in 100 μL of sterile Milli-Q water. DNA visual-ization and quantification were done in 1% agarose gel (Sigma,Oakland,USES)stainedwithethidiumbromide (10μg/mL).LowMassmolecularweightmarkerwasused (Invitrogen).
PCR amplification. For the amplification of theM. leprae specific repetitive sequence, a set of primers 5’-GCACGTAAGCCTGTCGGTGG-3’ and 5’-CGGCCG-GATCCTCGATGCAC-3’wereused.6,25PCRamplifications were performed in a Thermal Cycler PTC-100TM - 480 model (Peltier-Effect Cycling MJ Research, USA). DNA (10ng)wasmixedwith200mMofeachdeoxynucleotide triphosphate,10mMofeachprimer,50mMKCl,1,5mM MgCl2,10mMTris-HCl(pH9,0),1UofTaqpolymerase (GE Healthcare) and water to a final volume of 25 mL. Cyclingconsistedof92ºCfor3min,followedby40cycles of2.5minat55ºC,2minat72ºCand1.5minat92ºC, and a final extension cycle at 72º C for 7 min.7 Samples were analyzed in a 1.5% agarose gel (Sigma, Oakland, USA)stainedwithethidiumbromide(10mg/mL).
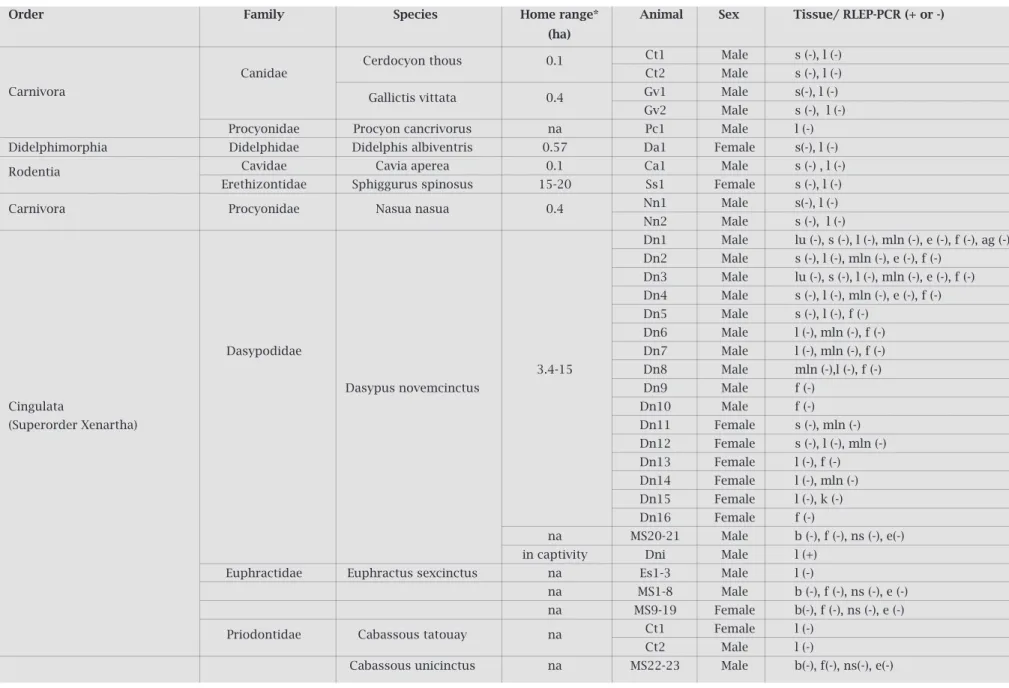
Table 1. Taxonomical data of the evaluated animals, including the species home range, sex, evaluated tissues, and RLEP-PCR results
Order Family Species Home range* Animal Sex Tissue/ rlEP-PCr (+ or -)
(ha)
Cerdocyon thous 0.1 Ct1 Male s (-), l (-)
Canidae Ct2 Male s (-), l (-)
Carnivora Gallictis vittata 0.4 Gv1 Male s(-), l (-)
Gv2 Male s (-), l (-)
Procyonidae Procyon cancrivorus na Pc1 Male l (-)
Didelphimorphia Didelphidae Didelphis albiventris 0.57 Da1 Female s(-), l (-)
Rodentia Cavidae Cavia aperea 0.1 Ca1 Male s (-) , l (-)
Erethizontidae Sphiggurus spinosus 15-20 Ss1 Female s (-), l (-)
Carnivora Procyonidae Nasua nasua 0.4 Nn1 Male s(-), l (-)
Nn2 Male s (-), l (-)
Dn1 Male lu (-), s (-), l (-), mln (-), e (-), f (-), ag (-) Dn2 Male s (-), l (-), mln (-), e (-), f (-)
Dn3 Male lu (-), s (-), l (-), mln (-), e (-), f (-) Dn4 Male s (-), l (-), mln (-), e (-), f (-) Dn5 Male s (-), l (-), f (-)
Dn6 Male l (-), mln (-), f (-)
Dasypodidae Dn7 Male l (-), mln (-), f (-)
3.4-15 Dn8 Male mln (-),l (-), f (-)
Dasypus novemcinctus Dn9 Male f (-)
Cingulata Dn10 Male f (-)
(Superorder Xenartha) Dn11 Female s (-), mln (-)
Dn12 Female s (-), l (-), mln (-) Dn13 Female l (-), f (-) Dn14 Female l (-), mln (-) Dn15 Female l (-), k (-)
Dn16 Female f (-)
na MS20-21 Male b (-), f (-), ns (-), e(-)
in captivity Dni Male l (+)
Euphractidae Euphractus sexcinctus na Es1-3 Male l (-)
na MS1-8 Male b (-), f (-), ns (-), e (-) na MS9-19 Female b(-), f (-), ns (-), e (-)
Priodontidae Cabassous tatouay na Ct1 Female l (-)
Ct2 Male l (-)
Cabassous unicinctus na MS22-23 Male b(-), f(-), ns(-), e(-)
lu – lung, s-spleen, b-blood, l-liver, k-kidney, h-heart, mln-mesenteric lymph node, ag-adrenal gland, e-ear, f-feces, ns-nostril swab, na: not available. * According to Eisenberg & Redford 1999.
M
yc
ob
ac
te
rium le
p
rae
inw
51 Braz J Infect Dis 2010;14(1):47-53
RESULTS
TherewasnotgrowthofMycobacteria insamplesincubat-edat37ºC.Nomycobacteriawerevisualizedintheorgan samples submitted to histopathological examination and bacilloscopy.
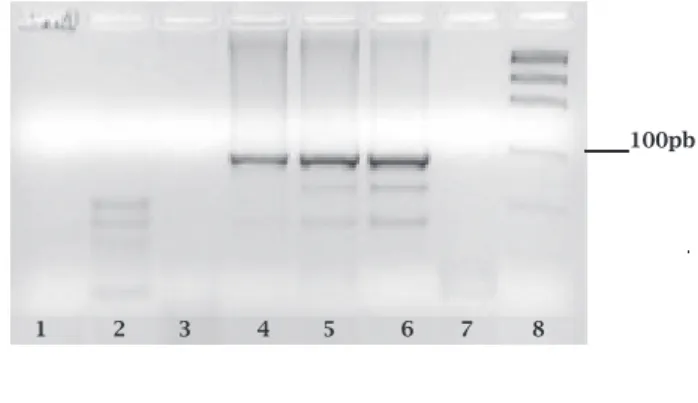
ThePCRreactionswerenegativeforM. lepraeinallwild mammals(Table1).26,27 Onlytheliversampleoftheexperi-mentallyinoculatedD. novemcinctuswasPCRpositivefor the RLEP region specific forM. leprae.A 372bp fragment could be visualized in the agarose gel. (Figure 1). The ge-nomicsequencingoftheamplifiedproductrevealed100% ofhomologywithM. leprae.
DISCUSSION
Genomic comparative studies show thatM. leprae pres-ents signatures characteristic of an extremely specialized and highly host dependent pathogen. The comparative analysisoftheM. tuberculosisgenome(completegenome with 4.411.532bp) withM. leprae genome (complete ge-nome with 3.268.203bp) reveals that only 49,5% of the
M. leprae genome contain genes that codify proteins and 27%ofthenrepresent“pseudogenes”,inotherwords,genes thatwereturnedoffinM. lepraebutarestillfunctionalin
M. tuberculosis.28
Similarly to leprosy, the Buruli ulcer caused by Myco-bacterium ulcerans isaseriousskindiseaseinhumansand its incidence overcomes that of leprosy in countries, such asAustraliaandPapuaNewGuinea.29TheM. ulcerans ge-nomeresemblestheM. lepraegenome,showingsignificant loss of genetic redundancy and of metabolic pathways.30 Through comparative analysis of genomic sequences, it wasdiscoveredthatM. ulcerans emergedfromM. marinum
by the acquisition of genes responsible for mycolactone-A
production, an immunosuppressive cytotoxin that pro- vokesaseriousnecroticulcerationinthesubcutaneoustis- sue.Theevolutionaryreductionandthegenomicrearrange-ment remodeledM. marinum, a fish and toads pathogen, intoM. ulcerans.Thisorganismisapparentlyadaptednow toadarkandaerobicatmosphere,whereitsreducedantige-nicity,slowgrowth,andmycolactoneproductionprovided advantagesforitssurvival.29
Thisprocessofgenomicreductionisalsodocumentedin otherobligateintracellularparasites,suchasRickettsiaand
Chlamydia spp.inwitchsomegenesbecameinactivated,once theirfunctionsarenolongernecessaryinhighlyspecialized niches.AlsoYersinia pestisapparentlydivergedrecentlyfrom Y.pseudotuberculosistoengagethemidgutoffleas,whereas
Bordetella pertussisderivedfromB. bronchisepticatobecome anobligatoryhumanpathogen.30
M. leprae may have followed this same evolutionary trend,withaminimumamountofactivegenesneededfor its adaptation to the host, without, losing their character-istic pathogenicity. This process of evolutionary reduction indicatesthatthemicroorganismtriedrearrangementsand deletionsofitsgenomeandsufferedanevolutionaryadap-tationprocessextremelywellestablishedinhumanhost.
Inthisway,theHansen’sbacillushasbeencohabitating withmanformorethan2,600years,anditremains,still today,intheseconddecadeofthe21stcentury,asanex- tremelypreoccupyingpathogenintropicalclimatecoun-tries,likeBrazil,India,Madagascar,andAfricancountries, withhighdetectionrates(morethan690,000newcasesof the disease registered annually in the world). Only, after the1980s,withthesuccessofmultidrugtherapy,transmis-sionrateswerereduced.
Despite reports of the existence of natural disease in some primates species in Africa (mangabeys monkeys,
Cynomolgus,chimpanzees,9andinarmadillosD.
novemcinc-tusincertaingeographicalareasoftheUnitedStates(Texas andLouisiana),17,18fewstudieswerecarriedoutinBrazil.We didn’tfindanynaturalinfectioninthestudiedwildmam- malsintheMiddleWesternregionofSãoPaulostate;nei-therinMatoGrossodoSulPantanal.Althoughotherwild species have never been implicated withM. leprae infec-tion,excludingarmadillosandsomenon-humanprimates, otherspeciesofarmadillosandBrazilianroadkilledmam-malshadneverbeenresearchedforthispurpose.Depset al. (2002)foundthatbloodsamplesof5outof14animalswere positiveforM. lepraebyPCR.31Inanotherstudy,Depset al. (2007)found11outof37(29.7%)positiveserumsamples usingtheMLFlowtest.32
Search forM. leprae in wild armadillosD. novem-cinctus of different geographical areas resulted negative inFlorida,Colombia,Paraguay,33-35andintheSoutheast areaoftheUnitedStates,correspondingtoAlabama,Ar-Figure 1: Specific repetitive element of M. leprae (RLEP),
demonstrated by visualization of a 372bp product. 1 – M.
smegmatis MC2 155; 2 – M. avium; 3 – Hamster M. auratus
inoculated with M. avium (liver DNA); 4, 5, and 6 – D. novemcinctus inoculated with M. leprae (liver); 7 – Negative control of reaction; 8 – Low Mass DNA ladder.
Pedrini,Rosa,Medri et al.
100pb
kansas,Florida,GeorgiaandMississipi,besidesother800 armadillosexaminedinFlorida.33,36
InthebiodiversityofLatinAmericafauna,oftheactual 20 known armadillo’s species, 17 are only found in Brazil and have been little studied. It has been shown that these animalscanbenaturalcarriersofimportantpathogenssuch asTrypanosoma cruzi,37,38 Histoplasma capsulatum,39
Leish-mania naiffi,40 Toxoplasma gondii,41 andParacoccidioides
brasiliensis.42-44
The nine-banded armadillos, familyDasypodidae, or-derCingulata, SuperorderXenarthra,13were introduced in NorthAmericaveryrecently,aroundtheyear1880.Thehu- mancivilization,thatprobablyappearedinAfrica400,000-200,000 years ago, arrived in America much later, being SouthAmericathelastpartofthecontinenttobecolonized bymanabout14,000yearsago.44,45
The discovery of armadillos from Louisiana naturally infectedwithM.leprae,SNPtype3strain,originatingfrom EuropeandNorthofAfrica,isanindicativethatthoseani-malswerecontaminatedbyhumansources.19TheM. leprae bacilluswasprobablycarriedtotheNewWorldabout500 years ago, after the arrival of settlers and through the Af-ricanslave’straffic.Asseveralspeciesofarmadillosalready lived in Latin America from the Paleocene era, when lep-rosy did not exist in humans, it is possible to infer that if thereareD. novemcinctusnaturallyinfectedincertainareas oftheUnitedStates;theseanimalsmusthavebeeninfected bymanandnotthecontrary.Prabhakaran,in1998,already questioned if armadillos were, besides man, reservoirs of
M. leprae,oncethediseasewasbroughtfromtheOldtothe New World where native inhabitants and armadillos lived withoutleprosy.46
Experimentally,notalltheanimalsaresusceptibletothe infection byM. leprae, because some do not develop the disease (20-30%), even when inoculated with high bacil-laryloads.However,itisadmittedthatthearmadillosmay show an immune status similar to that of some patients withtuberculosis,inwhichthediseaseisauto-limited.12,20,21 Inthatcase,however,armadilloswouldremainforsome timewithhighIgMantibodytitlesagainstPGL-1antigen. ThestudyofDepset al.(2007),usingML-flow,mayreflect thissituation.32
Theobjectiveofthepresentstudywastoidentifyindig- enousleprosyinwildanimals,lookingforabetterunder- standingoftransmission.Somequestionsareyettobean-swered:arearmadillosandotheranimalsofSouthAmerica indeednaturallyinfectedbytheM. leprae?Wouldtheybe involvedintheepidemicchainofthedisease?Werearmadil-losinfectedbyhumanbacillifromTexasandLouisianaor didman,onceagain,interfereintheenvironmentthuscon-taminatingarmadillosthatwerelivinginthoseregions?
OurstudysuggeststhatthearmadillosoftheSãoPaulo and Mato Grosso do Sul states were not contaminated by
man,because,despitethesmallsampling,nonaturallyin-fectedanimalswerefound.Theresultsobtainedshowedthat
M. lepraeisfundamentallyananthropophilicpathogen,and inthatrespectwildanimalsshouldplayasmallornullrole asnaturalreservoirs.
ACKNOWLEDGEMENTS
Financial support: this work was supported by Fundação PaulistaContraaHanseníase.
REFERENCES
1. BrowneSG.In:HastingsRC(ed),Leprosy.ChurchillLiving-stone,Edinburgh,394-5,1994.
2. Noordeen SK. The epidemiology. In: Hastings, Robert C. (Ed.)Leprosy.2.ed.Edinburgh:ChurchillLivingstone:29-34.1994.
3. WilliamsDL,GillisTP,FialloPet al .DetectionofMycobac-teriumlepraeandthepotentialformonitoringantileprosy drugtherapydirectlyfromskinbiopsiesbyPCR.Molecand CellProbes1992;6:401-10.
4. TelentiA,MarchesiF,BalzM,BallyF,BöttgerEC,BodmerT. Rapid Identification ofMycobacteria to the species level by PolymeraseChainReactionandrestrictionenzymeanalysis.J ClinMicrobiol1993;31:175-8.
5. Rastogi N, Goh KS, Berchel M. Species-specific identifica-tion of Mycobacterium leprae by PCR-Restricidentifica-tion Fragment LengthPolymorphismAnalysisofthehsp 65gene.JClinMi-crobiol1999;37:2016-9.
6. WoodsSA,ColeST.Arapidmethodforthedetectionofpo-tentially viable Mycobacterium leprae in human biopsies: anovelapplicationofPCR.FEMSMicrobiolLett1989;65: 305-10.
7. SantosAR,DeMirandaAB,SanoEN,SuffysPN,Degrave WM.UseofPCR-mediatedamplificationofMycobacte-rium leprae DNA in different types of clinical samples for the diagnosis of leprosy. J Med Microbiol 1993; 39: 298-304.
8. MeyersWM,GormusBJ,WalshGP,BaskimGB,HubbardGB. Naturally-acquiredandexperimentalleprosyinnonhumanpri-mates.AmJTropMedandHyg1991;44:24-7.
9. Rojas-EspinosaO,LovikM.MycobacteriumlepraeandMyco-bacterium lepramurium infections in domestic and wild ani-mals.RevSciTechOffIntEpiz2001;20:219-51.
10. Storrs EE. The nine-banded armadillo: a model for leprosy andotherbiomedicalresearch.IntJLeprOtherMycobactDis 1971;39:703-714.
11. KirchheimerWF,StorrsEE.Atttemptstoestablishthearma-dillo(Dasypusnovemcinctuslinn.)asamodelforthestudy of leprosy. Report of lepromatous leprosy in an experimen-tallyinfectedarmadillo.IntJLeprOtherMycobactDis1971; 39:692.
12. RosaPS,BeloneAFF,SilvaEA.Mitsudareactioninarmadillos Dasypus novemcinctus using human and armadillo derived antigens.HansenInt2005a;30:174-9.
53 Braz J Infect Dis 2010;14(1):47-53
14. Laguna AF. El cachicamo sabanero: aspectos de su biologia y ecologia.FondoEditActaCientVenezol15-7;1984.
15. NowakRM.Walker’sMammalsoftheWorld.6thEd.JohnsHop-kinsUniversityPress,Baltimore,515-535.1999.
16. WetzelRM.Theidentificationanddistributionofrecentxenarthra
(Edentata)andtaxonomyanddistribuitonofarmadillo, Dasypodi-dae .In:Montgomery,GG.(Ed),TheEvolutionandEcologyofAr- madillos,SlothsandVermilingua.SmithsonianInstitution,Wash-ington,279-92.1985.
17. MeyersWM,WalshGP,BrownHL,ReesRJW,ConvitJ.Natu-rally acquired leprosy-like disease in the nine-banded arma-dillo (Dasypus novemcinctus): Reactions in leprosy patients to lepromins prepared from naturally infected armadillos. J ReticuloendothelSoc1977;22:369.
18. TrumanRW,JobCK,HastingsRC.Antibodiestothephenolic glycolipid-1antigenforepidemiologicinvestigationsofenzo-oticleprosyinarmadillo(Dasypusnovemcinctus).LeprRev 1990;61:19-24.
19. MonotM,HonoréN,GarnierTet al .OntheOriginofLep-rosy.Science2005;308:1040-1042.
20. RosaPS,BeloneAFF,SilvaEA,PedriniSCB,PinkeCAE.Inocu-laçãoexperimentaldetatuscomoM. leprae :resultadospre-liminares.10ọCongressoBrasileirodeHansenologia,Recife, Brasil.2005b;19-22.
21. RosaPS,SilvaEA,PedriniSCB,PinkeCAE.TheEffectofIron SupplementationintheDietofDasypusnovemcinctusArma-dillosinCaptivity.BrazJBiol2009;69:1(inpress).
22. Brasil, Ministério da Saúde – FNS/CENEPI/CNPS/CRPHF. 1994.Manualdebacteriologiadatuberculose.20ạed.
23. LealNC,AlmeidaAMP.Diagnosisofplagueandidentification ofvirulencemarkersinYersinia pestisbymultiplex-PCR.Rev InstMedTropdeSãoPaulo1999;41:339-42.
24. Mahbubani MH, Bej AK. Applications of polimerase chain reactionmethodologyinclinicaldiagnostics.In:Griffin,L.C., Griffin, A.M. (Eds). PCR Technology: Current innovations. BocaRaton:307-326.1994.
25. GrosskinskyCM,JacobsJrWR,Clark-CurtissJE,BloomBR. Genetic Relationships among Mycobacterium leprae, My-cobacterium tuberculosis, and Candidate leprosy Vaccine StrainsDeterminedbyDNAHybridization:Identificationof anM. leprae-SpecificRepetitiveSequence.InfectImmun1989; 57:1535-41.
26. Eisenberg Jf, Redford K. eds. Mammals of the Neotrop-ics.TheUniversityofChicagoPress,ChicagoandLondon, 1999.
27. Medri,IM.EcologiaeHistóriaNaturaldoTatu-peba, Eu-phractus sexcinctus (Linnaeus,1758),noPantanaldaNhe- colândia,MatoGrossodoSul.(Thesis)UniversityofBra-sília,2008.
28. ColeS.T.,EigimeierK.,ParkhiiJ.et al.Massivegenedecayin theleprosybacillus.Nature2001;409:1007-11.
29. StinearT,JohnsonDR.FromMarinumtoUlcerans:a my-cobacteria lhumanpathogenemerges.Microbe2007;2:187-94.
30. StinearTP,SeemannT,PidotSet al.Reductiveevolutionandniche adaptationinferredfromthegenomeofMycobacteriumulcerans, thecausativeagentofBurulliulcer.GenRes2007;17:192-200.
31. Deps PD, Santos AR, Yamashita-Tomimori J. Detection of Mycobacterium leprae DNA by PCR in blood sample from nine-banded armadillo: Preliminary results. Int J Lepr other mycobactDis2002:70:34-5.
32. DepsPD,AntunesJMAP,Tomimori-YamashitaJ.Detectionof Mycobacteriumlepraeinfectioninwildnine-bandedarmadil-los(Dasypusnovemcinctus)usingtherapidMLFlowtest.Rev SocBrasMedTrop2007;40:86-7.
33. KirchheimerWF,SanchesRM.ExaminationofNorthAmeri- canarmadillosformycobacteriosis.LeprIndia1978;50:156-60.
34. Kirchheimer WF. Examination of north american armadil-los for mycobacteriosis – a further report. Lepr India 1979; 51:60-6.
35. Howerth EW, Stallknecht DE, Davidson WR, Wentworth EJ. Survey for leprosy in nine-banded armadillos (Dasypus novemcinctus)fromtheSoutheasternUnitedStates.JWildlife Dis1990;26:112-5.
36. Walsh GP, Storrs EE, Meyers WM, Binford CH. Naturally acquired leprosy-like disease in the nine-banded armadillo (Dasypusnovemcinctus):recentepizootiologicfindings.JRe-ticuloendothelSoc1977;22:363-7.
37. BarretoMP.ReservatóriosevetoresdeTrypanosoma cruzino Brasil.ArqHigSaúdePub.1963;28:43-66.
38. LainsonR,ShawJJ,FrahaH,MilesMA,DraperCC.Chagas’s diseaseintheAmazonbasin:I.Trypanossoma cruziinfections insilvaticmammals,triatominegugsandmaninstateofPará, northBrazil.TransRoyalTropMedHyg1979;73:193-204. 39.
AriasJR,NaiffRD,NaiffMF,MokWY,AlmeidaMMR.Isola-tionofHistoplasma capsulatumfromanarmadillo(Dasypus novemcinctus)intheeasternAmazonofBrazil.TransRoyal SocTropMedHyg1982;76:705-6.
40. NaiffRD,FreitasRA,NaiffMFet al .Epidemiologicalandno-sologicalaspectsofLeishmanianaiffi.MemInstOswaldoCruz 1991;86:317-21.
41. Sogorb F, Jamira LF, Guimarães EC. Toxoplasmose em ani-maisdeSãoPaulo,Brasil.RevInstMedTropSãoPaulo1977; 19:91-4.
42. NaiffRD,FerreiraLCP,BarreteTV,NaiffMF,AriasJR.Para-coccidioidomicose enzoótica em tatus (Dasypus novemcinc-tus) no estado do Pará. Rev Inst Med Trop São Paulo 1986; 53:19-27.
43. BagagliE,SanoA,CoelhoKILet al.Isolationof Paracoccid-ioides brasiliensis from armadillos (Dasypus novemcinctus) capturedinanendemicareaofparacoccidioidomycosis.Am JTropMedHyg1998;58:505-12.
44. BagagliE,BoscoSMG,TheodoroRC,FrancoM.Phylogenetic andevolutionaryaspectsofParacoccidioides brasiliensisreveal alongcoexistencewithanimalhoststhatexplainseveralbio-logicalfeaturesofthepathogen.InfectGenetandEvol2006;6: 344-51.
45. Leakey RE. Human Origins. Lodestar Books, New York: 87. 1982.
46. PrabhakaranK.WhyMycobacteriumlepraeinvadestheeyes? IndianJLepr1998;70:413.