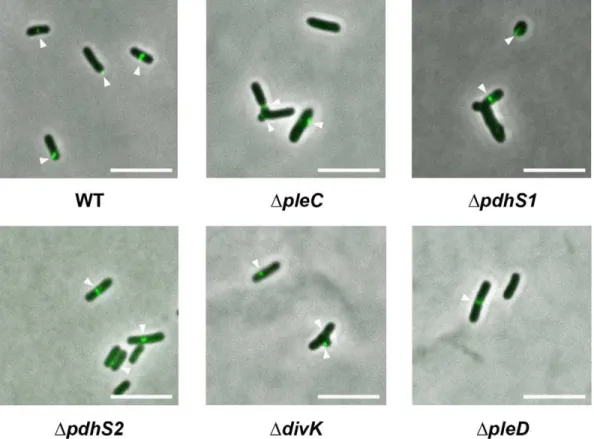
Coordination of division and development influences complex multicellular behavior in Agrobacterium tumefaciens.
Texto
Imagem

Documentos relacionados
Os objetivos específicos foram avaliar as propriedades físicas e mecânicas de chapas aglomeradas do tipo convencional e MDP, a influência da inclusão laminar e
«Começam a colher-se já os frutos do Concílio Ecuménico Vaticano II. […] Os fiéis, devidamente preparados, entram gostosamente no espírito e na letra da reforma litúrgica.
The recovered Candida species were simultaneously isolated from prawns and the aquatic environment and some of these isolates presented 100% similarity, even when
É nesta mudança, abruptamente solicitada e muitas das vezes legislada, que nos vão impondo, neste contexto de sociedades sem emprego; a ordem para a flexibilização como
As em endas que não reunirem o núm ero regim ental de assinaturas ou forem apresentadas fora d o prazo podem , entretanto, ser exam inedas pelos
Esta questão deixa em aberto a tipologia de ferramentas tecnológicas a utilizar. A opinião é quase unânime quer em relação às vantagens quer no que diz respeito
Bacteria were isolated from all wounds and 41 bacterial isolates could be identified based on culture of the materials collected by punch biopsy; 53.66% of the isolates
Desta forma, diante da importância do tema para a saúde pública, delineou-se como objetivos do es- tudo identificar, entre pessoas com hipertensão arterial, os