Identification of Tilletia species using rep-PCR fingerprinting technique
Texto
Imagem

Documentos relacionados
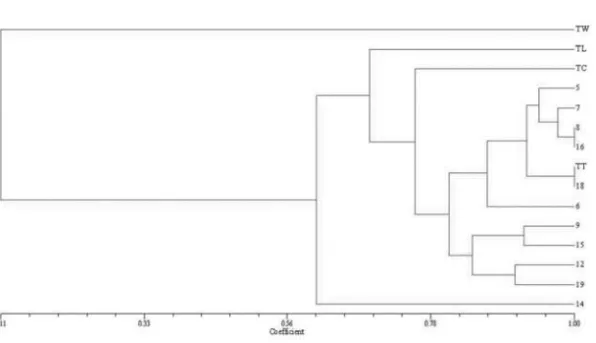
Combined analysis of the PCR patterns obtained with primers REP, ERIC and BOX, showed a high degree of similarity among Brazilian strains and the Indian type strain NCPPB 2475..
KEY WORDS: bites and stings, symptoms, species identification, neural network.. CONFLICTS OF INTEREST: There is
Unlike the analysis for morphological data, the absence of primer transferability between species (common zeros) are informative for species identification in the context of DNA
Para os caracteres morfoagronômicos, bromatológicos e minenalógicos estudados, verificou-se que os genótipos crioulos apresentaram, de modo geral, médias iguais
É importante salientar que ainda que um dos objetivos deste trabalho fosse utilizar os dados de vazão do SWAT como ferramenta para determinação de vazões mínimas de
Apêndice A: Análise de variação para o efeito dos fatores Adubação de pré-plantio e Cultivar sobre as variáveis de crescimento, produção e qualidade de
At the firm level, previous research suggests that the magnitude of the effect of the implementation of capital controls outweighs the impact of unforeseen
O presente trabalho pretende apontar a importância dos fatores relacionados ao Planejamento e à Logística aplicados à gestão pública, uma vez que tais fatores são