Hatch et al. Epigenetics & Chromatin (2017) 10:9
DOI 10.1186/s13072-017-0116-6
METHODOLOGY
Assessing histone demethylase
inhibitors in cells: lessons learned
Stephanie B. Hatch
1,2†, Clarence Yapp
1,2†, Raquel C. Montenegro
1,2,3, Pavel Savitsky
1, Vicki Gamble
1,2,
Anthony Tumber
1,2, Gian Filippo Ruda
1,2, Vassilios Bavetsias
4, Oleg Fedorov
1,2, Butrus Atrash
4,
Florence Raynaud
4, Rachel Lanigan
4, LeAnne Carmichael
4, Kathy Tomlin
4, Rosemary Burke
4,
Susan M. Westaway
5, Jack A. Brown
5, Rab K. Prinjha
5, Elisabeth D. Martinez
6, Udo Oppermann
1,7,
Christopher J. Schofield
8, Chas Bountra
1,2, Akane Kawamura
8,9, Julian Blagg
4, Paul E. Brennan
1,2,
Olivia Rossanese
4and Susanne Müller
1,2,10*Abstract
Background: Histone lysine demethylases (KDMs) are of interest as drug targets due to their regulatory roles in chro-matin organization and their tight associations with diseases including cancer and mental disorders. The first KDM inhibitors for KDM1 have entered clinical trials, and efforts are ongoing to develop potent, selective and cell-active ‘probe’ molecules for this target class. Robust cellular assays to assess the specific engagement of KDM inhibitors in cells as well as their cellular selectivity are a prerequisite for the development of high-quality inhibitors. Here we describe the use of a high-content cellular immunofluorescence assay as a method for demonstrating target engage-ment in cells.
Results: A panel of assays for the Jumonji C subfamily of KDMs was developed to encompass all major branches of the JmjC phylogenetic tree. These assays compare compound activity against wild-type KDM proteins to a catalyti-cally inactive version of the KDM, in which residues involved in the active-site iron coordination are mutated to inacti-vate the enzyme activity. These mutants are critical for assessing the specific effect of KDM inhibitors and for revealing indirect effects on histone methylation status. The reported assays make use of ectopically expressed demethylases, and we demonstrate their use to profile several recently identified classes of KDM inhibitors and their structurally matched inactive controls. The generated data correlate well with assay results assessing endogenous KDM inhibition and confirm the selectivity observed in biochemical assays with isolated enzymes. We find that both cellular perme-ability and competition with 2-oxoglutarate affect the translation of biochemical activity to cellular inhibition.
Conclusions: High-content-based immunofluorescence assays have been established for eight KDM members of the 2-oxoglutarate-dependent oxygenases covering all major branches of the JmjC-KDM phylogenetic tree. The usage of both full-length, wild-type and catalytically inactive mutant ectopically expressed protein, as well as struc-ture-matched inactive control compounds, allowed for detection of nonspecific effects causing changes in histone methylation as a result of compound toxicity. The developed assays offer a histone lysine demethylase family-wide tool for assessing KDM inhibitors for cell activity and on-target efficacy. In addition, the presented data may inform further studies to assess the cell-based activity of histone lysine methylation inhibitors.
© The Author(s) 2017. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/ publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
Open Access
Epigenetics & Chromatin
*Correspondence: susanne.mueller-knapp@bmls.de
†Stephanie B. Hatch and Clarence Yapp have contributed equally to this work
10 Buchmann Institute for Molecular Life Science, Goethe University Frankfurt, Riedberg Campus, Max-von-Laue-Straße 15, 60438 Frankfurt am Main, Germany
Page 2 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
Background
Histones are highly conserved proteins that play impor-tant roles, not only in compacting DNA in the cell, but which also have dynamic functions in many physiological and molecular processes. Covalent modiications of the histone tail, in particular on lysine residues, are an essen-tial part of epigenetic mechanisms to regulate processes like DNA repair and gene transcription [1, 2]. he com-plex code of histone tail modiications is tightly regulated and includes a variety of modiications, most prominently acetylation and methylation of lysine and arginine resi-dues. Enzymes that modify or bind to these residues have been implicated in a variety of diseases including cancer, inlammatory diseases, heart disease and neurodegen-erative diseases [3–5]. Methylation of histone lysines was once thought by many to be irreversible, but two classes of histone lysine demethylases (KDMs) have now been identiied that catalyse the removal of these methylation marks. Based on their enzymatic mechanism, the lavin-dependent demethylases of the KDM1 subfamily (lysine-speciic demethylases or LSDs) are distinguished from the Jumonji C (JmjC) family of KDMs, comprising the KDM2 to KDM7 subfamilies, which belong to the super-family of Fe(II)- and 2-oxoglutarate (2-OG)-dependent oxygenases. Demethylation of diferent marks has been described thereby inluencing the ratios of mono-, di- and tri-methylation on H3K4, H3K9, H3K27, H3K36 and H4K20, resulting in diferential efects on gene transcrip-tion. For example, H3K4me2/3 and H3K36me3 are pref-erentially associated with transcriptionally active genes, while H3K9me2/3, H3K27me2/3 and H4K20me3 corre-late with transcriptional repression [6–8].
here is growing interest in exploring the biology and disease relevance of the KDMs with the help of well-char-acterized ‘probe molecules’ that show potent activity in biochemical and cellular assays as well as selectivity over other KDMs [9]. Several inhibitors for the KDM1 fam-ily have been described, with the most advanced com-pounds entering clinical trials [6]. A number of inhibitors have also recently been discovered for the JmjC family of demethylases, e.g. [6, 10–17]. In order to draw conclu-sions regarding the biological activity of new inhibitors, it must irst be conirmed that their observed cellular efect is due to the inhibition of the respective enzyme rather than uncharacterized of-target activity or nonspeciic efects. We have developed and evaluated an immunolu-orescence-based high-content imaging assay to assess the
on-target efect of demethylase inhibitors and describe here the lessons learned.
Results
Several robust in vitro biochemical assays have been developed to assess the potency of potential KDM inhibi-tors; however, most of them do not use the full-length protein, but rather employ truncated constructs encom-passing mainly the catalytic domain. It has recently been shown that, in addition to the catalytic domain, reader domains, such as plant homeodomains (PHDs), are also important in overall KDM enzymatic activity [18, 19]. In order to assess the on-target efect of KDM inhibitors of the JmjC family in cells, we have developed a panel of high-content immunoluorescence-based assays cover-ing the major branches of the phylogenetic tree of the JmjC family of 2-OG-dependent oxygenases that rely on the full-length protein (Fig. 1) [20]. he assays utilize an overexpression system after transiently transfecting the full-length FLAG-tagged demethylase (WT) into cells and assaying the cells overexpressing the demethylase for the respective substrate mark in a high-content immuno-luorescence (IF) assay. Overexpression was found to be optimal after 20–24 h for the tested demethylases after which cell death was observed, even in the absence of inhibitor (Yapp et al., unpublished results). As control, catalytically inactive KDMs (MUT) were used, in which the speciic residues involved in iron coordination were substituted for alanine residues (or tyrosine for KDM3A) (Additional ile 1: Figure S1; Additional ile 2: Table S1). In the ideal case, the dose–response curve of the ectopi-cally produced WT KDM should meet the curve of the MUT KDM when the enzyme is fully inhibited by the inhibitor.
A global decrease in methylation was observed for HeLa cervical carcinoma cells overexpressing the WT demethylase as determined by reduction in the levels of methyl-lysine antibody staining (e.g. KDM5B overex-pression correlating with H3K4me3 nuclear staining in Fig. 2a iv–vi), relative to cells overexpressing the corre-sponding catalytically inactive MUT demethylase or non-transfected cells (Fig. 2a vii–ix).
Many reported inhibitors for the JmjC-KDMs display poor cellular activity [6, 11, 13, 14], and prodrug strategies have been utilized in an attempt to enhance cellular permeabil-ity and activpermeabil-ity [6, 21–23]. We synthesized and tested vari-ous inhibitors from our KDM inhibitor programme as well
Page 3 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
as inhibitors described in the literature (Additional ile 3: Figure S2). Recently, potent inhibitors of the KDM5 family with good cellular activity have been described by Epithera-peutics (Table 1; Additional ile 3: Figure S2; Additional ile 4: Table S2) [24–26]. We tested two of the reported inhibitors: KDOAM-20 and KDOAM-21 (also known as KDM5-C49 and KDM5-C70, respectively) [25], in our cellular high-content assays quantifying changes in the global methylation levels 24 h after addition of the compound. he pyridine car-boxylic acid KDOAM-20 is a highly potent KDM5 inhibitor
in vitro, whereas the more cell-active ethyl ester derivative KDOAM-21 most likely acts as prodrug, which is hydrolysed in cells to provide KDOAM-20, an approach utilized also in other carboxylic acid-bearing KDM inhibitor scafolds and 2-OG oxygenase inhibitors (Table 1) [21, 23, 27]. In parallel, we also tested a related control compound, KDOAM-32, in which the core scafold has been modiied to abrogate KDM-binding (Table 1; Additional ile 3: Figure S2).
Page 5 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
the highest compound concentration tested, the KDM5 enzymes were completely inhibited and H3K4me3 lev-els reached that of the cells transfected with the MUT KDM5 enzyme (Fig. 2a (i–iii), b). In line with the in vitro biochemical potency of the parent carboxylic acid (KDOAM-20), KDOAM-21 showed potent inhibition of H3K4me3 demethylation caused by overexpressed KDM5B with an EC50 < 1 µM. Surprisingly, KDOAM-20 also showed activity against KDM5B in cells with an EC50 of 20 µM despite having low cell permeability in Caco-2 assays (Additional ile 4: Table S2). he other KDM5 fam-ily members tested (KDM5A, KDM5C and KDM5D) were also potently inhibited by KDOAM-21 with EC50 values in the 3–5 μM range in line with the measured in vitro potency of the parent carboxylic acid KDOAM-20 (Fig. 2b; Table 1), but showed little inhibition upon treatment with KDOAM-20 consistent with poor cell penetrance of the parent carboxylic acid (Additional ile 4: Table S2). KDOAM-20/21 also showed inhibition of the KDM4 family, which was assessed by increase of the H3K9me3 mark. KDM4B was inhibited most potently with an EC50 value of 10 μM for KDOAM-21 (Fig. 2c; Table 1). For the KDM4 subfamily members tested, we also observed inhibition of the overexpressed enzymes by the compounds within the same potency rank order as observed in enzyme kinetic assays (Fig. 2c; Table 1; Addi-tional ile 4: Table S2). None of the inhibitors showed activity on members of the other demethylase families tested (KDM3A activity on H3K9me2, KDM6B activity on H3K27me3) (Fig. 2c; Table 1) consistent with the weak in vitro enzymatic potency of these compounds versus the respective demethylase assays.
Interestingly, at higher KDOAM-20/21 inhibitor con-centrations, a dose-dependent efect on the H3K4me3 mark was observed in cells overexpressing MUT KDM5. his efect was not seen in cells expressing WT or MUT KDM5 treated with the control compound KDOAM-32, possibly indicating that the observed efects were either due to inhibition of the endogenous enzyme(s) or due to a nonspeciic efect of KDOAM-21 and KDOAM-20
(Fig. 2b). We observed no toxicity as measured by cell count using HeLa cells at any of the measured concen-trations, indicated by a constant number of cells assessed in the high-content screen (Additional ile 5: Figure S3). In order to further study inhibition of endogenous H3K4me3 levels, nontransfected HeLa cells were treated with KDOAM-20, KDOAM-21 and the inactive control KDOAM-32 at the same concentrations used in assays where the protein was ectopically expressed. H3K4me3 was monitored after 72 h by IF staining. Observed EC50 (sub-μM potency for 21, ~50 μM for KDOAM-20 and no inhibition observed for KDOAM-32) were in good agreement with EC50 values measured using the overexpressed enzyme (Fig. 2d; Table 1). Similar results were obtained after treatment of nontransfected HeLa cells for 48 h, whereas at 24 h the observed EC50 values were slightly lower (Hatch et al. unpublished results). We therefore have established a robust panel of cell assays to test inhibitors for the Jumonji family of KDMs.
Treatment of cells with inhibitors of the recently disclosed cell-penetrant cyanopyrazole class such as CPI-455 [28, 29] (Table 1; Additional ile 3: Figure S2; Additional ile 4: Table S2) resulted in a dose-dependent increase of the H3K4me3 mark (Fig. 3a). An increase in methylation in both WT and MUT KDM5-expressing cells was observed, which was even more pronounced than noted with KDOAM-21, as described above. No reduction in cell numbers was observed after treatment with this inhibitor, indicating that the compound is not toxic at the exposures examined (Fig. 3a). We hypoth-esized that the observed CPI-455-dependent increase of the H3K4me3 mark in cells overexpressing the mutant KDM5B protein was due to compound-mediated inhibi-tion of the endogenous KDM5 enzyme(s) as observed for the KDOAM series.
In the weakly acidic pyridopyrimidinone series [13] (Table 1; Additional ile 3: Figure S2), we observed dose-response cellular activity in the KDM5B IF assay. Treat-ment with KDIPP51 (CCT364883) resulted in a marked spike indicating increased methylation levels in both (See figure on previous page.)
P
age 6 of 17
Ha
tch
et al
. Epigenetic
s & Chr
omatin (2017) 10:9
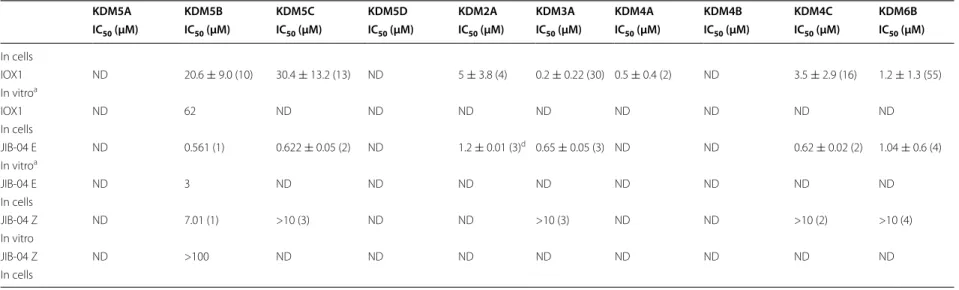
Table 1 In vitro and cellular IC50 of selected KDM inhibitors tested
KDM5A KDM5B KDM5C KDM5D KDM2A KDM3A KDM4A KDM4B KDM4C KDM6B
IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM)
KDOAM-20 0.010 ± 0.0030 (6)
0.007 ± 0.0030 (12)
0.019 ± 0.0050 (10)
0.019 ± 0.0040 (6)
2.2 ± 0.37 (2) 1.2 ± 0.91 (4) 0.67 ± 0.23 (2) 0.099 (1) 0.19 ± 0.081 (6) 1.4 ± 0.45 (2) In vitro
KDOAM-20 ND 20 ND ND ND ND ND ND ND ND
In cells
KDOAM-21 0.068 ± 0.010 (2) 0.094 ± 0.065 (6) 0.17 ± 0.073 (6) ND 19 ± 10 (2) 2.6 ± 1.3 (4) 11 ± 4.5 (2) 2.5 (1) 2.8 ± 1.3 (4) 11 ± 4.6 (2) In vitro
KDOAM-21 4.8 0.7 3.1 4 NA NA 70 10* 17.9 NA In cells
KDOAM-32 ND >10 (2) ND ND ND ND ND ND ND ND In vitro
KDOAM-32 >100 >100 >100 >100 >100 >100 >100 >100 >100 >100 In cells
KDIPP15 0.040 ± 0.016 (2) 0.063 ± 0.028 (2) 0.26 ± 0.039 (2) 0.23 ± 0.039 (2) >100 (2) >100 (2) 3.9 ± 1.4 (3) 1.4 ± 0.47 (3) 5.1 ± 0.062 (4) >100 (2) In vitro
KDIPP15 ND 20.7 ND ND ND ND ND ND ND ND In cells
KDIPP51 (CCT364883)
0.015 ± 0.0041 (4)
0.014 ± 0.0046 (4)
0.024 ± 0.0044 (4)
0.024 ± 0.0038 (4)
8.6 ± 5.9 (2) 6.0 ± 3.9 (4) 0.126 ± 0.013 (2)
0.050 ± 0.0038 (2)
0.10 ± 0.031 (4) 44 ± 17 (3) In vitro
KDIPP51 (CCT364883)
ND 39.9# ND ND ND ND ND ND ND ND
In cells
CPI-455 0.16 ± 0.068 (2) 0.26 ± 0.064 (2) 7.8 ± 1.6 (2) ND 23 ± 3.5 (2) >100 (2) 31 (1) 44 (1) 11 ± 2.3 (2) >100 (2) In vitro
CPI-455 ND 60 ND ND ND ND ND ND ND ND
iN cells
CCT365599 ND 0.023(2) 0.065(2) ND 12.9c 5.3c 0.102 ± 0.058 0.031 ± 0.012 ND 15% at 100 µMc
In vitroa
CCT365599 ND >50 ND ND ND ND 10 ND ND ND In cells
CCT366293 ND >10c >10c ND ND ND >10 (2) >10 (2) ND ND
In vitroa
P
age 7 of 17
Ha
tch
et al
. Epigenetic
s & Chr
omatin (2017) 10:9
In vitro
IC50 values ± SD (number of replicates) determined by the AlphaScreen assay
a In vitro biochemical proiling was previously reported [11]
b Biochemical assay protocols as previously reported [11]. CCT366293 displays assay interference above 10 μM c Results are from a single determination
d Results are determined by RapidFire under low iron condition (0.2 mM)
In cells
EC50 values were determined by IF assay
Results are mean values of at least 3 independent experiments unless indicated otherwise
* Results from 2 independent experiments
# Highest concentration was excluded from calculation
Table 1 continued
KDM5A KDM5B KDM5C KDM5D KDM2A KDM3A KDM4A KDM4B KDM4C KDM6B
IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM) IC50 (µM)
In cells
IOX1 ND 20.6 ± 9.0 (10) 30.4 ± 13.2 (13) ND 5 ± 3.8 (4) 0.2 ± 0.22 (30) 0.5 ± 0.4 (2) ND 3.5 ± 2.9 (16) 1.2 ± 1.3 (55) In vitroa
IOX1 ND 62 ND ND ND ND ND ND ND ND
In cells
JIB-04 E ND 0.561 (1) 0.622 ± 0.05 (2) ND 1.2 ± 0.01 (3)d 0.65 ± 0.05 (3) ND ND 0.62 ± 0.02 (2) 1.04 ± 0.6 (4)
In vitroa
JIB-04 E ND 3 ND ND ND ND ND ND ND ND
In cells
JIB-04 Z ND 7.01 (1) >10 (3) ND ND >10 (3) ND ND >10 (2) >10 (4) In vitro
Page 8 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
WT and MUT KDM5B-expressing cells at the highest inhibitor concentration (Fig. 3b). his spike in methyla-tion levels correlates with a steep drop in cell numbers (Fig. 3b). he potent inhibitor KDOAM-21 demonstrated that KDM5 inhibition per se is not toxic to HeLa cells, so the observed cell death is unlikely to be solely due to inhi-bition of KDM5 demethylation activity. Cell permeabil-ity for both compounds is good and better for KDIPP15 compared to KDIPP51 (Additional ile 4: Table S2). We therefore propose that the observed spike in H3K4 meth-ylation is most likely due to nonspeciic of-target efects of KDIPP51 and/or is due to the simultaneous inhibition of several KDMs. In cells overexpressing KDM5B and treated with the promiscuous KDM inhibitors IOX1 [30] and JIB-04 [16], we only observed cell toxicity at high concentrations (Fig. 4a; Additional ile 6: Figure S4). Both of these inhibitors elicited a dose-dependent increase of H3K4me3 in cells transfected with WT KDM5B. In addi-tion, treatment with the broad-spectrum KDM inhibi-tor JIB-04 at concentrations above ~20 µM resulted in an increase of H3K4me3 in cells expressing either the WT or the MUT enzyme similar to what was observed with KDIPP51 in cells overexpressing KDM5B (Table 1;
Fig. 4a) and accompanied by a reduction in cell number (Additional ile 6: Figure S4).
he KDM6A/B inhibitor GSK-J4 [21] is reported to also inhibit KDM5 at higher concentrations in vitro [31]. We tested GSK-J4 together with its inactive con-trol compound GSK-J5 and also observed an increase of the H3K4me3 mark upon treatment with the active KDM6A/B inhibitor, but not the inactive control. hese two compounds showed only minor toxicity at the high-est concentrations used, which is well above the efective doses used for KDM6A/B inhibition (~1 µM) (Fig. 4b; Additional ile 6: Figure S4). Assessment of inhibitors in the high-content assay therefore needs careful considera-tion of all parameters, WT and MUT data. In addiconsidera-tion, steep-slope increases of histone methylation mark are often accompanied by a reduction in cell number point-ing to cell toxicity of the compound. he results imply that simultaneous inhibition of several KDMs does not seem to be tolerated at higher compound concentrations by the tested cells.
Page 9 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
but that are known not to be associated with KDM inhib-itor activity (Additional ile 7: Table S3). Accordingly, we treated HeLa cells with the DNA intercalator doxorubicin and the microtubule stabilizer paclitaxel and monitored the efect of these cytotoxic agents on H3K4me3 levels (Fig. 5a). Both these compounds induced cell death with the expected EC50 values as estimated from the number of cells remaining per ield after ixation (Fig. 5b). Sur-prisingly, paclitaxel also led to a dose-dependent increase of H3K4me3, whereas doxorubicin had no efect on this mark indicating that cell death can be associated with an increase in H3K4me3 without direct inhibition of demethylases (Fig. 5c). We then tested the inluence of these two compounds on the other histone marks used to assess KDM activity (Fig. 5d). Both inhibitors had a profound efect on the repressive H3K27me3 mark. Con-versely, H3K9me2 levels were not afected by cell death induced by paclitaxel, but showed some increase upon treatment with doxorubicin, although the efects were more variable. Globally, H3K36me2 levels were only mar-ginally afected by cell death induced by either of the two inhibitors but individual cells could show a complete lack of this mark upon treatment (Additional ile 8: Figure S5).
Similar efects on histone methylation have been noted using other compounds causing cell death like the broad-spectrum kinase inhibitor staurosporine (unpublished observation). None of the three compounds had any spe-ciic efect on inhibition of KDM4C, KDM5B or KDM6B in in vitro demethylase assays (Additional ile 7: Table S3), indicating that mechanisms inducing cytotoxicity can commonly afect global histone methylation marks.
To assess the mode of cell death caused by these compounds and by the tested KDM inhibitors in more detail, we performed a high-content-based triple stain-ing protocol (Fig. 6). Cells were categorized into healthy cells (Hoechst staining only), apoptotic cells deined as Annexin V positive with or without Yo-Pro 3 uptake, or necrotic cells deined by Yo-Pro 3-positive Annexin V negative staining (Fig. 6a) [32]. After 24 h of treatment with doxorubicin, paclitaxel or the pan-kinase inhibi-tor staurosporine, cell death occurred and was accom-panied by the appearance of a predominant apoptotic staining, in line with their known mechanism of action. At higher concentrations the number of necrotic cells increased as monitored by a Yo-Pro 3-positive stain-ing. In contrast, cells treated with DMSO were deined as ‘healthy’ and showed predominantly a negative stain-ing for both Annexin V and Yo-Pro 3 (Fig. 6b). We then tested the diferent KDM inhibitors to assess their efect on cell viability in more detail. As expected, inhibitors of the KDOAM series (KDOAM-20, KDOAM-21 and KDOAM-32) did not induce cell death at any of the con-centrations measured, above the level of DMSO nor did CPI-455 or KDIPP15 (Fig. 6c, d). However, treatment of HeLa cells with KDIPP51 (at 60 µM) for 24 h resulted in 40% apoptotic and 20% necrotic cells as compared to 20% apoptotic and 2% apoptotic cells upon treatment with KDIPP15 (66 µM), similar to what was observed for DMSO-treated cells, indicating that the observed changes in methylation coincide with cell death accom-panied by increased apoptosis and necrosis (Fig. 6e). In the same assay, the pan-KDM inhibitors IOX1 and JIB-04 and its inactive control displayed little to no cell toxicity, with only a minor efect of JIB-04 on cellular viability at 30 µM. However, both the KDM6B inhibitor GSK-J4 and its inactive control GSK-J5 (Additional ile 9: Figure S6) showed about 60% necrotic cells at 50 µM concentration, which is well above the concentration required to inhibit KDM5/6 activity in cellular assays.
We have recently described the discovery of cell-pen-etrant analogues of the pyridopyrimidinone KDIPP51 as dual KDM4/5-subfamily inhibitors [11]. We have used the assays described herein to measure the cellular activity of these compounds and their matched negative control analogues that we report here for the irst time. Treatment of HeLa cells with CCT365599 resulted in a Fig. 4 Effect of various KDM inhibitors described in the literature on
Page 10 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
concentration-dependent increase of H3K9me3 and H3K4me3 levels, while treatment with the respective negative controls CCT366293 had no efect (Fig. 7a). No cytotoxicity was observed for these compounds at the concentrations used to assess H3K4me3 and H3K9me3 demethylation (Fig. 7b, c), suggesting that in these cases the increase in methylation is due to inhibition of the cor-responding demethylase. However, we notice for these compounds and the other compounds tested that there
Page 11 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
Page 12 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
Page 13 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
shifts in the presence of 2-OG, leading to values in the 3–5 μM range in the presence of physiologically relevant 2-OG concentrations (1000 μM).
Discussion
Almost all currently available JmjC-KDM inhibitors act via metal chelation and compete with the 2-OG co-substrate for binding at the active site. Poor cellular activity has been a problem; however, cell permeability is not the only reason for the poor cellular activities of the 2-OG competitive KDM inhibitors [11]. he results presented here further inform on the cellular activities of KDM inhibitors. hey indicate that the currently used in vitro assays may overestimate the cellular efects due to the use of lower than physiologically relevant 2-OG concentration.
A number of methods have been described to assess cellular activity of KDM inhibitors, each with advantages as well as disadvantages [20]. We have described a panel of cellular assays to assess the direct inhibition of KDMs as well as selectivity of KDM inhibitors in cultured human cells using a high-content-based overexpres-sion system. We assessed several known KDM inhibitors as well as newly described inhibitors and demonstrate the usefulness of structure-matched inactive control compounds to assess efects related to speciic KDM inhibition. In addition, the results obtained with overex-pression of the MUT protein in combination with assess-ment of cell numbers provide a useful guide to enzyme inhibition in cells, helping to identify on-target efects. It is now our observation across multiple compounds that, in our hands, a spike in histone mark methylation is characteristic of cytotoxicity. he results obtained using this system are in good agreement with data obtained assaying endogenous demethylases, but ofer the addi-tional advantage of distinguishing between a direct efect on KDM inhibition and changes to histone methylation upon induction of cell death.
Cell death observed upon treatment with toxic com-pounds (e.g. doxorubicin or paclitaxel) resulted in a dose-dependent increase in histone methylation, in particular of the repressive H3K27me3 mark, but also of other marks such as H3K4me3 and H3K9me2. A recent study reported the induction of global changes in histone lysine and arginine methylation and altered expression of lysine demethylases upon treatment of human cancer cells with HDAC inhibitors pointing to a complex network of histone crosstalk [34]. Increased methylation of histone H3K27 associated with apopto-sis induced by staurosporine has also been described in osteosarcoma cells, but the study did not examine any changes in dimethyl or trimethyl histone H3K4 and H3K9 [35].
Although not addressed in the current study, one can speculate regarding the underlying physiological mecha-nisms of these histone mark changes in apoptosis and general cell death. Little is known about changes in his-tone methylation upon induction of cell death in humans although a global decrease in histone acetylation has been observed during apoptosis [36, 37]. Loss of histone H3 methylation at lysine 4 has, however, been shown to trig-ger apoptosis in Saccharomyces cerevisiae [38]. In mela-noma cells, short-term treatment (4 h) with doxorubicin decreased H3K4me3 methylation but did not have an efect on H3K27me3 [39]. Detailed chromatin immuno-precipitation-based assays and gene expression analyses, as well as time-resolved experiments, will be necessary to investigate in more detail the interplay between cell death and chromatin changes addressing questions of speciic mechanisms—for example, if cells irst activate genes through increased density of H3K4me3 activating marks before increasing repressive marks like H3K27me3 when cell death cannot be avoided.
Conclusions
In summary, we have developed a series of overexpres-sion IF assays to assess the target engagement of KDM inhibitors in cells. he presented assays cover most major branches of the 2-OG JmjC-KDMs. Nonspeciic toxic-ity of tested inhibitors, in particular those resulting in apoptosis, leads to changes in histone tail modiications unrelated to the expected cellular activity of these com-pounds. he presented assay provides a suitable method to distinguish between direct and indirect efects of his-tone methylation.
Methods
Cloning
Full-length cDNA for KDM5A (JARID1A, P29375), KDM5B (JARID1B, Q9UGL1, cDNA), KDM5C (JARID1C, P41229, IMAGE 5492114), KDM4A (O75164) containing A482E substitution (natural variant rs586339), KDM4B (JMJD2B, O94953, gift from ICR), KDM4C (JMJD2C, Q9H3R0, 8143862), KDM3A (JMJD1A, Q9Y4C1, 4823253), KDM2A (FBXL11, Q9Y2K7, 5534384) and KDM6B (JMJD3, O15054, cDNA) were ampliied by PCR from either a MGC clone or commer-cial cDNA source or received as a gift from collaborators and cloned into pDONR-221 vector using Gateway BP reaction producing Gateway entry clones.
Page 14 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
H212A/D214A and KDM6B H1390A/E1392A). Muta-tions were introduced into full-length KDM Gateway entry clones using 15 cycles QuikChange II PCR protocol (Agilent Technologies).
Mammalian expression constructs encoding for N-ter-minal 3*FLAG potencies were constructed using the Gateway LR recombination reaction between pCDNA5-FRT/TO-3FLAG destination vector [40] and the wild-type or mutated KDM Gateway entry clone.
All constructs described are available upon request.
Cell culture/transfection for 24‑h IF assay
HeLa cells (ATCC) were cultured at 37 °C and 5% CO2 in Minimal Essential Media—Eagle (Sigma-Aldrich, UK) supplemented with 10% heat-inactivated foetal calf serum (FCS) (Gibco, UK), and 1% l-glutaMAX (Gibco, UK) in T75 lasks (Sarstedt, UK) and passaged regularly with TrypLE (Life Technologies, UK) before cells reached conluence. On day 0, 7000 cells per well were plated on a 96-well clear bottom CellCarrier plate (PerkinElmer, UK). he next day the cells were transfected with Lipo-fectamine 3000 (hermoFisher Scientiic, UK) following the manufacturer’s instructions. Briely, 0.1 µg of plas-mid DNA (encoding for the respective demethylase) and 0.15 µL of Lipofectamine 3000 were diluted in separate tubes containing 5 µL OptiMEM. 0.15 µL of P3000 was added to the tube with the DNA. he two solutions were then mixed gently and allowed to incubate for 5 min at room temperature. Next, the DNA-lipid complex was added to the cells for 4 h at 37 °C and 5% CO2.
Compound dilutions were made in a 96-well PCR plate (Starlab, UK). he culture medium was supplemented with 0.4% DMSO so that the solvent concentration was maintained throughout. Compounds were serially diluted at a 1:2 or 1:3 dilution ratio. Next, the cell culture plate was removed from the incubator and the medium was gently aspirated and replaced with the fresh medium containing diluted compounds.
After 24 h of compound incubation, cells were stained using the following protocol. First, the cells were washed once with Phosphate Bufered Saline (PBS) (Sigma-Aldrich, UK) and ixed for 20 min at room temperature using formaldehyde diluted in PBS to 4% w/v. Next, cells were rinsed once with PBS and permeabilized for 5 min at room temperature using TritonX-100 diluted in PBS at 0.5% v/v. hen cells were rinsed once with PBS and blocked for 30 min at room temperature using 3% v/v foetal calf serum diluted in PBS. his mixture was then replaced with the appropriate primary antibodies for the histone mark and FLAG-tag diluted in blocking solu-tion and left on overnight at 4 °C. he following day, cells were rinsed three times with PBS and stained with the
secondary antibody for 1 h at room temperature. Before imaging, cells were rinsed once with PBS, stained with DAPI (LifeTechnologies, UK) for 5 min and rinsed again with PBS at room temperature.
Endogenous IF assay
he experimental set-up was similar to the 24-h assay except for the following modiications. Cells were cul-tured in OptiMEM (LifeTechnologies, UK) and supple-mented with 0.5% foetal calf serum (FCS) (Gibco, UK) and 1% l-glutaMAX (Gibco, UK). HeLa cells were seeded at a density of 800 cells per well on day 0. he next day, a compound dilution plate (diluted in OptiMEM with sup-plements) was made and the medium from the day before replaced. he cells were incubated for 72 h at 37 °C and 5% CO2. After ixation, the immunoluorescence proto-col was similar as the 24-h assay with the omission of the FLAG-tag antibodies (Table 2).
Wideield luorescence microscopy
Transfection of cells was assessed on a Zeiss AxioObserver Z1 inverted luorescence microscope itted with an Axio-cam 506 monochrome Axio-camera, Colibri.2 LED system (385, 475, 561, 647 nm), 10× 0.45 N.A. and 20× 0.8 N.A. Plan Apochromat objectives. he system PC ran Windows 7 Ultimate 64-bit and had an Intel Core i5-2500 @3.30 GHz processor with 8 GB RAM and an integrated video card. he microscope was controlled with ZEN Blue.
High‑content analysis
Page 15 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
Hoechst 33342/Yo‑Pro 3/Annexin triple staining and live cell death pattern analysis
Cells exposed to the inhibitors for 24 h were stained with Hoechst 33342 (1 µM), Yo-Pro 3 (1 µM) and Annexin V (0.3 μL per well) for 1 h. Cellular luorescence was meas-ured using the IN Cell 6000 (GE) using the following set-up parameters: bright-ield transmitted light at 50% for 15 ms; Hoechst 33342 was excited by 35-ms exposure(Ex 360–400 nm/Em 410–480 nm), Yo-Pro 3 by 35-ms exposure(Ex 560–580 nm/Em 650–760 nm) and Annexin V (Alexa 488) by 45-ms exposure(Ex 460–490 nm/Em 500–550 nm). All the generated data were analysed using the IN Cell Analyser software, and three categories were distinguished: healthy cells, apoptosis and necrosis—cal-culated as percentage of each class for every concentra-tion used.
Caco‑2 permeability
Papp (apparent permeability) was determined in the Caco-2 human colon carcinoma cell line essentially as described in [11].
AlphaScreen
IC50 values were determined as previously described [11]. he following amendments were made to the protocol for carrying out 2-OG competition assays.
For each compound, a 10 mM stock concentration in 100% DMSO was used. Compounds (100 nL) were dis-pensed with an ECHO®
550 acoustic dispenser (Labcyte Inc™
, Sunnyvale, CA, USA) to generate 10–12 pt dilu-tion curves directly into 384-well Proxiplates (#6008289, PerkinElmer, Waltham, MA, USA) to give inal assay concentrations in the range 0.00015–30 µM in 2%(v/v) DMSO where appropriate. Compounds were pre-incu-bated with their enzymes (2.5 µL, 1.5 nM KDM4A, 0.5 nM KDM4B or for 2 nM KDM5B inal assay concen-tration) for 10 min at room temperature before the addi-tion of their peptide substrates (2.5 µL).
Peptide substrates (60 nM ARTKQTARK(me3)STG-GKAPRKQLA-GGK-biotin for KDM4A and KDM4B or 200 nM ARTK(me3)QTARKSTGGKAPRKQLA-GGK-biotin for KDM5B) were made up in peptide bufer consisting of HEPES (50 mM, pH 7.5), BSA (0.1%), sodium ascorbate (200 µM), ammonium iron(II) sul-phate hexahydrate (2 µM for KDM4A/KDM4B and 10 µM for KDM5B), Tween 20 (0.01%) and 11 2-OG concentrations ranging from 2000 to 0.5 µM. Peptide and bufer components were diluted twofold upon the addition to the compound plate containing enzyme in assay bufer [HEPES (50 mM, pH 7.5), BSA (0.1%) and Tween 20 (0.01%)]. he plate was sealed, then centri-fuged at 1000 rpm for 1 min before being left for 15 min at room temperature. he reaction was stopped with the addition of 2.5 µL EDTA (30 mM EDTA in water for KDM4A and KDM4B or 30 mM EDTA, 1600 mM NaCl, 50 mM HEPES pH 7.5 and 0.01% Tween 20 for KDM5B) before the addition of 2.5 µL AlphaScreen detection reagents and antibody appropriate for the substrate being detected (mouse IgG beads (#6760606M, Perki-nElmer) and anti-dimethyl H3K9 antibody (#Ab1220, Abcam) for KDM4A and KDM4B and Protein A beads (#6760617M, PerkinElmer) and anti-dimethyl H3K4 antibody (#9725S, Cell Signalling Technology) for KDM5B). AlphaScreen beads and antibody mixes were prepared and pre-incubated for 1 h prior to addition to the plate. Plates were left for 1.5 h in the dark before being read on the EnVision®
Multilabel Reader (Perki-nElmer Life Sciences). All assays were run at a inal assay reaction volume of 5 µL and a total assay volume of 10 µL.
Data were normalized to the controls without 2-OG and DMSO wells containing all reagents as the totals. IC50 values at each 2-OG concentration were determined using a nonlinear regression it of the log (inhibitor) ver-sus response with variable slope equation in GraphPad Prism 6.0.
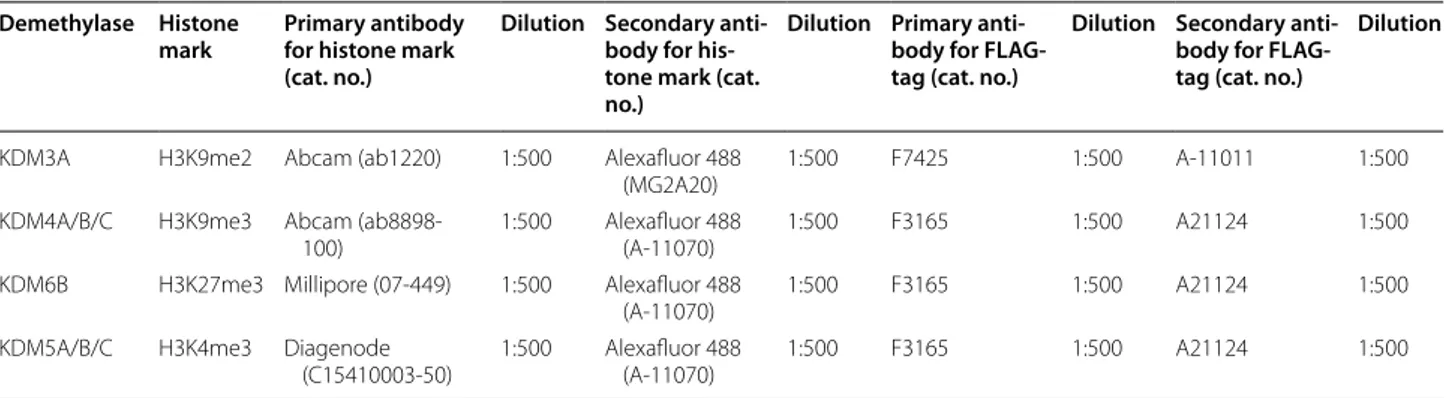
Table 2 Antibody speciications for each demethylase IF assay
Demethylase Histone
mark
Primary antibody for histone mark (cat. no.)
Dilution Secondary anti‑ body for his‑ tone mark (cat. no.)
Dilution Primary anti‑ body for FLAG‑ tag (cat. no.)
Dilution Secondary anti‑ body for FLAG‑ tag (cat. no.)
Dilution
KDM3A H3K9me2 Abcam (ab1220) 1:500 Alexafluor 488 (MG2A20)
1:500 F7425 1:500 A-11011 1:500
KDM4A/B/C H3K9me3 Abcam (ab8898-100)
1:500 Alexafluor 488 (A-11070)
1:500 F3165 1:500 A21124 1:500
KDM6B H3K27me3 Millipore (07-449) 1:500 Alexafluor 488 (A-11070)
1:500 F3165 1:500 A21124 1:500
KDM5A/B/C H3K4me3 Diagenode (C15410003-50)
1:500 Alexafluor 488 (A-11070)
Page 16 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
Chemical synthesis
Additional ile 10 Methods.
Abbreviations
KDM: histone lysine demethylase; JmjC: Jumonji C; 2-OG: 2-oxoglutarate; IF: immunofluorescence; HCA: high-content assay.
Authors’ contributions
SBH and CY set up and performed IF assays and contributed to drafting of the manuscript. RCM set up and performed cytotoxicity assays and contributed to manuscript writing. PS cloned and sequenced all KDM constructs and contributed to manuscript writing. VG performed cytotoxicity assays. AT performed in vitro screening of compounds and evaluation of data. GFR and RL performed chemical synthesis. VB designed chemical inhibitors used in the study and contributed to drafting the manuscript. OF contributed to in vitro assays and evaluation of data. BA designed and synthesized chemical inhibitors. FR evaluated cell permeability data. LAC performed IF assay. KT performed in vitro 2-OG-dependent assay. RB evaluated in vitro assay data and contributed to manuscript writing. SMW designed chemical synthesis and contributed to manuscript writing. JAB, RKP, EDM UO, CJS, AK, JEB, CB and PEB contributed to manuscript writing. OR contributed to study design and par-ticipated in coordination and manuscript writing. SM devised the study and wrote the manuscript. All authors read and approved the final manuscript. Author details
1 Nuffield Department of Clinical Medicine, Structural Genomics Consortium, University of Oxford, Old Road Campus Research Building, Roosevelt Drive, Oxford OX3 7DQ, UK. 2 Nuffield Department of Medicine, Target Discovery Institute, University of Oxford, Roosevelt Drive, Oxford OX3 7FZ, UK. 3 Medical Faculty, Research and Drug Development Center, Federal University of Ceará, Rua Cel. Nunes de Melo n.1000—Rodolfo Teófilo, 60, Fortaleza, CE 430-270, Brazil. 4 Cancer Research UK Cancer Therapeutics Unit, The Institute of Cancer Research, 15 Cotswold Road, London, SM2 5NG, UK. 5 Epigenetics Discov-ery Performance Unit, Medicines Research Centre, GlaxoSmithKline R&D, Stevenage SG1 2NY, UK. 6 Hamon Center for Therapeutic Oncology Research, and Department of Pharmacology, UT Southwestern Medical Center at Dallas, Dallas, TX 75390, USA. 7 Nuffield Department of Orthopedics, Rheumatol-ogy and Musculoskeletal Sciences, Botnar Research Centre, NIHR Oxford Additional iles
Additional ile 1: Figure S1. Ribbon structure of KDM4A based on pdb: 2OQ6. Shown are residues involved in iron coordination with mutated amino acids annotated in black.
Additional ile 2: Table S1. KDMs and mutant KDMs for which assays are reported including histone mark assessed.
Additional ile 3: Figure S2. Chemical structure of the KDM inhibitors used in the study.
Additional ile 4: Table S2. Cell permeability data of compounds tested. Additional ile 5: Figure S3. Effect of KDOAM compounds on HeLa cell number transfected with different KDMs. The number of cells in 20 fields, based on DAPI staining, is shown.
Additional ile 6: Figure S4. Effect of various compounds on HeLa cell number transfected with different KDMs. The number of cells in 20 fields, based on DAPI staining, is shown.
Additional ile 7: Table S3. Effect of Doxorubicin, Paclitaxel and Stauro-sporine on KDM activity measured in in vitro KDM assay.
Additional ile 8: Figure S5. Cell images of HeLa cells treated with 5 µM Doxorubicin or 100 nM Paclitaxel and stained with an anti H3K36me2 antibody.
Additional ile 9: Figure S6. Percentage of healthy, apoptotic and necrotic HeLa cells treated with KDM inhibitors and negative controls for 24 h in a dose-dependent manner.
Additional ile 10. Methods and chemical synthesis.
Biomedical Research Unit, University of Oxford, Oxford OX3 7LD, UK. 8 Chem-istry Research Laboratory, 12 Mansfield Road, Oxford OX1 3TA, UK. 9 Division of Cardiovascular Medicine, Radcliffe Department of Medicine, Wellcome Trust Centre for Human Genetics, Roosevelt Drive, Oxford OX3 7BN, UK. 10 Buch-mann Institute for Molecular Life Science, Goethe University Frankfurt, Ried-berg Campus, Max-von-Laue-Straße 15, 60438 Frankfurt am Main, Germany. Acknowledgements
We thank Stefan Knapp for carefully reading of the manuscript and Adam Donovan for the provision of Caco-2 cell permeability assay data. Competing interests
The authors declare that they have no competing interests. Funding
The research has received support from the Innovative Medicines Initiative Joint Undertaking under Grant Agreement No. 115766, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and EFPIA companies’ in kind con-tribution. Authors from the Cancer Research UK Cancer Therapeutics Unit at The Institute of Cancer Research are supported by Cancer Research UK. Grant C309/A11566.
Received: 13 December 2016 Accepted: 21 February 2017
References
1. Tessarz P, Kouzarides T. Histone core modifications regulating nucleo-some structure and dynamics. Nat Rev Mol Cell Biol. 2014;15(11):703–8. 2. Lawrence M, Daujat S, Schneider R. Lateral thinking: how histone
modifi-cations regulate gene expression. Trends Genet. 2016;32(1):42–56. 3. Sen N. Epigenetic regulation of memory by acetylation and methylation
of chromatin: implications in neurological disorders, aging, and addiction. Neuromol Med. 2015;17(2):97–110.
4. Muller S, Filippakopoulos P, Knapp S. Bromodomains as therapeutic targets. Expert Rev Mol Med. 2011;13:e29.
5. Johansson C, Tumber A, Che K, Cain P, Nowak R, Gileadi C, Oppermann U. The roles of Jumonji-type oxygenases in human disease. Epigenomics. 2014;6(1):89–120.
6. McAllister TE, England KS, Hopkinson RJ, Brennan PE, Kawamura A, Scho-field CJ. Recent progress in histone demethylase inhibitors. J Med Chem. 2016;59(4):1308–29.
7. Dimitrova E, Turberfield AH, Klose RJ. Histone demethylases in chromatin biology and beyond. EMBO Rep. 2015;16(12):1620–39.
8. Zhang T, Cooper S, Brockdorff N. The interplay of histone modifications— writers that read. EMBO Rep. 2015;16(11):1467–81.
9. Muller S, Brown PJ. Epigenetic chemical probes. Clin Pharmacol Ther. 2012;92(6):689–93.
10. Maes T, Carceller E, Salas J, Ortega A, Buesa C. Advances in the develop-ment of histone lysine demethylase inhibitors. Curr Opin Pharmacol. 2015;23:52–60.
11. Bavetsias V, Lanigan RM, Ruda GF, Atrash B, McLaughlin MG, Tumber A, Mok NY, Le Bihan YV, Dempster S, Boxall KJ, et al. 8-Substituted pyrido[3,4-d]pyrimidin-4(3H)-one derivatives as potent, cell permeable, KDM4 (JMJD2) and KDM5 (JARID1) histone lysine demethylase inhibitors. J Med Chem. 2016;59(4):1388–409.
12. Thinnes CC, Tumber A, Yapp C, Scozzafava G, Yeh T, Chan MC, Tran TA, Hsu K, Tarhonskaya H, Walport LJ, et al. Betti reaction enables efficient synthesis of 8-hydroxyquinoline inhibitors of 2-oxoglutarate oxygenases. Chem Commun (Camb). 2015;51(84):15458–61.
13. Westaway SM, Preston AG, Barker MD, Brown F, Brown JA, Camp-bell M, Chung CW, Drewes G, Eagle R, Garton N, et al. Cell penetrant inhibitors of the KDM4 and KDM5 families of histone lysine demethy-lases. 2. Pyrido[3,4-d]pyrimidin-4(3H)-one derivatives. J Med Chem. 2016;59(4):1370–87.
Page 17 of 17 Hatch et al. Epigenetics & Chromatin (2017) 10:9
• We accept pre-submission inquiries
• Our selector tool helps you to find the most relevant journal • We provide round the clock customer support
• Convenient online submission • Thorough peer review
• Inclusion in PubMed and all major indexing services • Maximum visibility for your research
Submit your manuscript at www.biomedcentral.com/submit
Submit your next manuscript to BioMed Central
and we will help you at every step:
demethylases. 1. 3-Amino-4-pyridine carboxylate derivatives. J Med Chem. 2016;59(4):1357–69.
15. McGrath J, Trojer P. Targeting histone lysine methylation in cancer. Phar-macol Ther. 2015;150:1–22.
16. Wang L, Chang J, Varghese D, Dellinger M, Kumar S, Best AM, Ruiz J, Bruick R, Pena-Llopis S, Xu J, et al. A small molecule modulates Jumonji histone demethylase activity and selectively inhibits cancer growth. Nat Com-mun. 2013;4:2035.
17. Liang J, Zhang B, Labadie S, Ortwine DF, Vinogradova M, Kiefer JR, Gehling VS, Harmange JC, Cummings R, Lai T, et al. Lead optimization of a pyrazolo[1,5-a]pyrimidin-7(4H)-one scaffold to identify potent, selective and orally bioavailable KDM5 inhibitors suitable for in vivo biological studies. Bioorg Med Chem Lett. 2016;26(16):4036–41.
18. Torres IO, Kuchenbecker KM, Nnadi CI, Fletterick RJ, Kelly MJ, Fujimori DG. Histone demethylase KDM5A is regulated by its reader domain through a positive-feedback mechanism. Nat Commun. 2015;6:6204.
19. Horton JR, Upadhyay AK, Qi HH, Zhang X, Shi Y, Cheng X. Enzymatic and structural insights for substrate specificity of a family of Jumonji histone lysine demethylases. Nat Struct Mol Biol. 2010;17(1):38–43.
20. Martinez NJ, Simeonov A. Cell-based assays to support the profiling of small molecules with histone methyltransferase and demethylase modu-latory activity. Drug Discov Today Technol. 2015;18:9–17.
21. Kruidenier L, Chung CW, Cheng Z, Liddle J, Che K, Joberty G, Bantscheff M, Bountra C, Bridges A, Diallo H, et al. A selective Jumonji H3K27 dem-ethylase inhibitor modulates the proinflammatory macrophage response. Nature. 2012;488(7411):404–8.
22. Itoh Y, Sawada H, Suzuki M, Tojo T, Sasaki R, Hasegawa M, Mizukami T, Suzuki T. Identification of Jumonji AT-rich interactive domain 1A inhibitors and their effect on cancer cells. ACS Med Chem Lett. 2015;6(6):665–70. 23. Schiller R, Scozzafava G, Tumber A, Wickens JR, Bush JT, Rai G, Lejeune
C, Choi H, Yeh TL, Chan MC, et al. A cell-permeable ester derivative of the JmjC histone demethylase inhibitor IOX1. ChemMedChem. 2014;9(3):566–71.
24. Labelle M, Boesen T, Mehrotra M, Khan Q, Ullah F. Inhibitors of histone demethylases. International Patent WO2014053491A1, 2014.
25. Johansson C, Velupillai S, Tumber A, Szykowska A, Hookway ES, Nowak RP, Strain-Damerell C, Gileadi C, Philpott M, Burgess-Brown N, et al. Structural analysis of human KDM5B guides histone demethylase inhibitor develop-ment. Nat Chem Biol. 2016;12:539–45.
26. Horton JR, Engstrom A, Zoeller EL, Liu X, Shanks JR, Zhang X, Johns MA, Vertino PM, Fu H, Cheng X. Characterization of a linked Jumonji domain of the KDM5/JARID1 family of histone H3 lysine 4 demethylases. J Biol Chem. 2016;291(6):2631–46.
27. Luo X, Liu Y, Kubicek S, Myllyharju J, Tumber A, Ng S, Che KH, Podoll J, Heightman TD, Oppermann U, et al. A selective inhibitor and probe of the cellular functions of Jumonji C domain-containing histone demethylases. J Am Chem Soc. 2011;133(24):9451–6.
28. Vinogradova M, Gehling VS, Gustafson A, Arora S, Tindell CA, Wilson C, Williamson KE, Guler GD, Gangurde P, Manieri W, et al. An inhibitor of KDM5 demethylases reduces survival of drug-tolerant cancer cells. Nat Chem Biol. 2016;12(7):531–8.
29. Gehling VS, Bellon SF, Harmange JC, LeBlanc Y, Poy F, Odate S, Buker S, Lan F, Arora S, Williamson KE, et al. Identification of potent, selective KDM5 inhibitors. Bioorg Med Chem Lett. 2016;26(17):4350–4.
30. Hopkinson RJ, Tumber A, Yapp C, Chowdhury R, Aik W, Che KH, Li XS, Kristensen JB, King ON, Chan MC, et al. 5-Carboxy-8-hydroxyquinoline is a broad spectrum 2-oxoglutarate oxygenase inhibitor which causes iron translocation. Chem Sci. 2013;4(8):3110–7.
31. Heinemann B, Nielsen JM, Hudlebusch HR, Lees MJ, Larsen DV, Boesen T, Labelle M, Gerlach LO, Birk P, Helin K. Inhibition of demethylases by GSK-J1/J4. Nature. 2014;514(7520):E1–2.
32. Montenegro RC, Clark PG, Howarth A, Wan X, Ceroni A, Siejka P, Nunez-Alonso GA, Monteiro O, Rogers C, Gamble V, et al. BET inhibi-tion as a new strategy for the treatment of gastric cancer. Oncotarget. 2016;7(28):43997–4012.
33. Joberty G, Boesche M, Brown JA, Eberhard D, Garton NS, Mathieson T, Muelbaier M, Ramsden NG, Reader V, Rueger A, et al. Interrogating the druggability of the 2-oxoglutarate-dependent dioxygenase target class by chemical proteomics. ACS Chem Biol. 2016;11(7):2002–10. 34. Lillico R, Sobral MG, Stesco N, Lakowski TM. HDAC inhibitors induce
global changes in histone lysine and arginine methylation and alter expression of lysine demethylases. J Proteom. 2016;133:125–33. 35. Cheng MF, Lee CH, Hsia KT, Huang GS, Lee HS. Methylation of histone H3
lysine 27 associated with apoptosis in osteosarcoma cells induced by staurosporine. Histol Histopathol. 2009;24(9):1105–11.
36. Allera C, Lazzarini G, Patrone E, Alberti I, Barboro P, Sanna P, Melchiori A, Parodi S, Balbi C. The condensation of chromatin in apoptotic thymocytes shows a specific structural change. J Biol Chem. 1997;272(16):10817–22. 37. Scaffidi P, Misteli T, Bianchi ME. Release of chromatin protein HMGB1 by
necrotic cells triggers inflammation. Nature. 2002;418(6894):191–5. 38. Walter D, Matter A, Fahrenkrog B. Loss of histone H3 methylation at
lysine 4 triggers apoptosis in Saccharomyces cerevisiae. PLoS Genet. 2014;10(1):e1004095.
39. Pang B, Qiao X, Janssen L, Velds A, Groothuis T, Kerkhoven R, Nieuwland M, Ovaa H, Rottenberg S, van Tellingen O, et al. Drug-induced histone eviction from open chromatin contributes to the chemotherapeutic effects of doxorubicin. Nat Commun. 2013;4:1908.