A network-based gene expression signature informs prognosis and treatment for colorectal cancer patients.
Texto
Imagem

Documentos relacionados
Gene Ontology (GO) and pathway enrichment analyses of DEGs were performed, followed by protein-protein interaction (PPI) network, transcriptional regulatory network as well
LYSO-7 treatment enhanced PPAR gene and protein expression in the stomach, and impaired local neutrophil influx and stasis of the microcirculatory network caused by
Combined analysis of E-cadherin gene (CDH1) promoter hypermethylation and E-cadherin protein expression in patients with gastric cancer: implications for treatment with demethylating
By using knowledge-based gene network techniques and promoter sequence analysis, we also uncovered mechanisms that might regulate the expression of CNS genes involved in the
Considering the limited gene expression signature differences (number of probes and genes) we could identify as influenced by the location of the tumor or by the type of
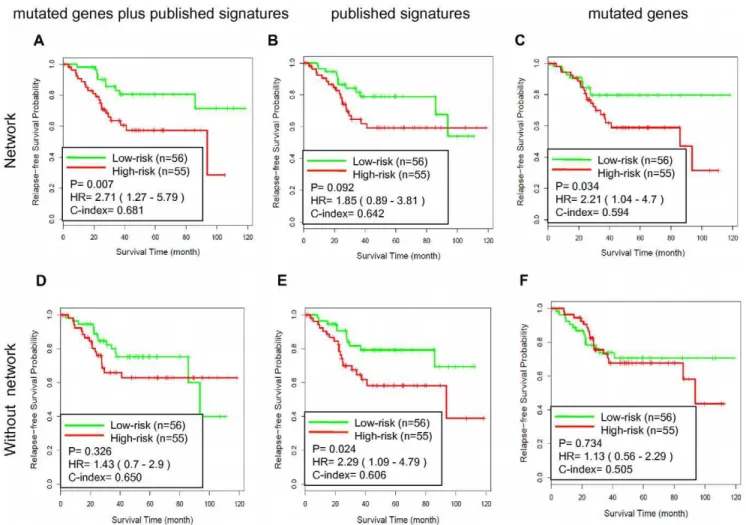
In this study, we developed a computational method to identify colorectal cancer-related genes based on (i) the gene expression profiles, and (ii) the shortest path analysis
We derived the expression profiles from 871 colorectal cancer patients with stage II and stage III samples to test associations between the 9-gene signature and clinical outcomes...
Keywords gene silencing; gene regulation; Boolean network; cell-cycle; gene expression; cellular phenotype..